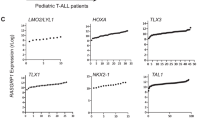
Abstract
The RUNX1/ETO (RE) fusion protein, which originates from the t(8;21) chromosomal rearrangement, is one of the most frequent translocation products found in de novo acute myeloid leukemia (AML). In RE leukemias, activated forms of the c-KIT tyrosine kinase receptor are frequently found, thereby suggesting oncogenic cooperativity between these oncoproteins in the development and maintenance of t(8;21) malignancies. In this report, we show that activated c-KIT cooperates with a C-terminal truncated variant of RE, REtr, to expand human CD34+ hematopoietic progenitors ex vivo. CD34+ cells expressing both oncogenes resemble the AML-M2 myeloblastic cell phenotype, in contrast to REtr-expressing cells which largely undergo granulocytic differentiation. Oncogenic c-KIT amplifies REtr-depended clonogenic growth and protects cells from exhaustion. Activated c-KIT reverts REtr-induced DNA damage and apoptosis. In the presence of activated c-KIT, REtr-downregulated DNA-repair genes are re-expressed leading to an enhancement of DNA-repair efficiency via homologous recombination. Together, our results provide new mechanistic insight into REtr and c-KIT oncogenic cooperativity and suggest that augmented DNA repair accounts for the increased chemoresistance observed in t(8;21)-positive AML patients with activated c-KIT mutations. This cell-protective mechanism might represent a new therapeutic target, as REtr cells with activated c-KIT are highly sensitive to pharmacological inhibitors of DNA repair.
Similar content being viewed by others
Introduction
Chromosomal translocations involving members of the core-binding factor (CBF) family are among the most frequent cytogenetic abnormalities found in acute myeloid leukemia (AML). The best characterized translocation involving CBF family members is the translocation t(8;21). In this translocation, the hematopoietic master regulator gene RUNX1 (also known as AML1, CBFα2 or PEBP2αB) on chromosome 21 is fused to almost the entire ETO gene (also known as MTG8 or RUNX1T1) on chromosome 8, thereby generating the fusion protein RUNX1/ETO (RE). RE retains the DNA-binding domain of RUNX1, while ETO provides multiple docking sites for members of the nuclear corepressor families such as N-CoR, mSin3, SMRT and various histone deacetylases and thus acts primarily as a transcriptional repressor of RUNX1 target genes leading to an epigenetic-driven block of myeloid differentiation.1, 2, 3 As the translocation t(8;21) has been found in utero and in the blood of newborns several years before disease onset, additional genetic alterations must occur for full blast cell transformation.4 Indeed, leukemia development has not been observed in immunocompromised mice transplanted with human hematopoietic CD34+ precursors expressing RE.5
Genetic aberrations affecting transcription factors and genes involved in signal transduction represent two classes of the most frequently detected mutational events in human leukemia. Murine experiment data have favored a model of AML pathogenesis in which the two groups of genetic alterations are required for the induction of full-blown disease. In human t(8;21)+ leukemia samples, mutations in receptor tyrosine kinase genes, primarily c-KIT and to a lesser extent FLT3, or mutations in N-RAS are recurrently found,6 suggesting that RE requires altered signal transduction pathways for leukemia development. In particular, c-KIT mutations are found at high frequencies (up to 48%) in AML samples harboring the translocation t(8;21), and they are associated with a high incidence of relapse after chemotherapy.6, 7, 8 Moreover, cooperativity between RE and activated forms of c-KIT in the induction of AML has been recently demonstrated using mouse transplantation models.9, 10
The most common class of c-KIT mutations in t(8;21)-associated AML occurs in the activation loop (A-loop) of the kinase domain, which has an auto-inhibitory function, most frequently resulting in D816V or N822K mutations.11 High expression levels of activated c-KIT mutants correlate with poor disease outcome, especially when co-expressed with a C-terminal truncated splice variant form of RE, referred to as RE9a.6 AML patients with high levels of both RE9a and activated c-KIT exhibit a significantly shorter event-free and overall survival time compared with patients with low RE9a expression levels.6
In this work, we analyzed potential oncogenic cooperativity between mutated c-KIT and a truncated variant of RE (REtr) lacking the C-terminal NHR3 and NHR4 domains. We show a robust oncogenic cooperativity between activated forms of c-KIT and REtr in human CD34+ progenitor cells using a recently published ex vivo progenitor cell expansion assay.5, 12, 13 Together with REtr, c-KIT(N822K) expression blocks cellular differentiation of progenitor cells at the myeloblastic stage, enhances long-term culture and clonogenic growth and protects cells from RE-induced DNA damage and apoptosis by activating the DNA-damage repair machinery.
Materials and methods
Cloning of MSCV vectors
The expression plasmid MSCV-REtr-IRES-eGFP has been described previously.14 In this plasmid, a tandem Tomato (tdTomato) cDNA was inserted to replace eGFP, thereby resulting in MSCV-REtr-IRES-tdTomato. C-KIT A-loop mutations were generated via site-directed mutagenesis (Stratagene, La Jolla, CA, USA). All constructs were verified via sequence analysis.
Cell culture and retroviral transduction
Kasumi-1, 293T and U937 cells were cultured as previously described.14 G-CSF (granulocyte colony-stimulating factor)-mobilized peripheral blood CD34+ cells were cultured in Iscove’s modified Dulbecco’s medium (Life Technologies, Karlsruhe, Germany) supplemented with 20% fetal calf serum, 20 ng/ml Flt-3 l, 20 ng/ml GM-CSF, 20 ng/ml stem cell factor (SCF), 20 ng/ml thrombopoietin, 20 ng/ml interleukin (IL)-6, 10 ng/ml IL-3 (all cytokines were obtained Peprotech, Hamburg, Germany), 100 U/ml penicillin/streptomycin and 2 mM L-glutamine. Retroviral transduction and long-term cultivation were performed as previously described.5 IMR-90 cells (ATCC-CCL-186) were cultured as suggested by the supplier.
Analysis of genomic instability
Cells were analyzed as described previously.15 Briefly, cells were fixed in 2% paraformaldehyde and permeabilized for 5 min in phosphate-buffered saline with 0.1% Triton-X-100. The primary antibody anti-γ-H2AX 3F2 antibody (1:200; Abcam, Cambridge, Great Britain) was used. Alexa 488-conjugated donkey anti-mouse (1:400; Invitrogen, Carlsbad, CA, USA) or Alexa 546-conjugated goat anti-mouse (1:200; Invitrogen) were used as secondary antibodies. For γ-tubulin staining, we used the ab11317 antibody (Abcam) at a 1:400 dilution and Alexa 546-conjugated goat anti-mouse antibody as the secondary antibody (1:200; Invitrogen). DNA was counterstained with 50 ng/ml DAPI (4,6-diamidino-2-phenylindole; Invitrogen). For γ-H2AX and Rad51 repair kinetics, cells were irradiated with 4 Gy (γ-irradiation) and incubated for 2, 4, 8 and 24 h at 37 °C followed by fixation and staining as described above.
Fluorescence-activated cell sorting (FACS) analysis, cell cycle and apoptosis assays
For the analysis of cell surface markers, we used FITC (fluorescein isothiocyanate)-, phycoerythrin-, phycoerythrin-Cy7 or allophycocyanin-conjugated anti-human HLA-DR, CD3, CD10, CD11b, CD11c, CD13, CD14, CD15, CD19, CD33, CD34, CD38, CD41a and CD117 antibody as well as mouse monoclonal immunoglobulin G1 or mouse immunoglobulin G1 isotype control antibodies (all obtained from BD Pharmingen, Heidelberg, Germany). For cell cycle analysis, cells were incubated for 15 min with 2 μM DRAQ5 (Alexis Biochemicals, San Diego, CA, USA) at 37 °C followed by FACS analysis.14
Analysis of homologous recombination
Homologous recombination events were measured essentially as described in Epinat et al.16 Briefly, IMR-90 cells expressing REtr or REtr+c-KIT(N822K) were transfected with a plasmid containing a split β-galactosidase gene. After TALEN-induced double-strand break, proper homologous recombination rescued β-galactosidase activity.
Transcriptome analysis
G-CSF-mobilized peripheral blood CD34+ cells (LONZA, Walkersville, MD, USA) were transduced with the MSCV-c-KIT(N822K) vector or mock-transduced and eGFP+ cells were FACS sorted 6 days after transduction. Stably transduced U937 cells were FACS sorted before analysis. Total RNA (input: 40 ng) from sorted cells (>90% purity) was amplified and cDNA labeled using standardized protocols. Microarray hybridization (4 × 44 K human GE Array, Agilent (Böblingen, Germany) (CD34+ cells) or GeneChip Human Gene 1.0 ST V1 arrays, Affymetrix (Santa Clara, CA, USA), (U937 cells)), washing steps and scanning of the microarrays were performed according to the suppliers’ protocol. Statistical analysis was done either with the statistical computing environment R version 2.12.17 or DAVID (http://david.abcc.ncifcrf.gov/). Additional software packages were taken from the Bioconductor project.18
Results
RUNX1/ETOtr and c-KIT(N822K) cooperate in the ex vivo expansion of CD34+ hematopoietic progenitor cells
The t(8;21)-associated gene product RE and the activated mutant of c-KIT (c-KIT(N822K)) were stably expressed in primary human CD34+ cells using retroviral vectors co-expressing fluorescence marker genes, which allowed for continuous detection and analysis of gene-modified cells during ex vivo log-term culture (Figure 1a). In our studies, we used a truncated version of RE, REtr, as a recent report has shown a strong correlation between the occurrence of activating mutations in c-KIT and high expression levels of RE9A, an equivalent splice variant of RE lacking the C-terminal NHR3 and NHR4 domains of ETO.6, 19 We selected the c-KIT mutation N822K for our studies, as N822K represents the most frequent individual mutation among all c-KIT mutations found in t(8;21) AML (Supplementary Figure S1a).6 Co-expression of REtr and c-KIT(N822K) in human CD34+ enriched progenitor cells led to a robust and highly reproducible SCF-dependent outgrowth of cells expressing both oncogenes in ex vivo expansion cultures, which resulted in an almost pure population (>90%) of REtr+c-KIT(N822K) double-positive cells (Figures 1b and c and Supplementary Figure S1b). Similar results were obtained with full-length RUNX1/ETO (data not shown). In contrast, CD34+ cells transduced with c-KIT(N822K) alone failed to expand and exhibited a behavior similar to mock-transduced cells (Figure 1c). The oncogenic cooperativity could not be reproduced with wild-type c-KIT, suggesting a critical role of the activating mutation N822K in CD34+ expansion (Figure 1d). Substitution of amino-acid 568 (Y568F), a juxtamembrane amino acid critical for kinase activation, completely abolished the function of c-KIT(N822K), thereby indicating that proper kinase activity is required for the observed effects of c-KIT(N822K) (Supplementary Figure S1c). Similar to the effects observed with c-KIT(N822K), other c-KIT-activating mutations also conferred oncogenic support of REtr-enhanced CD34+ cell expansion. The c-KIT A-loop mutations D816V, A814S and V825A (Figure 1e and Supplementary Figure S1a) also cooperated with REtr in enhancing the expansion of double-transduced cells to almost 100% from an initial minor fraction (1%) of REtr and c-KIT-mutant co-expressing cells (Figure 1f). In contrast, the c-KIT mutation L793S, which has only been described in a single t(8;21) patient, did not confer oncogenic cooperativity (Figures 1e and f, Supplementary Figure S1a and Jiao et al.6). Similar to the N822K mutant, the c-KIT mutants V825A, D816V and A814S did not generate an outgrowth of transduced CD34+ cells (Supplementary Figure S2a). Similarly, co-expression of FLT3-ITD, which is rarely found in t(8;21)+ leukemia, or an activated form of signal transducer and activator of transcription factor 5 (STAT5) did not enhance REtr-mediated growth of transduced cells. Under these conditions, only REtr single-transduced cells expanded in culture (Supplementary Figure S2b). Oncogenic cooperativity could also be observed in a stepwise expansion and selection process. REtr-expressing cells were also transduced with either c-KIT(N822K) or activated forms of STAT3 (caSTAT3) or STAT5 (caSTAT5). Only c-KIT(N822K)- and REtr-expressing cells expanded in culture and overgrew single REtr-transduced cells 26 days after transduction (Supplementary Figures S2c and d). This selection process for REtr- and c-KIT(N822K)-expressing cells could be further enhanced with a low-dose AraC treatment, a chemotherapeutic drug that has been shown to induce cellular differentiation at low concentrations (Supplementary Figure S3; Castaigne et al.20).
Outgrowth of human CD34+ cells co-expressing REtr and activated c-KIT mutants in ex vivo cultures. (a) Scheme of the REtr and c-KIT(N822K) MSCV retroviral expression constructs used in this study. (b) Expression of REtr and c-KIT(N822K) in transduced CD34+ cell over time. (c) Proliferation kinetics of REtr, c-KIT(N822K) and REtr+c-KIT(N822K) and mock-transduced cells over time. (d) FACS blots of REtr and wild-type c-KIT or c-KIT(N822K) co-expressing CD34+ cells at days 6 and 33 after transduction. (e) Structure and annotation of c-KIT A-loop mutations (red) commonly found in t(8;21)-positive AMLs. The mutants tested in our study are marked in bold. Grey: A-loop structure of the c-KIT kinase domain; Y823: pseudosubstrate tyrosine for autoinhibitory function. (f) FACS blots showing the outgrowth of CD34+ cells expressing REtr and different c-KIT mutants at days 4 and 41 after co-transduction of human CD34+ cells.
REtr- and c-KIT(N822K)-expanded progenitor cells exhibit a phenotype similar to FAB AML-M2 myeloblastic cells
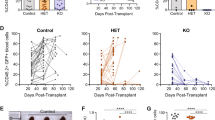
After ex vivo expansion, REtr- and REtr+c-KIT(N822K)-expressing cells were analyzed via FACS for cellular differentiation markers. In agreement with the published data, only a minor fraction of REtr-expressing cells remained CD34+, while most cells expressed myeloid differentiation markers.5, 12 In contrast, REtr+c-KIT(N822K) co-expressing cells were highly positive for CD11c and CD13 and resembled myeloblastic cells as originally described for AML FAB-M2 primary leukemia (Figure 2a). Based on the forward scatter/side scatter profiles, REtr+c-KIT(N822K) co-expressing cells were homogeneous and smaller in size with reduced levels of cell debris and differentiation. This enhanced differentiation block was also evident during the selection process by gating on the individual populations as well as for c-KIT(V825A), c-KIT(D816V) and c-KIT(A814S) co-expressing cells (Supplementary Figures S4a and b). Wright–Giemsa-stained cytospins further confirmed that REtr+c-KIT(N822K) co-expressing cells exhibited a more primitive cell morphology compared with REtr cells as demonstrated by the less segmented nuclei and a higher nucleus/cytoplasm ratio (Figure 2b). In REtr+c-KIT(N822K) co-expressing cells, a broad CD34+ cell population became evident even after 60 days of ex vivo culture, with 20% CD34+/CD38− cells, a cell surface marker combination characteristic of primitive progenitor cells (Figure 2b). Morphologically, the REtr+c-KIT(N822K)+ population showed a distinct shift towards undifferentiated blasts and myeloblasts, while REtr-expressing cells preferentially differentiated into macrophage-like cells (Figure 2c). To understand whether REtr+c-KIT(N822K)+ cells remained dependent on REtr expression, we generated double-positive cells expressing a floxed REtr-IRES-tdTomato construct (Figure 2d). After outgrowth, the double-positive cells were transduced with vectors expressing Cre recombinase. Excised REtr became evident by the loss of tdTomato fluorescence, which was co-expressed with REtr. Over time, the Cre-expressing, REtr-negative cells were lost owing to cell cycle arrest at G1 and concomitant myeloid differentiation (Figures 2e and f), thereby suggesting that REtr expression has an essential role in the maintenance of cell growth in REtr-c-KIT(N822K) long-term cultures.
Cell surface markers of REtr and REtr+c-KIT(N822K) cells after outgrowth. (a) Multi-parameter FACS blots illustrating the differentiation status of REtr- and REtr+c-KIT(N822K)-expressing cells after outgrowth. (b) Wright–Giemsa staining and CD34/CD38 FACS blots of REtr and REtr+c-KIT(N822K) cells. (c) Cell morphology distribution of REtr and REtr+c-KIT(N822K) cells (representative values). (d) Generation of progenitor cells carrying floxed REtr (REtrflox) and c-KIT(N822K) after outgrowth. (e) Growth kinetics of controls and Cre/YFP-expressing REtrflox+c-KIT(N822K) cells. (f) Differentiation profile and DRAQ5 cell cycle distribution following REtrflox excision at day 8 after Cre-T2A-YFP transduction.
C-KIT(N822K) enhances REtr-induced progenitor cell expansion and clonogenic growth
Compared with REtr counterparts, REtr+c-KIT(N822K)-expressing CD34+ cells displayed a growth advantage in cumulative cell growth in dense (1 × 10e6 cells/ml) ex vivo culture conditions (Figure 3a). Approximately 30 days after initial transduction, the growth advantage of double-transduced cells became visible. From day 60 onwards, REtr-expressing cells and double-transduced cells were assessed for clonogenic potential in methylcellulose-based colony-forming unit assays. Both populations were able to generate colonies until day 102 posttransduction; however, the quantity and size of colony-forming units derived from the REtr+c-KIT(N822K) co-expressing cells suggested an increased number of progenitor cells with colony-forming capacity in this population (Figures 3b and c). In mixed cultures containing both REtr and REtr+c-KIT(N822K) cells in the same well, REtr+c-KIT(N822K) co-expressing cells overgrew REtr cells within 10 days (Supplementary Figure S4c). In a serial dilution assay, REtr+c-KIT(N822K) co-expressing cells could be replated for several rounds, whereas REtr-expressing cells stopped growing after a few replating rounds, thereby strengthening the observation of an increased number of progenitor cells with increased self-renewal potential in the REtr+c-KIT(N822K) cell population (Figure 3d). A similar effect was observed after seeding a limited number of cells per well. REtr+c-KIT(N822K) co-expressing cells replenished the culture even from a small number of seeded cells (1000 cells/48-well plate), whereas REtr-expressing cells did not sustain growth (Figure 3e). REtr+c-KIT(N822K) cells remained cytokine dependent but were able to grow even in the presence of 1/10 of the initial cytokine concentration (20 ng/ml), albeit with slower kinetics (Supplementary Figure S4d). However, complete cytokine deprivation led to cell death in both REtr- and REtr+c-KIT(N822K)-expressing cells.
Proliferation kinetics of REtr and REtr+c-KIT(N822K) cells in long-term cultures and limited culture conditions. (a) Cumulative growth of REtr and REtr+c-KIT(N822K) cells under dense culture conditions (1 × 106 cells/ml). (b) colony-forming units at days 60, 74, 88 and 102 after long-term culture of cells shown in panel (a). (c) Colony morphology of secondary plated REtr- and REtr+c-KIT(N822K)-expressing cells after 74 days in culture. (d) Absolute cell numbers of cultures upon dilution to 1 × 105 cells/ml after every 4 days. (e) Cell numbers at day 7 after seeding 1000, 5000 and 10 000 REtr- and REtr+c-KIT(N822K)-expressing cells in 24-well plates.
C-KIT(N822K) does not propel the cell cycle but reduces the basal apoptosis rate in REtr-expressing cells
As activated tyrosine kinases are regarded as proliferation drivers in leukemia,21 we compared the cell cycle distribution of single versus double oncogene-expressing cells during the course of ex vivo expansion. Interestingly, no obvious difference was observed between the populations expressing REtr and REtr+c-KIT(N822K) at day 10 after transduction, a time point at which REtr- and REtr+c-KIT(N822K)-expressing cells co-exist (Figure 4a). However, after outgrowth, analysis of the individual REtr single or REtr+c-KIT(N822K) double-positive populations revealed a clear difference in cell cycle distribution with 26% of the REtr+c-KIT(N822K) cells in the S/G2M phase of the cell cycle, most likely reflecting a higher percentage of proliferation-competent progenitor cells in this population. In contrast, REtr cells were found to be primarily arrested in the G1 phase (Figure 4b). This increased proliferation rate was accompanied by a reduction in the basal apoptosis level as estimated by analyzing Annexin-V, subG1 levels and intracellular binding of a FITC-tagged ZVAD peptide in REtr+c-KIT(N822K) co-expressing cells (Figure 4c). In parallel, western blotting analysis revealed increased levels of the anti-apoptotic proteins AVEN, BCL2, BCLXL and MCL1 with reduced levels of activated caspase-3 and caspase-9 in REtr+c-KIT(N822K) cells, which also implies a decrease in the basal apoptosis rate. Furthermore, we found increased levels of AKT(S473) phosphorylation in c-KIT(N822K) co-expressing cells concomitant with increased phosphorylation of the adaptor protein GAB2 (Grb2-associated binder 2), which mediates the interaction between activated forms of c-KIT and phosphatidylinositol 3′-kinase (PI3K), thereby enhancing c-KIT signaling via the PI3K/AKT pathway (Figure 4d).
Cell cycle analysis and apoptosis in REtr and REtr+c-KIT(N822K) cells after outgrowth. (a) Cell cycle analysis of mixed cell populations at day 10 after transduction of CD34+ cells. Cells were stained with DRAQ5 and measured by FACS. (b) Cell cycle analysis (DRAQ5) after outgrowth of REtr- and REtr+c-KIT(N822K)-expressing cells. (c) Level of apoptosis in REtr- and REtr+c-KIT(N822K)-expressing cells by Annexin-V staining, subG1 (DRAQ5) and ZVAD-FITC staining. (d) Western blotting analysis of apoptosis-relevant proteins in REtr- and REtr+c-KIT(N822K)-expressing cells after outgrowth.
As activated caspases target REtr for degradation,22, 23, 24 we aimed to determine whether the downregulation of caspase activity in REtr+c-KIT(N822K) cells would stabilize the REtr protein. As expected, no REtr cleavage products were observed in REtr+c-KIT(N822K) cells (Supplementary Figures S5a and b). As the initiator caspase, caspase-9 was found activated exclusively in REtr cells but not in cultures co-expressing several activated c-KIT forms, we treated REtr cells with the pan-caspase inhibitor ZVAD-fmk and assessed REtr protein integrity via western blotting. In fact, we found reduced REtr cleavage after 3-day preincubation with ZVAD-fmk, indicating that caspases are indeed involved in REtr fragmentation (Supplementary Figure S5c).
REtr+c-KIT(N822K)-co-expressing progenitors are resistant towards imatinib but respond to PI3K inhibitors
As c-KIT(N822K) potentiates the transformation abilities of REtr, we aimed to inhibit oncogenic c-KIT in growing cultures by administration of imatinib. Surprisingly, REtr+c-KIT(N822K) cells did not respond to imatinib, even at high concentrations (Figure 5a). At the molecular level, PI3K signaling was slightly reduced at 5 μM imatinib, as assessed by measuring AKT phosphorylation but was completely abrogated in the presence of the PI3K inhibitor Ly294002 (Figures 5b and c). In the presence of Ly294002, induction of apoptosis became obvious in cells expressing both oncogenes as measured with Annexin-V staining (Figure 5d). Furthermore, Ly294002 acted synergistically with daunorubicin, to induce apoptosis in REtr+c-KIT(N822K)-expressing cells (Figure 5e). To understand the role that c-KIT(N822K) activity has in continuous progenitor cell growth, a floxed c-KIT(N822K) was introduced into CD34+ cells together with REtr. After outgrowth of double-positive cells, a Cre-T2A-YFP construct was retrovirally introduced. Four days thereafter, YFP-expressing cells were negative for c-KIT as analyzed via CD117 staining and PCR of genomic DNA, thereby indicating proper Cre-mediated excision of c-KIT(N822K) (Figures 5f and g). C-KIT(N822K) deletion resulted in a rapid loss of cells with increased Annexin-V positivity (Figures 5h and i), indicating that c-KIT(N822K) is indeed required for the maintenance of cell growth of REtr+c-KIT(N822K) progenitors.
Responsiveness of REtr+c-KIT(N822K) cells towards imatinib treatment and PI3K inhibition. (a) Growth of REtr+c-KIT(N822K) cells in the presence of 1 and 2 μM imatinib. (b) AKT(S473) phosphorylation in REtr and REtr+c-KIT(N822K) cells before or after treatment with 50 μM Ly294002. (c) AKT(S473) phosphorylation in REtr- and REtr+c-KIT(N822K)-expressing cells after treatment with 1, 2 and 5 μM imatinib. (d) Percentage of Annexin-V-positive REtr+c-KIT(N822K) cells after 3-day treatment with 10 and 50 μM Ly294002 (Ly29). (e) Percentage of Annexin-V-positive REtr+c-KIT(N822K) cells after 4-day treatment with 250 nM Daunorubicin, 50 μM Ly294002 and the combination of both. (f) Quantification of c-KIT(N822K) expression on transduced REtr+c-KIT(N822K) cells at days 3, 4 and 5 after transduction with Cre recombinase or control vectors. (g) PCR on genomic DNA before and after (day 4) Cre-mediated excision of c-KIT(N822K) in Cre-T2A-eYFP transduced REtr+c-KIT(N822K)-positive CD34+ cells. (h) Growth of REtr+c-KIT(N822K)flox CD34+ cells after transduction with a vector expressing Cre/FP or control vectors. (i) Quantification of apoptosis rates in the cell populations shown in panels (f) and (h). (***P<0.001; **0.001<P<0.01; *P<0.05).
C-KIT(N822K) accelerates DNA repair and counteracts REtr-induced DNA damage
Among the multiple genes transcriptionally repressed by RUNX1/ETO, genes involved in the DNA-damage response have an important role in RE-induced leukemogenesis via the induction of genomic instability. In fact, mutants of RUNX1 lacking the C-terminal amino acids trigger the formation of DNA double-strand breaks.25, 26, 27, 28, 29
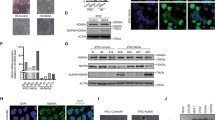
Therefore, we analyzed genomic instability in CD34+ cells expressing either REtr or REtr and c-KIT(N822K). As reported recently, we found increased numbers of γ-H2AX foci in REtr-expressing CD34+ cells (Figures 6a and b and Krejci et al.27). Surprisingly, we detected a decreased overall number of γ-H2AX foci in REtr+c-KIT(N822K)-positive cells, suggesting either an attenuated DNA-damage response or increased kinetics of double-strand break repair (Figures 6a and b). Genomic instability was not unique to REtr-expressing CD34+ cells, as similar effects were observed in the p53-positive human diploid IMR-90 cell line. REtr expression in these cells led to a significant increase in centrosomal aberrations as detected via γ-tubulin staining (Figure 6c). Co-expression of c-KIT(N822K) in REtr IMR-90 cells resulted in a significant and concentration-dependent reduction in abnormal centrosome numbers (Figure 6c). We next aimed to determine whether the increased apoptosis rate observed in REtr+c-KIT(N822K) cells upon LY294002 treatment was caused by increased numbers of DNA double-strand breaks. As expected, a significant increase in γ-H2AX foci numbers was observed in REtr+c-KIT(N822K) cells after LY294002 treatment, thereby linking the effect of the PI3K inhibitor to the increased DNA damage in c-KIT(N822K)-expressing cells. Of note, wild-type CD34+ cells were not affected by this treatment (Figure 6d). To investigate the molecular basis of c-KIT(N822K)-induced changes in the DNA-damage response, we assessed the kinetics of DNA double-strand break repair in REtr+c-KIT(N822K)-positive cells after γ-irradiation with 4 Gy. Compared with wild-type cells, we found an enhanced rate of DNA repair in CD34+ cells expressing REtr+c-KIT(N822K) (Figure 6e). To decipher the mechanisms of enhanced double-strand repair, we analyzed the rate of repair by non-homologues end joining (NHEJ) and homologous recombination (HRR) in REtr cells co-expressing c-KIT(N822K) using a cellular assay based on the reconstitution of a functional β-galactosidase cDNA.16 Although we did not observe differences in the kinetics of repair by NHEJ (data not shown), we found a significant reduction (P<0.0001) in the frequency of HRR in IMR-90 cells expressing REtr, which was entirely reversed upon expression of c-KIT(N822K) (Figure 6f). Compared with control and REtr-expressing cells, REtr+c-KIT(N822K) co-expressing IMR-90 cells showed increased nuclear Rad51 levels, thereby supporting the notion of enhancement of DNA-repair process via homologous recombination (Figure 6g). The observed increased levels of phospho-Rad51 and the accelerated rate of HRR as measured via Rad51 repair kinetics after gamma-irradiation further strengthened this hypothesis (Figures 6h–j). Furthermore, several important DNA-repair genes involved in homologous recombination that are known to be downregulated by REtr, such as OGG1, ATM and BRCA2,27 were re-expressed in REtr cells co-expressing c-KIT(N822K) (Figure 6k). The increased repair kinetics observed in REtr+c-KIT(N822K)-expressing cells should increase the sensitivity of these cells towards DNA-repair inhibitors. Indeed, treatment of REtr+c-KIT(N822K)- and REtr-expressing IMR-90 cells with the ATM inhibitors KU55933 and KU60019 and the PARP-inhibitor Olaparib resulted in increased levels of apoptotic cells, thereby implying that the DNA-repair machinery represents a novel therapeutic target for the treatment of t(8;21)+ leukemias with activated c-KIT mutations (Figure 6l). Apoptosis induction triggered by the ATM inhibitor KU55933 was also observed in CD34+ cells expressing REtr+cKIT(N822K), thus validating the use of DNA-repair inhibitors for the treatment of RE leukemias (Figure 6m).
DNA-damage response in REtr- and REtr+c-KIT(N822K)-expressing cells. (a) Immunofluorescence staining for γ-H2AX foci (red) in transduced CD34+ cells after outgrowth. Nuclei were counterstained with DAPI (blue). (b) Quantification of γ-H2AX foci in transduced CD34+ cells. Foci of 800–1000 individual nuclei were quantified on confocal picture stacks. Irradiated (1 Gy) CD34+ cells were used as control. (c) Quantification of centrosome numbers in IMR-90 cells transfected with the indicated plasmids. (d) Quantification of γ-H2AX foci in transduced CD34+ cells after 2 days of treatment with the PI3K inhibitor Ly294002 or DMSO. Foci of 150–200 individual nuclei were quantified on confocal picture stacks. (e) Quantification of γ-H2AX foci in transduced CD34+ cells at different time points after γ-irradiation (4 Gy). Foci of 200–400 individual nuclei were quantified on confocal picture stacks. (f) Reconstitution of β-galactosidase activity after homologous recombination in IMR-90 cells expressing REtr or REtr+c-KIT(N822K). Wild-type (WT) cells were used as controls. (g) Cellular localization of Rad51 in REtr- and REtr+c-KIT(N822K)-expressing cells or WT IMR-90 cells. Actin, tubulin and lamin were used to control for cellular localization and loading. (h) Analysis of Rad51 phosphorylation in REtr- and REtr+c-KIT(N822K)-expressing IMR-90 cells. (i) Quantification of Rad51 phosphorylation in REtr- and REtr+c-KIT(N822K)-expressing or WT IMR-90 cells. (j) Quantification of Rad51 foci in transduced IMR-90 cells at different time points after γ-irradiation (4 Gy). Foci of 200–400 individual nuclei were quantified on confocal picture stacks. (k) Relative expression of genes involved in HRR in cells expressing REtr or REtr+c-KIT(N822K). WT U937 cells were used as control. Values are normalized on TBP expression levels. (l) Apoptosis level in REtr- and REtr+c-KIT(N822K)-expressing or WT IMR-90 cells after daily treatment with DDR inhibitors. (m) Apoptosis levels in WT or REtr+c-KIT(N822K)-expressing CD34+ cells after daily treatment with Ku55933. Annexin-V staining was performed after 5 days of treatment. (n=3). (***P<0.001, **0.001<P<0.01, *P<0.05).
Transcriptome analysis of REtr+c-KIT(N822K)-expressing cells compared with REtr-only-expressing cells revealed a large number of deregulated genes. Besides signal transduction, genes involved in immunity, host defense and inflammation were selectively deregulated in c-KIT(N822K) expressing cells, suggesting additional potential targets for a specific therapeutic intervention in t(8;21) leukemia with activated c-KIT mutations (Supplementary Figure S6 and Supplementary Tables S1–S3).
Discussion
We show that activated c-KIT mutants strongly enhance REtr-mediated human CD34+ progenitor cell expansion in long-term ex vivo cultures. Within 30 days, REtr+c-KIT(N822K)-expressing cells predominated in culture and generated immature cell populations with increased self-renewal and survival potential. The expanded cell populations remain SCF- and cytokine-dependent, although lower concentrations were required for proliferation, thereby indicating the necessity of complementary signaling pathways for cell growth. Although overexpression of wild-type c-KIT has been found to be associated with t(8;21)+ AML6 and has the potential to drive oncogenic cooperativity with RE in a murine leukemia model,30 wild-type c-KIT did not cooperate with REtr-mediated progenitor expansion in our humanized CD34+ cell expansion model. Therefore, our cell culture system can be used to discriminate between silent and activating mutations of c-KIT. Expression of activated c-KIT alone, however, did not confer progenitor cell expansion properties comparable with RE. In this aspect, our model differs from two recent reports published during the course of our studies showing cooperativity between RE and activated c-KIT in mouse models.9, 10 In these studies, activated c-KIT alone triggered leukemia development, and when co-expressed with RE, it slightly accelerated leukemia onset. Although RE- and c-KIT(N822K)-expressing human primary CD34+ cells accurately resembled the AML FAB-M2 myeloblastic phenotype, intravenous and intrafemoral injection of these cells into sublethally irradiated NSG mice did not lead to leukemia development (Supplementary Figure S7), thereby arguing either for the necessity of additional hits or the absence of specialized bone marrow niches for full blast cell transformation of human hematopoietic progenitors in NSG mice.
As imatinib has been shown to bind to the catalytic center of c-KIT, we investigated its effect on REtr- and c-KIT(N822K)-co-expressing human progenitor cells. Previous work has demonstrated that imatinib can revert the effects of activated c-KIT on factor-independent growth of IL-3-dependent murine hematopoietic cell lines, which become IL-3-independent upon expression of activated c-KIT in the absence of SCF.9, 31 In contrast, imatinib applied as a single agent has only moderate effects in human primary t(8;21)+ AML cells harboring the cKIT mutation N822K.7 Our experiments also suggest that imatinib has a limited inhibitory effect on the activated forms of c-KIT, as REtr+c-KIT(N822K) cells do not respond to imatinib treatment. The permanent presence of SCF in our cell culture media most likely renders the kinase in a constitutive activated conformation, which prevents binding of the inhibitor, as imatinib preferentially binds to the catalytic site of the enzyme in its inactive conformation.32, 33 Moreover, we found activated GAB2 in REtr+c-KIT(N822K) cells, which has recently been shown to confer resistance to multiple kinase inhibitors, including imatinib, nilotinib and dasatinib, in BCR/ABL-positive cells.34 In contrast, inhibition of PI3K signaling was highly effective in REtr+c-KIT(N822K) cells and showed synergistic effects when administrated together with daunorubicin. Indeed, the PI3K/AKT pathway is a major signaling node activated by several receptor tyrosine kinases, including c-KIT, thus providing a rational for a therapeutic intervention.35, 36, 37, 38, 39 However, the PI3K inhibitor used in our study, Ly294002, has been shown to bind to several other therapeutic targets, and therefore we cannot attribute the effects of Ly294002 to the exclusive inhibition of PI3K activity.40, 41
Several recent reports have demonstrated that RE has a direct role in the induction of DNA damage linking RE to genomic instability. In fact, RE alone is sufficient to induce increased numbers of γ-H2AX foci in human CD34+ long-term cultures together with an increase in the basal apoptosis rate.26, 27 However, induction of apoptosis would counteract the pro-leukemic properties of RE. Therefore we anticipated that activated c-KIT could have a role in dampening the DNA-damage response, thereby lowering the rates of cell death. We found that activated c-KIT was able to restore the expression of RE-induced downregulation of ATM, OGG1 and BRCA2, which are important members of the DNA-repair machinery.26 Moreover, the c-KIT-mediated effects on the DNA-damage response were reverted by the PI3K inhibitor Ly294002. PI3K and AKT are critical for DNA repair,42 and PI3K has been shown to upregulate BRCA2 expression thereby stabilizing homologous recombination.43 OGG1, which acts as an 8-oxoG glycosylase and endonuclease, was also described to be induced by the PI3K signaling pathway.44 Furthermore, we found that the expression of ATM was restored in REtr+c-KIT(N822K)-expressing cells. ATM, in turn, is involved in the activation of the DNA-damage response and contributes to the recruitment of the DNA-repair machinery to DNA strand breaks resulting in HRR or NHEJ.45 Increased DNA-repair rates in REtr+c-KIT(N822K) co-expressing cells were accompanied by reduced apoptosis levels and caspase activity. The latter may explain the reduced REtr cleavage rates observed in c-KIT(N822K) co-expressing cells, as REtr has been shown to be a target of caspases.23 Increased REtr levels may then account for the more immature cell phenotype as observed in primary t(8;21)+ leukemia cells with high RE expression levels.6 Additionally, caspases also trigger cellular differentiation and are involved in RAD51 degradation.46, 47, 48
Our results show that C-KIT(N822K) triggers a series of pleiotropic effects in CD34+ cells expressing REtr, including reduced apoptosis, enhancement of chemoresistance, reactivation of DNA-repair genes and accelerated repair of DNA damage. These effects act in synergy to generate a preleukemic stage, which may predispose primary CD34+ cells to overt transformation. The continuous expression of both oncogenes was necessary for the observed effects, as deletion of either REtr or c-KIT(N822K) led to a significant increase in differentiation and apoptosis rates. This observation provides a rationale for a direct targeting of RE and c-KIT for therapeutic purposes. Indeed, several approaches targeting AEtr or c-KIT oncogenic activity and processing have been reported.14, 24, 49, 50, 51, 52
In conclusion, we propose that activated c-KIT confers oncogenic cooperativity for RE leading to the selection of myeloblastic cells with increased progenitor cell characteristics, an enhanced block in cellular differentiation and resistance to stress stimuli and DNA-damage signals. The later observation may also explain the higher relapse rates in c-KIT positive t(8;21)+ AML patients after conventional chemotherapy.6, 53 In addition to approaches inhibiting the oncogenic activity of RE and c-KIT, targeting the DNA-damage repair machinery might represent a new promising path to interfere with t(8;21)-positive leukemia with activated c-KIT.
References
Yan M, Burel SA, Peterson LF, Kanbe E, Iwasaki H, Boyapati A et al. Deletion of an AML1-ETO C-terminal NcoR/SMRT-interacting region strongly induces leukemia development. Proc Natl Acad Sci USA 2004; 101: 17186–17191.
Wildonger J, Mann RS . The t(8;21) translocation converts AML1 into a constitutive transcriptional repressor. Development 2005; 132: 2263–2272.
Ptasinska A, Assi SA, Mannari D, James SR, Williamson D, Dunne J et al. Depletion of RUNX1/ETO in t(8;21) AML cells leads to genome-wide changes in chromatin structure and transcription factor binding. Leukemia 2012; 26: 1829–1841.
Song J, Mercer D, Hu X, Liu H, Li MM . Common leukemia- and lymphoma-associated genetic aberrations in healthy individuals. J Mol Diagn 2011; 13: 213–219.
Mulloy JC, Cammenga J, Berguido FJ, Wu K, Zhou P, Comenzo RL et al. Maintaining the self-renewal and differentiation potential of human CD34+ hematopoietic cells using a single genetic element. Blood 2003; 102: 4369–4376.
Jiao B, Wu CF, Liang Y, Chen HM, Xiong SM, Chen B et al. AML1-ETO9a is correlated with C-KIT overexpression/mutations and indicates poor disease outcome in t(8;21) acute myeloid leukemia-M2. Leukemia 2009; 23: 1598–1604.
Wang YY, Zhou GB, Yin T, Chen B, Shi JY, Liang WX et al. AML1-ETO and C-KIT mutation/overexpression in t(8;21) leukemia: implication in stepwise leukemogenesis and response to Gleevec. Proc Natl Acad Sci USA 2005; 102: 1104–1109.
Wakita S, Yamaguchi H, Miyake K, Mitamura Y, Kosaka F, Dan K et al. Importance of c-kit mutation detection method sensitivity in prognostic analyses of t(8;21)(q22;q22) acute myeloid leukemia. Leukemia 2011; 25: 1423–1432.
Wang YY, Zhao LJ, Wu CF, Liu P, Shi L, Liang Y et al. C-KIT mutation cooperates with full-length AML1-ETO to induce acute myeloid leukemia in mice. Proc Natl Acad Sci USA 2011; 108: 2450–2455.
Nick HJ, Kim HG, Chang CW, Harris KW, Reddy V, Klug CA . Distinct classes of c-Kit-activating mutations differ in their ability to promote RUNX1-ETO-associated acute myeloid leukemia. Blood 2012; 119: 1522–1531.
Lennartsson J, Ronnstrand L . Stem cell factor receptor/c-Kit: from basic science to clinical implications. Physiol Rev 2012; 92: 1619–1649.
Mulloy JC, Cammenga J, MacKenzie KL, Berguido FJ, Moore MA, Nimer SD . The AML1-ETO fusion protein promotes the expansion of human hematopoietic stem cells. Blood 2002; 99: 15–23.
Chou FS, Wunderlich M, Griesinger A, Mulloy JC . N-Ras(G12D) induces features of stepwise transformation in preleukemic human umbilical cord blood cultures expressing the AML1-ETO fusion gene. Blood 2011; 117: 2237–2240.
Wichmann C, Becker Y, Chen-Wichmann L, Vogel V, Vojtkova A, Herglotz J et al. Dimer-tetramer transition controls RUNX1/ETO leukemogenic activity. Blood 2010; 116: 603–613.
Stein S, Ott MG, Schultze-Strasser S, Jauch A, Burwinkel B, Kinner A et al. Genomic instability and myelodysplasia with monosomy 7 consequent to EVI1 activation after gene therapy for chronic granulomatous disease. Nat Med 2010; 16: 198–204.
Epinat JC, Arnould S, Chames P, Rochaix P, Desfontaines D, Puzin C et al. A novel engineered meganuclease induces homologous recombination in yeast and mammalian cells. Nucleic Acids Res 2003; 31: 2952–2962.
Team RDC. A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria, 2005. http://www.R-project.org.
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 2004; 5: R80.
Yan M, Kanbe E, Peterson LF, Boyapati A, Miao Y, Wang Y et al. A previously unidentified alternatively spliced isoform of t(8;21) transcript promotes leukemogenesis. Nat Med 2006; 12: 945–949.
Castaigne S, Daniel MT, Tilly H, Herait P, Degos L . Does treatment with ARA-C in low dosage cause differentiation of leukemic cells? Blood 1983; 62: 85–86.
Gilliland DG, Griffin JD . Role of FLT3 in leukemia. Curr Opin Hematol 2002; 9: 274–281.
Wang L, Zhao WL, Yan JS, Liu P, Sun HP, Zhou GB et al. Eriocalyxin B induces apoptosis of t(8;21) leukemia cells through NF-kappaB and MAPK signaling pathways and triggers degradation of AML1-ETO oncoprotein in a caspase-3-dependent manner. Cell Death Differ 2007; 14: 306–317.
Lu Y, Peng ZG, Yuan TT, Yin QQ, Xia L, Chen GQ . Multi-sites cleavage of leukemogenic AML1-ETO fusion protein by caspase-3 and its contribution to increased apoptotic sensitivity. Leukemia 2008; 22: 378–386.
Zhen T, Wu CF, Liu P, Wu HY, Zhou GB, Lu Y et al. Targeting of AML1-ETO in t(8;21) leukemia by oridonin generates a tumor suppressor-like protein. Sci Transl Med 2012; 4: 127ra138.
Burel SA, Harakawa N, Zhou L, Pabst T, Tenen DG, Zhang DE . Dichotomy of AML1-ETO functions: growth arrest versus block of differentiation. Mol Cell Biol 2001; 21: 5577–5590.
Alcalay M, Meani N, Gelmetti V, Fantozzi A, Fagioli M, Orleth A et al. Acute myeloid leukemia fusion proteins deregulate genes involved in stem cell maintenance and DNA repair. J Clin Invest 2003; 112: 1751–1761.
Krejci O, Wunderlich M, Geiger H, Chou FS, Schleimer D, Jansen M et al. p53 signaling in response to increased DNA damage sensitizes AML1-ETO cells to stress-induced death. Blood 2008; 111: 2190–2199.
Lu Y, Xu YB, Yuan TT, Song MG, Lubbert M, Fliegauf M et al. Inducible expression of AML1-ETO fusion protein endows leukemic cells with susceptibility to extrinsic and intrinsic apoptosis. Leukemia 2006; 20: 987–993.
Satoh Y, Matsumura I, Tanaka H, Harada H, Harada Y, Matsui K et al. C-terminal mutation of RUNX1 attenuates the DNA-damage repair response in hematopoietic stem cells. Leukemia 2012; 26: 303–311.
Zheng X, Oancea C, Henschler R, Ruthardt M . Cooperation between constitutively activated c-Kit signaling and leukemogenic transcription factors in the determination of the leukemic phenotype in murine hematopoietic stem cells. Int J Oncol 2009; 34: 1521–1531.
Cammenga J, Horn S, Bergholz U, Sommer G, Besmer P, Fiedler W et al. Extracellular KIT receptor mutants, commonly found in core binding factor AML, are constitutively active and respond to imatinib mesylate. Blood 2005; 106: 3958–3961.
Foster R, Griffith R, Ferrao P, Ashman L . Molecular basis of the constitutive activity and STI571 resistance of Asp816Val mutant KIT receptor tyrosine kinase. J Mol Graph Model 2004; 23: 139–152.
Schindler T, Bornmann W, Pellicena P, Miller WT, Clarkson B, Kuriyan J . Structural mechanism for STI-571 inhibition of abelson tyrosine kinase. Science 2000; 289: 1938–1942.
Wohrle FU, Halbach S, Aumann K, Schwemmers S, Braun S, Auberger P et al. Gab2 signaling in chronic myeloid leukemia cells confers resistance to multiple Bcr-Abl inhibitors. Leukemia 2013; 27: 118–129.
Hashimoto K, Matsumura I, Tsujimura T, Kim DK, Ogihara H, Ikeda H et al. Necessity of tyrosine 719 and phosphatidylinositol 3'-kinase-mediated signal pathway in constitutive activation and oncogenic potential of c-kit receptor tyrosine kinase with the Asp814Val mutation. Blood 2003; 101: 1094–1102.
Lennartsson J, Ronnstrand L . The stem cell factor receptor/c-Kit as a drug target in cancer. Curr Cancer Drug Targets 2006; 6: 65–75.
Bertacchini J, Guida M, Accordi B, Mediani L, Martelli AM, Barozzi P et al. Feedbacks and adaptive capabilities of the PI3K/Akt/mTOR axis in acute myeloid leukemia revealed by pathway selective inhibition and phosphoproteome analysis. Leukemia 2014; e-pub ahead of print 4 April 2014; doi:10.1038/leu.2014.123.
Reikvam H, Tamburini J, Skrede S, Holdhus R, Poulain L, Ersvaer E et al. Antileukaemic effect of PI3K-mTOR inhibitors in acute myeloid leukaemia-gene expression profiles reveal CDC25B expression as determinate of pharmacological effect. Br J Haematol 2014; 164: 200–211.
Thomas D, Powell JA, Vergez F, Segal DH, Nguyen NY, Baker A et al. Targeting acute myeloid leukemia by dual inhibition of PI3K signaling and Cdk9-mediated Mcl-1 transcription. Blood 2013; 122: 738–748.
Bain J, Plater L, Elliott M, Shpiro N, Hastie CJ, McLauchlan H et al. The selectivity of protein kinase inhibitors: a further update. Biochem J 2007; 408: 297–315.
Gharbi SI, Zvelebil MJ, Shuttleworth SJ, Hancox T, Saghir N, Timms JF et al. Exploring the specificity of the PI3K family inhibitor LY294002. Biochem J 2007; 404: 15–21.
Kumar A, Fernandez-Capetillo O, Carrera AC . Nuclear phosphoinositide 3-kinase beta controls double-strand break DNA repair. Proc Natl Acad Sci USA 2010; 107: 7491–7496.
Ibrahim YH, Garcia-Garcia C, Serra V, He L, Torres-Lockhart K, Prat A et al. PI3K inhibition impairs BRCA1/2 expression and sensitizes BRCA-proficient triple-negative breast cancer to PARP inhibition. Cancer Discov 2012; 2: 1036–1047.
Kim KC, Lee IK, Kang KA, Cha JW, Cho SJ, Na SY et al. 7,8-Dihydroxyflavone suppresses oxidative stress-induced base modification in DNA via induction of the repair enzyme 8-oxoguanine DNA glycosylase-1. Biomed Res Int 2013; 2013: 863720.
Helleday T . Homologous recombination in cancer development, treatment and development of drug resistance. Carcinogenesis 2010; 31: 955–960.
Fujita J, Crane AM, Souza MK, Dejosez M, Kyba M, Flavell RA et al. Caspase activity mediates the differentiation of embryonic stem cells. Cell Stem Cell 2008; 2: 595–601.
Leong SM, Tan BX, Bte Ahmad B, Yan T, Chee LY, Ang ST et al. Mutant nucleophosmin deregulates cell death and myeloid differentiation through excessive caspase-6 and -8 inhibition. Blood 2010; 116: 3286–3296.
Huang Y, Nakada S, Ishiko T, Utsugisawa T, Datta R, Kharbanda S et al. Role for caspase-mediated cleavage of Rad51 in induction of apoptosis by DNA damage. Mol Cell Biol 1999; 19: 2986–2997.
Bartel Y, Grez M, Wichmann C . Interference with RUNX1/ETO leukemogenic function by cell-penetrating peptides targeting the NHR2 oligomerization domain. Biomed Res Int 2013; 2013: 297692.
Metz A, Schanda J, Grez M, Wichmann C, Gohlke H . From determinants of RUNX1/ETO tetramerization to small-molecule protein-protein interaction inhibitors targeting acute myeloid leukemia. J Chem Inf Model 2013; 53: 2197–2202.
Wichmann C, Chen L, Heinrich M, Baus D, Pfitzner E, Zornig M et al. Targeting the oligomerization domain of ETO interferes with RUNX1/ETO oncogenic activity in t(8;21)-positive leukemic cells. Cancer Res 2007; 67: 2280–2289.
Fang HT, Zhang B, Pan XF, Gao L, Zhen T, Zhao HX et al. Bortezomib interferes with C-KIT processing and transforms the t(8;21)-generated fusion proteins into tumor-suppressing fragments in leukemia cells. Proc Natl Acad Sci USA 2012; 109: 2521–2526.
Paschka P, Marcucci G, Ruppert AS, Mrozek K, Chen H, Kittles RA et al. Adverse prognostic significance of KIT mutations in adult acute myeloid leukemia with inv(16) and t(8;21): a Cancer and Leukemia Group B Study. J Clin Oncol 2006; 24: 3904–3911.
Acknowledgements
We thank Sandra Moore for critical comments on the manuscript and Dr Frank Schnüttgen (Department of Hematology, University Hospital Frankfurt/Main, Germany) for providing the plasmids pCMV-HPRT, pCMV-mHPRTTAL1 and pCMV-mHPRTTAL2. We are supported by research grants from the NGFN Cancer Network (grant 01GS0450, TP-10 to MG), the LOEWE Center for Cell and Gene Therapy Frankfurt funded by the Hessisches Ministerium für Wissenschaft und Kunst (HMWK; funding reference number: III L 4–518/17.004 (2010) (startup grant to CW)), the LOEWE OSF (TP-C3) and the Jose’ Carreras Leukemia Foundation (DJCLS R 12/28, to CW). The Georg-Speyer-Haus is funded jointly by the German Federal Ministry of Health (BMG) and the Ministry of Higher Education, Research and the Arts of the State of Hessen (HMWK).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Wichmann, C., Quagliano-Lo Coco, I., Yildiz, Ö. et al. Activating c-KIT mutations confer oncogenic cooperativity and rescue RUNX1/ETO-induced DNA damage and apoptosis in human primary CD34+ hematopoietic progenitors. Leukemia 29, 279–289 (2015). https://doi.org/10.1038/leu.2014.179
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2014.179
This article is cited by
-
Combination of eriocalyxin B and homoharringtonine exerts synergistic anti-tumor effects against t(8;21) AML
Acta Pharmacologica Sinica (2024)
-
AML1/ETO and its function as a regulator of gene transcription via epigenetic mechanisms
Oncogene (2021)
-
ZBTB7A prevents RUNX1-RUNX1T1-dependent clonal expansion of human hematopoietic stem and progenitor cells
Oncogene (2020)
-
Compatibility of RUNX1/ETO fusion protein modules driving CD34+ human progenitor cell expansion
Oncogene (2019)
-
UBASH3B/Sts-1-CBL axis regulates myeloid proliferation in human preleukemia induced by AML1-ETO
Leukemia (2016)