Abstract
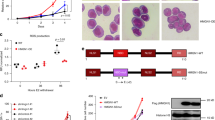
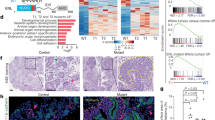
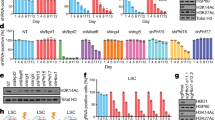
Histone H3 lysine 4 methylation (H3K4me) has been proposed as a critical component in regulating gene expression, epigenetic states, and cellular identities1. The biological meaning of H3K4me is interpreted by conserved modules including plant homeodomain (PHD) fingers that recognize varied H3K4me states1,2. The dysregulation of PHD fingers has been implicated in several human diseases, including cancers and immune or neurological disorders3. Here we report that fusing an H3K4-trimethylation (H3K4me3)-binding PHD finger, such as the carboxy-terminal PHD finger of PHF23 or JARID1A (also known as KDM5A or RBBP2), to a common fusion partner nucleoporin-98 (NUP98) as identified in human leukaemias4,5, generated potent oncoproteins that arrested haematopoietic differentiation and induced acute myeloid leukaemia in murine models. In these processes, a PHD finger that specifically recognizes H3K4me3/2 marks was essential for leukaemogenesis. Mutations in PHD fingers that abrogated H3K4me3 binding also abolished leukaemic transformation. NUP98–PHD fusion prevented the differentiation-associated removal of H3K4me3 at many loci encoding lineage-specific transcription factors (Hox(s), Gata3, Meis1, Eya1 and Pbx1), and enforced their active gene transcription in murine haematopoietic stem/progenitor cells. Mechanistically, NUP98–PHD fusions act as ‘chromatin boundary factors’, dominating over polycomb-mediated gene silencing to ‘lock’ developmentally critical loci into an active chromatin state (H3K4me3 with induced histone acetylation), a state that defined leukaemia stem cells. Collectively, our studies represent, to our knowledge, the first report that deregulation of the PHD finger, an ‘effector’ of specific histone modification, perturbs the epigenetic dynamics on developmentally critical loci, catastrophizes cellular fate decision-making, and even causes oncogenesis during mammalian development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
The structural coordinates of JARID1A-PHD3 in the H3-bound or free state have been submitted to the Protein Data Bank under accessions 3GL6, 2KGG and 2KGI, and the chemical shift assignment of NMR structures to BioMagResBank under accessions 16209 and 16210.
Change history
11 June 2009
In Fig. 1e, the x-axis label was changed from 'Cd115 (c-Kit)' to 'Cd117 (c-Kit)'.
References
Ruthenburg, A. J., Allis, C. D. & Wysocka, J. Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol. Cell 25, 15–30 (2007)
Taverna, S. D., Li, H., Ruthenburg, A. J., Allis, C. D. & Patel, D. J. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nature Struct. Mol. Biol. 14, 1025–1040 (2007)
Baker, L. A., Allis, C. D. & Wang, G. G. PHD fingers in human diseases: Disorders arising from misinterpreting epigenetic marks. Mutat. Res. 647, 3–12 (2008)
van Zutven, L. J. et al. Identification of NUP98 abnormalities in acute leukemia: JARID1A (12p13) as a new partner gene. Genes Chromosom. Cancer 45, 437–446 (2006)
Reader, J. C., Meekins, J. S., Gojo, I. & Ning, Y. A novel NUP98–PHF23 fusion resulting from a cryptic translocation t(11;17)(p15;p13) in acute myeloid leukemia. Leukemia 21, 842–844 (2007)
Wysocka, J. et al. A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442, 86–90 (2006)
Li, H. et al. Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nature 442, 91–95 (2006)
Pena, P. V. et al. Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 442, 100–103 (2006)
Shi, X. et al. ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442, 96–99 (2006)
Vermeulen, M. et al. Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell 131, 58–69 (2007)
Lan, F. et al. Recognition of unmethylated histone H3 lysine 4 links BHC80 to LSD1-mediated gene repression. Nature 448, 718–722 (2007)
Ooi, S. K. et al. DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature 448, 714–717 (2007)
Matthews, A. G. et al. RAG2 PHD finger couples histone H3 lysine 4 trimethylation with V(D)J recombination. Nature 450, 1106–1110 (2007)
Gong, W., Suzuki, K., Russell, M. & Riabowol, K. Function of the ING family of PHD proteins in cancer. Int. J. Biochem. Cell Biol. 37, 1054–1065 (2005)
Kasper, L. H. et al. CREB binding protein interacts with nucleoporin-specific FG repeats that activate transcription and mediate NUP98-HOXA9 oncogenicity. Mol. Cell. Biol. 19, 764–776 (1999)
Wang, G. G., Cai, L., Pasillas, M. P. & Kamps, M. P. NUP98–NSD1 links H3K36 methylation to Hox-A gene activation and leukaemogenesis. Nature Cell Biol. 9, 804–812 (2007)
Moore, M. A. et al. NUP98 dysregulation in myeloid leukemogenesis. Ann. NY Acad. Sci. 1106, 114–142 (2007)
Wang, G. G., Pasillas, M. P. & Kamps, M. P. Meis1 programs transcription of FLT3 and cancer stem cell character, using a mechanism that requires interaction with Pbx and a novel function of the Meis1 C-terminus. Blood 106, 254–264 (2005)
Taverna, S. D. et al. Yng1 PHD finger binding to H3 trimethylated at K4 promotes NuA3 HAT activity at K14 of H3 and transcription at a subset of targeted ORFs. Mol. Cell 24, 785–796 (2006)
Lee, T. I. et al. Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 125, 301–313 (2006)
Boyer, L. A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, 349–353 (2006)
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007)
Kroon, E. et al. Hoxa9 transforms primary bone marrow cells through specific collaboration with Meis1a but not Pbx1b . EMBO J. 17, 3714–3725 (1998)
Pineault, N., Helgason, C. D., Lawrence, H. J. & Humphries, R. K. Differential expression of Hox, Meis1, and Pbx1 genes in primitive cells throughout murine hematopoietic ontogeny. Exp. Hematol. 30, 49–57 (2002)
Martin, D. G. et al. The Yng1p plant homeodomain finger is a methyl-histone binding module that recognizes lysine 4-methylated histone H3. Mol. Cell. Biol. 26, 7871–7879 (2006)
Doyon, Y. et al. ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol. Cell 21, 51–64 (2006)
Wysocka, J. et al. WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell 121, 859–872 (2005)
Lee, M. G. et al. Demethylation of H3K27 regulates polycomb recruitment and H2A ubiquitination. Science 318, 447–450 (2007)
Cloos, P. A., Christensen, J., Agger, K. & Helin, K. Erasing the methyl mark: histone demethylases at the center of cellular differentiation and disease. Genes Dev. 22, 1115–1140 (2008)
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000)
Wang, G. G. et al. Quantitative production of macrophages or neutrophils ex vivo using conditional Hoxb8. Nature Methods 3, 287–293 (2006)
Sawka-Verhelle, D. et al. PE-1/METS, an antiproliferative Ets repressor factor, is induced by CREB-1/CREM-1 during macrophage differentiation. J. Biol. Chem. 279, 17772–17784 (2004)
Calvo, K. R., Sykes, D. B., Pasillas, M. & Kamps, M. P. Hoxa9 immortalizes a granulocyte-macrophage colony-stimulating factor-dependent promyelocyte capable of biphenotypic differentiation to neutrophils or macrophages, independent of enforced meis expression. Mol. Cell. Biol. 20, 3274–3285 (2000)
Okada, Y. et al. hDOT1L links histone methylation to leukemogenesis. Cell 121, 167–178 (2005)
Wallace, W., Schaefer, L. H. & Swedlow, J. R. A workingperson’s guide to deconvolution in light microscopy. Biotechniques 31, 1076–1078 (2001)
Bolte, S. & Cordelieres, F. P. A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 224, 213–232 (2006)
Fischle, W. et al. Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438, 1116–1122 (2005)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
McCoy, A. J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D 63, 32–41 (2007)
Brunger, A. T. Version 1.2 of the crystallography and NMR system. Nature Protocols 2, 2728–2733 (2007)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Zwahlen, C. et al. Methods for measurement of intermolecular NOEs by multinuclear NMR spectroscopy: application to a bacteriophage λ N-peptide/boxB RNA complex. J. Am. Chem. Soc. 119, 6711–6721 (1997)
Ottiger, M., Delaglio, F. & Bax, A. Measurement of J and dipolar couplings from simplified two-dimensional NMR spectra. J. Magn. Reson. 131, 373–378 (1998)
Farrow, N. A. et al. Backbone dynamics of a free and phosphopeptide-complexed Src homology 2 domain studied by 15N NMR relaxation. Biochemistry 33, 5984–6003 (1994)
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995)
Guntert, P., Mumenthaler, C. & Wuthrich, K. Torsion angle dynamics for NMR structure calculation with the new program DYANA. J. Mol. Biol. 273, 283–298 (1997)
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999)
Schwieters, C. D., Kuszewski, J. J., Tjandra, N. & Clore, G. M. The Xplor-NIH NMR molecular structure determination package. J. Magn. Reson. 160, 65–73 (2003)
Linge, J. P., Williams, M. A., Spronk, C. A., Bonvin, A. M. & Nilges, M. Refinement of protein structures in explicit solvent. Proteins 50, 496–506 (2003)
Laskowski, R. A., Rullmannn, J. A., MacArthur, M. W., Kaptein, R. & Thornton, J. M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996)
Lee, L. K., Rance, M., Chazin, W. J. & Palmer, A. G. Rotational diffusion anisotropy of proteins from simultaneous analysis of 15N and 13C alpha nuclear spin relaxation. J. Biomol. NMR 9, 287–298 (1997)
Acknowledgements
We thank L. Baker, P. Chi, A. Ruthenberg and A. Xiao for critical reading and help in manuscript preparation, C. Hughes for antibodies, and other Allis laboratory members for their advice. We are extremely grateful to M. Kamps and S. Rafii for shared plasmids and expert advice. Thanks to E. Coutavas, W. Chen, E. Ezhkova and the R. Roeder laboratory for help on cell and animal work. G.G.W. is supported by a Leukemia & Lymphoma Society Fellow award and a Choh-Hao Li Memorial Fund Scholar award; H.L.D. is supported by a predoctoral Boehringer Ingelheim Foundation fellowship; This research was supported by the National Institutes of Health (NIH) grant and funds of Rockefeller University to C.D.A., funds of Abby Rockefeller Mauze Trust and the Dewitt Wallace and Maloris Foundations to D.J.P., US Department of Defense CDMRP grant to J.-L.L., and a joint Starr Foundation Cancer Consortium grant.
Author Contributions G.G.W. and C.D.A. designed the study. G.G.W. performed most of the cellular and molecular experiments, and wrote the paper; J.S. performed protein preparation, NMR structure determination and crystallization; Z.W. performed crystallographic analyses; H.L.D. and G.G.W. did the immunostaining; F.C. participated in plasmid/protein preparation; H.L. performed isothermal titration calorimetry measurement; G.G.W. and J.-L.L. performed animal studies; D.J.P. and C.D.A. supervised the structural and functional aspects of the project, respectively, and helped with manuscript preparation.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-12 with Legends, Supplementary Tables 1-5 and a Supplementary Reference. (PDF 6325 kb)
Rights and permissions
About this article
Cite this article
Wang, G., Song, J., Wang, Z. et al. Haematopoietic malignancies caused by dysregulation of a chromatin-binding PHD finger. Nature 459, 847–851 (2009). https://doi.org/10.1038/nature08036
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08036
This article is cited by
-
The emerging roles of histone demethylases in cancers
Cancer and Metastasis Reviews (2024)
-
Proximity labeling reveals a new in vivo network of interactors for the histone demethylase KDM5
Epigenetics & Chromatin (2023)
-
The pediatric leukemia oncoprotein NUP98-KDM5A induces genomic instability that may facilitate malignant transformation
Cell Death & Disease (2023)
-
Regulation, functions and transmission of bivalent chromatin during mammalian development
Nature Reviews Molecular Cell Biology (2023)
-
Epigenetic regulation in hematopoiesis and its implications in the targeted therapy of hematologic malignancies
Signal Transduction and Targeted Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.