Abstract
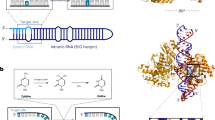
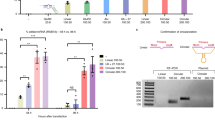
Site-directed RNA editing is an important technique for correcting gene sequences and ultimately tuning protein function. In this study, we engineered the deaminase domain of adenosine deaminase acting on RNA (ADAR1) and the MS2 system to target-specific adenosines, with the goal of correcting G-to-A mutations at the RNA level. For this purpose, the ADAR1 deaminase domain was fused downstream of the RNA-binding protein MS2, which has affinity for the MS2 RNA. To direct editing to specific targets, we designed guide RNAs complementary to target RNAs. The guide RNAs directed the ADAR1 deaminase to the desired editing site, where it converted adenosine to inosine. To provide proof of principle, we used an allele of enhanced green fluorescent protein (EGFP) bearing a mutation at the 58th amino acid (TGG), encoding Trp, into an amber (TAG) or ochre (TAA) stop codon. In HEK-293 cells, our system could convert stop codons to read-through codons, thereby turning on fluorescence. We confirmed the specificity of editing at the DNA level by restriction fragment length polymorphism analysis and sequencing, and at the protein level by western blotting. The editing efficiency of this enzyme system was ~5%. We believe that this system could be used to treat genetic diseases resulting from G-to-A point mutations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Kim H, Kim J-S . A guide to genome engineering with programmable nucleases. Nat Rev Genet 2014; 15: 321–334.
Doudna J . Perspective: embryo editing needs scrutiny. Nature 2015; 528: 230–236.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E . A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012; 337: 816–821.
Leisegang M, Engels B, Schreiber K, Yew PY, Kiyotani K, Idel C et al. Eradication of large solid tumors by gene therapy with a T-cell receptor targeting a single cancer-specific point mutation. Clin Cancer Res 2016; 22: 2734–2743.
Ferec C, Audrezet MP, Mercier B, Guillermit H, Moullier P, Quere I et al. Detection of over 98% cystic fibrosis mutations in a Celtic population. Nat Genet 1992; 1: 188–1891.
Bolscher BG, M de Boer, A de Klein, Weening RS, Roos D . Point mutations in the beta-subunit of cytochrome b558 leading to X-linked chronic granulomatous disease. Blood 1991; 77: 2482–2487.
Higuchi M, Kazazian HH, Kasch L, Warren TC, McGinniss MJ, Phillips JA et al. Molecular characterization of severe hemophilia A suggests that about half the mutations are not within the coding regions and splice junctions of the factor VIII gene. Proc Natl Acad Sci USA 1991; 88: 7405–7409.
Maas S, Rich A . Changing genetic information through RNA editing. BioEssays 2000; 22: 790–802.
Nishikura K . Functions and regulation of RNA editing by ADAR deaminases. Annu Rev Biochem 2010; 79: 321–349.
Nishikura K . A-to-I editing of coding and non-coding RNAs by ADARs. Nat Rev Mol Cell Biol 2016; 17: 83–96.
Slotkin W, Nishikura K . Adenosine-to-inosine RNA editing and human disease. Genome Med 2013; 5: 1–13.
Rosenberg BR, Hamilton CE, Mwangi MM, Dewell S, Papavasiliou FN . Transcriptome-wide sequencing reveals numerous APOBEC1 mRNA-editing targets in transcript 3′ UTRs. Nat Struct Mol Biol 2011; 18: 230–236.
Zabner J, Couture LA, Gregory RJ, Graham SM, Smith AE, Welsh MJ . Adenovirus-mediated gene transfer transiently corrects the chloride transport defect in nasal epithelia of patients with cystic fibrosis. Cell 1993; 75: 207–216.
Davies JC, Alton EW . Gene therapy for cystic fibrosis. Proc Am Thorac Soc 2010; 7: 408–14.
Blow MJ, Grocock RJ, van Dongen S, Enright AJ, Dicks E, Futreal PA et al. RNA editing of human microRNAs. Genome Biol 2006; 7: 27.1–27.8.
Kawahara Y, Zinshteyn B, Sethupathy P, Iizasa H, Hatzigeorgiou AG, Nishikura K . Redirection of silencing targets by adenosine-to-inosine editing of miRNAs. Science 2007; 315: 1137–1140.
Kim U, Wang Y, Sanford T, Zeng Y, Nishikura K . Molecular cloning of cDNA for double-stranded RNA adenosine deaminase, a candidate enzyme for nuclear RNA editing. Proc Natl Acad Sci USA 1994; 91: 11457–11461.
Melcher T, Maas S, Herb A, Sprengel R, Seeburg PH, Higuchi M . A mammalian RNA editing enzyme. Nature 1996; 379: 460–464.
Chen CX, Cho DS, Wang Q, Lai F, Carter KC, Nishikura K . A third member of the RNA-specific adenosine deaminase gene family, ADAR3, contains both single and double-stranded RNA binding domains. RNA 2000; 6: 755–767.
Schneider MF, Wettengel J, Hoffmann PC, Stafforst T . Optimal guideRNAs for re-directing deaminase activity of hADAR1 and hADAR2 in trans. Nucleic Acids Res 2014; 42: 1–9.
Stefl R, Xu M, Skrisovska L, Emeson RB, Allain FH . Structure and specific RNA binding of ADAR2 double-stranded RNA binding motifs. Structure 2006; 14: 345–355.
Herbert A, Alfken J, Kim YG, Mian IS, Nishikura K, Rich A . A Z-DNA binding domain present in the human editing enzyme, double-stranded RNA adenosine deaminase. Proc Natl Acad Sci USA 1997; 94: 8421–8426.
Cho DSC, Yang WD, Lee JT, Shiekhattar R, Murray JM, Nishikura K . Requirement of dimerization for RNA editing activity of adenosine deaminases acting on RNA. J Biol Chem 2003; 278: 17093–17102.
Valente L, Nishikura K . RNA binding-independent dimerization of adenosine deaminases acting on RNA and dominant negative effects of nonfunctional subunits on dimer functions. J Biol Chem 2007; 282: 16054–16061.
Ota H, Sakurai M, Gupta R, Valente L, Wulff BE, Ariyoshi K et al. ADAR1 forms a complex with dicer to promote microRNA processing and RNA-induced gene silencing. Cell 2013; 153: 575–589.
Chen L, Li Y, Lin CH, Chan TH, Chow RK, Song Y et al. Recoding RNA editing of AZIN1 predisposes to hepatocellular carcinoma. Nat Med 2013; 19: 209–216.
Shimokawa T, Rahman MF, Tostar U, Sonkoly E, Stahle M, Pivarcsi A et al. RNA editing of the GLI1 transcription factor modulates the output of Hedgehog signaling. RNA Biol 2013; 10: 321–333.
Paz-Yaacov N, Bazak L, Buchumenski I, Porath HT, Danan-Gotthold M, Knisbacher BA et al. Elevated RNA editing activity is a major contributor to transcriptomic diversity in tumors. Cell Rep 2015; 13: 267–276.
Han L, Diao L, Yu S, Xu X, Li J, Zhang R et al. The genomic landscape and clinical relevance of A-to-I RNA editing in human cancers. Cancer Cell 2015; 28: 515–528.
Morabito MV, Abbas AI, Hood JL, Kesterson RA, Jacobs MM, Kump DS et al. Mice with altered serotonin 2C receptor RNA editing display characteristics of Prader–Willi syndrome. Neurobiol Dis 2010; 39: 169–180.
Silberberg G, Lundin D, Navon R, Ohman M . Deregulation of the A-to-I RNA editing mechanism in psychiatric disorders. Hum Mol Genet 2012; 21: 311–321.
Hartner JC, Schmittwolf C, Kispert A, Muller AM, Higuchi M, Seeburg PH . Liver disintegration in the mouse embryo caused by deficiency in the RNA-editing enzyme ADAR1. J Biol Chem 2004; 279: 4894–4902.
Wang Q, Miyakoda M, Yang W, Khillan J, Stachura DL, Weiss MJ et al. Stress-induced apoptosis associated with null mutation of ADAR1 RNA editing deaminase gene. J Biol Chem 2004; 279: 4952–4961.
Zhu C, Zhu K, Zhou Y, Fan Y . A novel insertion mutation in the ADAR1 gene of a Chinese family with dyschromatosis symmetrica hereditaria. Genet Mol Res 2013; 12: 2858–2862.
Vogel P, Stafforst T . Site-directed RNA editing with antagomir deaminases-a tool to study protein and RNA function. Chem Med Chem 2014; 9: 2021–2025.
Wettengel J, Reautschnig P, Geisler S, Kahle PJ, Stafforst T . Harnessing human ADAR2 for RNA repair–recoding a PINK1 mutation rescues mitophagy. Nucleic Acids Res 2017; 45: 2797–2808.
Woolf TM, Chase JM, Stinchcomb DT . Toward the therapeutic editing of mutated RNA sequences. Proc Natl Acad Sci USA 1995; 92: 8298–8302.
Montiel-Gonzalez MF, Vallecillo-Viejo I, Yudowski GA, Rosenthal JJ . Correction of mutations within the cystic fibrosis transmembrane conductance regulator by site-directed RNA editing. Proc Natl Acad Sci USA 2013; 110: 18285–18290.
Hanswillemenke A, Kuzdere T, Vogel P, Jekely G, Stafforst T . Site-directed RNA 11editing in vivo can be triggered by the light-driven assembly of an artificial riboprotein. J Am Chem Soc 2015; 137: 15875–15881.
Vu LT, Nguyen TTK, Thoufic AAM, Suzuki H, Tsukahara T . Chemical RNA editing for genetic restoration: the relationship between the structure and deamination efficiency of carboxyvinyldeoxyuridine oligodeoxynucleotides. Chem Biol Drug Des 2016; 87: 583–593.
Vu LT, Nguyen TTK, Alam S, Sakamoto T, Fujimoto K, Suzuki H et al. Changing blue fluorescent protein to green fluorescent protein using chemical RNA editing as a novel strategy in genetic restoration. Chem Biol Drug Des 2015; 86: 1242–1252.
Keryer-Bibens C, Barreau C, Osborne HB . Tethering of proteins to RNAs by bacteriophage proteins. Biol Cell 2008; 100: 125–138.
Buxbaum AR, Haimovich G, Singer RH . In the right place at the right time: visualizing and understanding mRNA localization. Nat Rev Mol Cell Biol 2015; 16: 95–109.
Desterro JMP, Keegan LR, Lafarga M, Berciano MT, O'Connell M, Carmo-Fonseca M . Dynamic association of RNA-editing enzymes with the nucleolus. J Cell Sci 2003; 116: 1805–1818.
Strehblow A, Hallegger M, Jantsch MF . Nucleocytoplasmic distribution of human RNA-editing enzyme ADAR1 is modulated by double-stranded RNA-binding domains, a leucine-rich export signal, and a putative dimerization domain. Mol Biol Cell 2002; 13: 3822–3835.
Nurpeisov V, Hurwitz SJ, Sharma PL . Fluorescent dye terminator sequencing methods for quantitative determination of replication fitness of human immunodeficiency virus type 1 containing the codon 74 and 184 mutations in reverse transcriptase. J Clin Microbiol 2003; 41: 3306–3311.
Eggington JM, Greene T, Bass BL . Predicting sites of ADAR editing in double-stranded RNA. Nat Commun 2011; 2: 1–9.
Rinkevich FD, Schweitzer PA, Scott JG . Antisense sequencing improves the accuracy and precision of A-to-I editing measurements using the peak height ratio method. BMC Res Notes 2012; 5: 1–6.
Hakim NH, Kounishi T, Alam AH, Tsukahara T, Suzuki H . Alternative splicing of Mef2c promoted by Fox-1 during neural differentiation in P19 cells. Genes Cells 2010; 15: 255–67.
Acknowledgements
We gratefully acknowledge scholarships from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT), Japan. This research was supported in part by a Grant-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (25290072 and 26670167 to TT).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Gene Therapy website
Supplementary information
Rights and permissions
About this article
Cite this article
Azad, M., Bhakta, S. & Tsukahara, T. Site-directed RNA editing by adenosine deaminase acting on RNA for correction of the genetic code in gene therapy. Gene Ther 24, 779–786 (2017). https://doi.org/10.1038/gt.2017.90
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gt.2017.90
This article is cited by
-
Programmable RNA base editing via targeted modifications
Nature Chemical Biology (2024)
-
RNA Editing Therapeutics: Advances, Challenges and Perspectives on Combating Heart Disease
Cardiovascular Drugs and Therapy (2023)
-
Site-directed RNA editing by harnessing ADARs: advances and challenges
Functional & Integrative Genomics (2022)
-
RNA editing of BFP, a point mutant of GFP, using artificial APOBEC1 deaminase to restore the genetic code
Scientific Reports (2020)
-
In vivo RNA editing of point mutations via RNA-guided adenosine deaminases
Nature Methods (2019)