Abstract
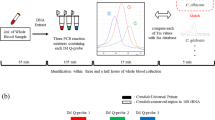
Invasive fungal disease (IFD) is a life-threatening event in immunocompromised patients, and there is an urgent need for reliable screening methods facilitating rapid and broad detection of pathogenic fungi. We have established a two-reaction real-time PCR assay permitting highly sensitive detection of more than 80 fungal pathogens, covering a large spectrum of moulds, yeasts and Zygomycetes. To assess the clinical potential of the assay, more than 600 consecutive specimens from 125 pediatric patients carrying a high risk of IFD were analyzed. An excellent correlation between PCR positivity and the presence of proven, probable or possible fungal infection according to the European Organization for Research and Treatment of Cancer criteria was demonstrated, as revealed by the sensitivity of the assay of 96% (95% CI: 82–99%). The negative predictive value of the panfungal PCR assay presented was 98% (95% CI: 90–100%), while the specificity and the positive predictive value were 77% (95% CI: 66–85%) and 62% (95% CI: 47–75%), respectively. The results indicate that molecular screening of patients during febrile neutropenic episodes by the assay presented could help prevent unnecessary toxicity resulting from empirical antifungal treatment in individuals who may not be at risk of imminent fungal disease. Our observations raise the possibility that rapid species identification may be required to increase the positive predictive value for impending fungus-related disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Enoch DA, Ludlam HA, Brown NM . Invasive fungal infections: a review of epidemiology and management options. J Med Microbiol 2006; 55: 809–818.
Kauffman CA . Fungal infections. Proc Am Thorac Soc 2006; 3: 35–40.
Walsh TJ, Groll A, Hiemenz J, Fleming R, Roilides E, Anaissie E . Infections due to emerging and uncommon medically important fungal pathogens. Clin Microbiol Infect 2004; 10 (Suppl 1): 48–66.
Warnock DW . Trends in the epidemiology of invasive fungal infections. Nippon Ishinkin Gakkai Zasshi 2007; 48: 1–12.
Richardson M, Lass-Florl C . Changing epidemiology of systemic fungal infections. Clin Microbiol Infect 2008; 14 (Suppl 4): 5–24.
Golan Y . Overview of transplant mycology. Am J Health Syst Pharm 2005; 62: S17–S21.
Pfaller MA, Diekema DJ, Gibbs DL, Newell VA, Bijie H, Dzierzanowska D et al. Results from the ARTEMIS disk global antifungal surveillance study, 1997 to 2007: 10.5-year analysis of susceptibilities of noncandidal yeast species to fluconazole and voriconazole determined by CLSI standardized disk diffusion testing. J Clin Microbiol 2009; 47: 117–123.
Nucci M, Spector N, Bueno AP, Solza C, Perecmanis T, Bacha PC et al. Risk factors and attributable mortality associated with superinfections in neutropenic patients with cancer. Clin Infect Dis 1997; 24: 575–579.
Rekha A, Kindo AJ, Ravi A . Fusarium solani in the post-transplant patient: an unusual fungus. Int J Low Extrem Wounds 2008; 7: 38–40.
Jayamohan Y, Ribes JA . Pseudallescheriasis: a summary of patients from 1980-2003 in a tertiary care center. Arch Pathol Lab Med 2006; 130: 1843–1846.
Cortez KJ, Roilides E, Quiroz-Telles F, Meletiadis J, Antachopoulos C, Knudsen T et al. Infections caused by Scedosporium spp. Clin Microbiol Rev 2008; 21: 157–197.
Malani AN, Kauffman CA . Changing epidemiology of rare mould infections: implications for therapy. Drugs 2007; 67: 1803–1812.
Dunyach C, Bertout S, Phelipeau C, Drakulovski P, Reynes J, Mallie M . Detection and identification of Candida spp. in human serum by LightCycler(R) real-time polymerase chain reaction. Diagn Microbiol Infect Dis 2008; 60: 263–271.
White PL, Shetty A, Barnes RA . Detection of seven Candida species using the Light-Cycler system. J Med Microbiol 2003; 52: 229–238.
Costa C, Costa JM, Desterke C, Botterel F, Cordonnier C, Bretagne S . Real-time PCR coupled with automated DNA extraction and detection of galactomannan antigen in serum by enzyme-linked immunosorbent assay for diagnosis of invasive aspergillosis. J Clin Microbiol 2002; 40: 2224–2227.
Kami M, Fukui T, Ogawa S, Kazuyama Y, Machida U, Tanaka Y et al. Use of real-time PCR on blood samples for diagnosis of invasive aspergillosis. Clin Infect Dis 2001; 33: 1504–1512.
Baskova L, Landlinger C, Preuner S, Lion T . The Pan-AC assay: a single-reaction real-time PCR test for quantitative detection of a broad range of Aspergillus and Candida species. J Med Microbiol 2007; 56: 1167–1173.
Jordanides NE, Allan EK, McLintock LA, Copland M, Devaney M, Stewart K et al. A prospective study of real-time panfungal PCR for the early diagnosis of invasive fungal infection in haemato-oncology patients. Bone Marrow Transplant 2005; 35: 389–395.
Klingspor L, Jalal S . Molecular detection and identification of Candida and Aspergillus spp. from clinical samples using real-time PCR. Clin Microbiol Infect 2006; 12: 745–753.
Schabereiter-Gurtner C, Selitsch B, Rotter ML, Hirschl AM, Willinger B . Development of novel real-time PCR assays for detection and differentiation of eleven medically important Aspergillus and Candida species in clinical specimens. J Clin Microbiol 2007; 45: 906–914.
Vollmer T, Stormer M, Kleesiek K, Dreier J . Evaluation of novel broad-range real-time PCR assay for rapid detection of human pathogenic fungi in various clinical specimens. J Clin Microbiol 2008; 46: 1919–1926.
Tolstrup N, Nielsen PS, Kolberg JG, Frankel AM, Vissing H, Kauppinen S . OligoDesign: optimal design of LNA (locked nucleic acid) oligonucleotide capture probes for gene expression profiling. Nucleic Acids Res 2003; 31: 3758–3762.
Watzinger F, Suda M, Preuner S, Baumgartinger R, Ebner K, Baskova L et al. Real-time quantitative PCR assays for detection and monitoring of pathogenic human viruses in immunosuppressed pediatric patients. J Clin Microbiol 2004; 42: 5189–5198.
Altmann DG, Machin D, Bryant TN, Gardner MJ . Statistics with Confidence, 2nd edn. BMJ Books, 2000.
De Pauw B, Walsh TJ, Donnelly JP, Stevens DA, Edwards JE, Calandra T et al. Revised definitions of invasive fungal disease from the European Organization for Research and Treatment of Cancer/Invasive Fungal Infections Cooperative Group and the National Institute of Allergy and Infectious Diseases Mycoses Study Group (EORTC/MSG) Consensus Group. Clin Infect Dis 2008; 46: 1813–1821.
Hebart H, Loffler J, Reitze H, Engel A, Schumacher U, Klingebiel T et al. Prospective screening by a panfungal polymerase chain reaction assay in patients at risk for fungal infections: implications for the management of febrile neutropenia. Br J Haematol 2000; 111: 635–640.
Boudewijns M, Verweij PE, Melchers WJ . Molecular diagnosis of invasive aspergillosis: the long and winding road. Future Microbiol 2006; 1: 283–293.
Buchheidt D, Baust C, Skladny H, Ritter J, Suedhoff T, Baldus M et al. Detection of Aspergillus species in blood and bronchoalveolar lavage samples from immunocompromised patients by means of 2-step polymerase chain reaction: clinical results. Clin Infect Dis 2001; 33: 428–435.
Cuenca-Estrella M, Meije Y, Diaz-Pedroche C, Gomez-Lopez A, Buitrago MJ, Bernal-Martinez L et al. Value of serial quantification of fungal DNA by a real-time PCR-based technique for early diagnosis of invasive Aspergillosis in patients with febrile neutropenia. J Clin Microbiol 2009; 47: 379–384.
Mengoli C, Cruciani M, Barnes RA, Loeffler J, Donnelly JP . Use of PCR for diagnosis of invasive aspergillosis: systematic review and meta-analysis. Lancet Infect Dis 2009; 9: 89–96.
White PL, Barton R, Guiver M, Linton CJ, Wilson S, Smith M et al. A consensus on fungal polymerase chain reaction diagnosis?: a United Kingdom-Ireland evaluation of polymerase chain reaction methods for detection of systemic fungal infections. J Mol Diagn 2006; 8: 376–384.
White PL, Linton CJ, Perry MD, Johnson EM, Barnes RA . The evolution and evaluation of a whole blood polymerase chain reaction assay for the detection of invasive aspergillosis in hematology patients in a routine clinical setting. Clin Infect Dis 2006; 42: 479–486.
Aimanianda V, Bayry J, Bozza S, Kniemeyer O, Perruccio K, Elluru SR et al. Surface hydrophobin prevents immune recognition of airborne fungal spores. Nature 2009; 460: 1117–1121.
Pastor FJ, Guarro J . Alternaria infections: laboratory diagnosis and relevant clinical features. Clin Microbiol Infect 2008; 14: 734–746.
Chen YC, Eisner JD, Kattar MM, Rassoulian-Barrett SL, LaFe K, Yarfitz SL et al. Identification of medically important yeasts using PCR-based detection of DNA sequence polymorphisms in the internal transcribed spacer 2 region of the rRNA genes. J Clin Microbiol 2000; 38: 2302–2310.
De Baere T, Claeys G, Swinne D, Verschraegen G, Muylaert A, Massonet C et al. Identification of cultured isolates of clinically important yeast species using fluorescent fragment length analysis of the amplified internally transcribed rRNA spacer 2 region (ITS2). BMC Microbiol 2002; 2: 21.
Landlinger C, Preuner S, Willinger B, Haberpursch B, Racil Z, Mayer J et al. Species-specific identification of a wide range of clinically relevant fungal pathogens by the Luminex xMAPTM technology. J Clin Microbiol 2009; 47: 1063–1073.
Turenne CY, Sanche SE, Hoban DJ, Karlowsky JA, Kabani AM . Rapid identification of fungi by using the ITS2 genetic region and an automated fluorescent capillary electrophoresis system. J Clin Microbiol 1999; 37: 1846–1851.
Das S, Brown TM, Kellar KL, Holloway BP, Morrison CJ . DNA probes for the rapid identification of medically important Candida species using a multianalyte profiling system. FEMS Immunol Med Microbiol 2006; 46: 244–250.
Diaz MR, Fell JW . High-throughput detection of pathogenic yeasts of the genus trichosporon. J Clin Microbiol 2004; 42: 3696–3706.
Boyanton Jr BL, Luna RA, Fasciano LR, Menne KG, Versalovic J . DNA pyrosequencing-based identification of pathogenic Candida species by using the internal transcribed spacer 2 region. Arch Pathol Lab Med 2008; 132: 667–674.
Putignani L, Paglia MG, Bordi E, Nebuloso E, Pucillo LP, Visca P . Identification of clinically relevant yeast species by DNA sequence analysis of the D2 variable region of the 25-28S rRNA gene. Mycoses 2008; 51: 209–227.
Campa D, Tavanti A, Gemignani F, Mogavero CS, Bellini I, Bottari F et al. DNA microarray based on arrayed-primer extension technique for identification of pathogenic fungi responsible for invasive and superficial mycoses. J Clin Microbiol 2008; 46: 909–915.
Spiess B, Seifarth W, Hummel M, Frank O, Fabarius A, Zheng C et al. DNA microarray-based detection and identification of fungal pathogens in clinical samples from neutropenic patients. J Clin Microbiol 2007; 45: 3743–3753.
Zeng X, Kong F, Halliday C, Chen S, Lau A, Playford G et al. Reverse line blot hybridization assay for identification of medically important fungi from culture and clinical specimens. J Clin Microbiol 2007; 45: 2872–2880.
Gangneux JP, Lavarde D, Bretagne S, Guiguen C, Gandemer V . Transient aspergillus antigenaemia: think of milk. Lancet 2002; 359: 1251.
Steinbach WJ . Pediatric aspergillosis: disease and treatment differences in children. Pediatr Infect Dis J 2005; 24: 358–364.
Murashige N, Kami M, Kishi Y, Fujisaki G, Tanosaki R . False-positive results of Aspergillus enzyme-linked immunosorbent assays for a patient with gastrointestinal graft-versus-host disease taking a nutrient containing soybean protein. Clin Infect Dis 2005; 40: 333–334.
Adam O, Auperin A, Wilquin F, Bourhis JH, Gachot B, Chachaty E . Treatment with piperacillin-tazobactam and false-positive Aspergillus galactomannan antigen test results for patients with hematological malignancies. Clin Infect Dis 2004; 38: 917–920.
Zandijk E, Mewis A, Magerman K, Cartuyvels R . False-positive results by the platelia Aspergillus galactomannan antigen test for patients treated with amoxicillin-clavulanate. Clin Vaccine Immunol 2008; 15: 1132–1133.
Acknowledgements
This work was supported by grants from the Austrian Center for Innovation and Technology (ZIT), the Austrian Science Fund (FWF; Project No. P16929-B13) and Fonds der Stadt Wien für innovative interdisziplinäre Krebsforschung. We are grateful to Pia Reindl for collecting clinical data relating to the patient specimens obtained from the St Anna Children's Hospital, Vienna. Furthermore, we want to thank Ulrike Pötschger for her help with statistical analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Landlinger, C., Preuner, S., Bašková, L. et al. Diagnosis of invasive fungal infections by a real-time panfungal PCR assay in immunocompromised pediatric patients. Leukemia 24, 2032–2038 (2010). https://doi.org/10.1038/leu.2010.209
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2010.209
Keywords
This article is cited by
-
Clinical evaluation of an in-house panfungal real-time PCR assay for the detection of fungal pathogens
Infection (2020)
-
Detection and identification of fungi in bronchoalveolar lavage fluid from immunocompromised patients using panfungal PCR
Folia Microbiologica (2019)
-
Progress in the Diagnosis of Invasive Fungal Disease in Children
Current Fungal Infection Reports (2017)
-
Similar efficacy of broad-range ITS PCR and conventional fungal culture for diagnosing fungal infections in non-immunocompromised patients
BMC Microbiology (2016)
-
Molecular Diagnosis in Fungal Infection Control
Current Treatment Options in Infectious Diseases (2015)