Abstract
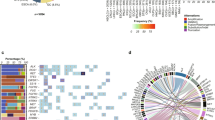
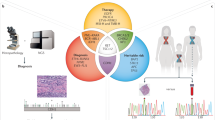
As more clinically relevant cancer genes are identified, comprehensive diagnostic approaches are needed to match patients to therapies, raising the challenge of optimization and analytical validation of assays that interrogate millions of bases of cancer genomes altered by multiple mechanisms. Here we describe a test based on massively parallel DNA sequencing to characterize base substitutions, short insertions and deletions (indels), copy number alterations and selected fusions across 287 cancer-related genes from routine formalin-fixed and paraffin-embedded (FFPE) clinical specimens. We implemented a practical validation strategy with reference samples of pooled cell lines that model key determinants of accuracy, including mutant allele frequency, indel length and amplitude of copy change. Test sensitivity achieved was 95–99% across alteration types, with high specificity (positive predictive value >99%). We confirmed accuracy using 249 FFPE cancer specimens characterized by established assays. Application of the test to 2,221 clinical cases revealed clinically actionable alterations in 76% of tumors, three times the number of actionable alterations detected by current diagnostic tests.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Stegmeier, F., Warmuth, M., Sellers, W.R. & Dorsch, M. Targeted cancer therapies in the twenty-first century: lessons from imatinib. Clin. Pharmacol. Ther. 87, 543–552 (2010).
Chin, L. & Gray, J.W. Translating insights from the cancer genome into clinical practice. Nature 452, 553–563 (2008).
Stratton, M.R., Campbell, P.J. & Futreal, P.A. The cancer genome. Nature 458, 719–724 (2009).
Hudson, T.J. et al. International network of cancer genome projects. Nature 464, 993–998 (2010).
Mardis, E.R. Genome sequencing and cancer. Curr. Opin. Genet. Dev. 22, 245–250 (2012).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Garnett, M.J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575 (2012).
Pao, W. New approaches to targeted therapy in lung cancer. Proc. Am. Thorac. Soc. 9, 72–73 (2012).
Thomas, R.K. et al. High-throughput oncogene mutation profiling in human cancer. Nat. Genet. 39, 347–351 (2007).
MacConaill, L.E. et al. Profiling critical cancer gene mutations in clinical tumor samples. PLoS ONE 4, e7887 (2009).
Dias-Santagata, D. et al. Rapid targeted mutational analysis of human tumours: a clinical platform to guide personalized cancer medicine. EMBO Mol. Med. 2, 146–158 (2010).
Ross, J.S. Update on HER2 testing for breast and upper gastrointestinal tract cancers. Biomark. Med. 5, 307–318 (2011).
McCourt, C.M., Boyle, D., James, J. & Salto-Tellez, M. Immunohistochemistry in the era of personalised medicine. J. Clin. Pathol. 66, 58–61 (2013).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Ding, L. et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481, 506–510 (2012).
Stephens, P.J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012).
Hammerman, P.S. et al. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525 (2012).
Roychowdhury, S. et al. Personalized oncology through integrative high-throughput sequencing: a pilot study. Sci. Transl. Med. 3, ra121 (2011).
Craig, D.W. et al. Genome and transcriptome sequencing in prospective refractory metastatic triple negative breast cancer uncovers therapeutic vulnerabilities. Mol. Cancer Ther. 12, 104–116 (2013).
Liang, W.S. et al. Genome-wide characterization of pancreatic adenocarcinoma patients using next generation sequencing. PLoS ONE 7, e43192 (2012).
Hadd, A.G. et al. Targeted, high-depth, next-generation sequencing of cancer genes in formalin-fixed, paraffin-embedded and fine-needle aspiration tumor specimens. J. Mol. Diagn. 15, 234–247 (2013).
Kerick, M. et al. Targeted high throughput sequencing in clinical cancer settings: formaldehyde fixed-paraffin embedded (FFPE) tumor tissues, input amount and tumor heterogeneity. BMC Med. Genomics 4, 68 (2011).
Hiatt, J.B., Pritchard, C.C., Salipante, S.J., O'Roak, B.J. & Shendure, J. Single molecule molecular inversion probes for targeted, high-accuracy detection of low-frequency variation. Genome Res. 23, 843–854 (2013).
Gargis, A.S. et al. Assuring the quality of next-generation sequencing in clinical laboratory practice. Nat. Biotechnol. 30, 1033–1036 (2012).
Wagle, N. et al. High-throughput detection of actionable genomic alterations in clinical tumor samples by targeted, massively parallel sequencing. Cancer Discov. 2, 82–93 (2012).
Thomas, R.K. et al. Sensitive mutation detection in heterogeneous cancer specimens by massively parallel picoliter reactor sequencing. Nat. Med. 12, 852–855 (2006).
Beltran, H. et al. Targeted next-generation sequencing of advanced prostate cancer identifies potential therapeutic targets and disease heterogeneity. Eur. Urol. 63, 920–926 (2013).
Giulino-Roth, L. et al. Targeted genomic sequencing of pediatric Burkitt lymphoma identifies recurrent alterations in anti-apoptotic and chromatin-remodeling genes. Blood 120, 5181–5184 (2012).
Lipson, D. et al. Identification of new ALK and RET gene fusions from colorectal and lung cancer biopsies. Nat. Med. 18, 382–384 (2012).
Siva, N. 1000 Genomes project. Nat. Biotechnol. 26, 256 (2008).
Lovly, C.M. et al. Potentially actionable kinase fusions in inflammatory myofibroblastic tumors. J. Clin. Oncol. 31, supplement, abstract 10513 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Albers, C.A. et al. Dindel: accurate indel calls from short-read data. Genome Res. 21, 961–973 (2011).
Carter, S.L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl. Acad. Sci. USA 107, 16910–16915 (2010).
Yau, C. et al. A statistical approach for detecting genomic aberrations in heterogeneous tumor samples from single nucleotide polymorphism genotyping data. Genome Biol. 11, R92 (2010).
Kim, E.S. et al. The BATTLE trial: personalizing therapy for lung cancer. Cancer Discov. 1, 44–53 (2011).
Forbes, S.A. et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 39, D945–D950 (2011).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
MacConaill, L.E. & Garraway, L.A. Clinical implications of the cancer genome. J. Clin. Oncol. 28, 5219–5228 (2010).
Swanton, C. My Cancer Genome: a unified genomics and clinical trial portal. Lancet Oncol. 13, 668–669 (2012).
Beadling, C. et al. Combining highly multiplexed PCR with semiconductor-based sequencing for rapid cancer genotyping. J. Mol. Diagn. 15, 171–176 (2013).
Bose, R. et al. Activating HER2 mutations in HER2 gene amplification negative breast cancer. Cancer Discov. 3, 224–237 (2013).
Fisher, S. et al. A scalable, fully automated process for construction of sequence-ready human exome targeted capture libraries. Genome Biol. 12, R1 (2011).
Karolchik, D. et al. The UCSC Table Browser data retrieval tool. Nucleic Acids Res. 32, D493–D496 (2004).
Gnirke, A. et al. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nat. Biotechnol. 27, 182–189 (2009).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Compeau, P.E., Pevzner, P.A. & Tesler, G. How to apply de Bruijn graphs to genome assembly. Nat. Biotechnol. 29, 987–991 (2011).
Djordjevic, B. et al. Clinical assessment of PTEN loss in endometrial carcinoma: immunohistochemistry outperforms gene sequencing. Mod. Pathol. 25, 699–708 (2012).
Reis-Filho, J.S. et al. Cyclin D1 protein overexpression and CCND1 amplification in breast carcinomas: an immunohistochemical and chromogenic in situ hybridisation analysis. Mod. Pathol. 19, 999–1009 (2006).
Sherry, S.T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Acknowledgements
The authors would like to acknowledge L. Gay for advice and assistance with manuscript preparation. H.B. is the Damon Runyon-Gordon Family Clinical Investigator supported (in part) by the Damon Runyon Cancer Research Foundation (CI-67-13). M.L. was supported by a Wellcome Trust Fellowship (WT093855MA) and by the Austrian Science Fund (J2856).
Author information
Authors and Affiliations
Contributions
G.M.F., D.L. and R.Y. designed the study, wrote the manuscript and developed and/or performed analyses. M.T.C. and P.J.S. designed the study and wrote the manuscript. V.A.M., J.S.R. and M.F.B. wrote the manuscript. G.A.O. designed the study. A.F., K.W., J.H., M.S.-L., J.W., E.M.S., P.A., J.S. and C.V. developed and/or performed analyses. G.A.O., S.R.D., K.B., F.J., V.B., S.B., J.B., A.D., L.G., K.I., A.M., K.M., T.R., S.T., E.W., M.Z., Z.Z., M.J., A.P., J.S.R. and J.C. planned and/or performed laboratory experiments. H.B., J.M.M., M.A.R., S.D., C.V.H., M.F.B., L.P., M.L. and C.B. planned and/or performed confirmatory experiments.
Corresponding authors
Ethics declarations
Competing interests
All authors with a Foundation Medicine affiliation are current or former employees of and stockholders in Foundation Medicine. M.B. is a former consultant to Foundation Medicine.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1–15 (PDF 2254 kb)
Supplementary Table 4C
Supplementary Table 4 (XLSX 366 kb)
Supplementary Table 7C
Supplementary Table 7 (XLSX 43 kb)
Supplementary Table 15
Supplementary Table 15 (XLSX 49 kb)
Rights and permissions
About this article
Cite this article
Frampton, G., Fichtenholtz, A., Otto, G. et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol 31, 1023–1031 (2013). https://doi.org/10.1038/nbt.2696
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2696
This article is cited by
-
Germline BRCA2 pathogenic variants in pediatric ganglioglioma: Case report and review of the literature
Child's Nervous System (2024)
-
Biomarker Testing Journey Among Patients with Advanced Solid Tumors and Treatment Patterns by Homologous Recombination Repair Status: A Clinico-Genomic Database Study
Advances in Therapy (2024)
-
Multiple PIK3CA mutation clonality correlates with outcomes in taselisib + fulvestrant-treated ER+/HER2–, PIK3CA-mutated breast cancers
Genome Medicine (2023)
-
Circulating tumor DNA landscape and prognostic impact of acquired resistance to targeted therapies in cancer patients: a national center for precision medicine (PRISM) study
Molecular Cancer (2023)
-
Overall survival in patients with advanced non-small cell lung cancer with KRAS G12C mutation with or without STK11 and/or KEAP1 mutations in a real-world setting
BMC Cancer (2023)