Abstract
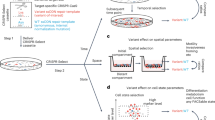
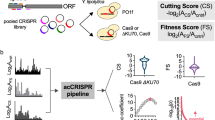
Construction and characterization of large genetic variant libraries is essential for understanding genome function, but remains challenging. Here, we introduce a Cas9-based approach for generating pools of mutants with defined genetic alterations (deletions, substitutions, and insertions) with an efficiency of 80–100% in yeast, along with methods for tracking their fitness en masse. We demonstrate the utility of our approach by characterizing the DNA helicase SGS1 with small tiling deletion mutants that span the length of the protein and a series of point mutations against highly conserved residues in the protein. In addition, we created a genome-wide library targeting 315 poorly characterized small open reading frames (smORFs, <100 amino acids in length) scattered throughout the yeast genome, and assessed which are vital for growth under various environmental conditions. Our strategy allows fundamental biological questions to be investigated in a high-throughput manner with precision.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Ben-Shitrit, T. et al. Systematic identification of gene annotation errors in the widely used yeast mutation collections. Nat. Methods 9, 373–378 (2012).
DiCarlo, J.E. et al. Genome engineering in Saccharomyces cerevisiae using CRISPR-Cas systems. Nucleic Acids Res. 41, 4336–4343 (2013).
Garst, A.D. et al. Genome-wide mapping of mutations at single-nucleotide resolution for protein, metabolic and genome engineering. Nat. Biotechnol. 35, 48–55 (2017).
Bao, Z. et al. Homology-integrated CRISPR-Cas (HI-CRISPR) system for one-step multigene disruption in Saccharomyces cerevisiae. ACS Synth. Biol. 4, 585–594 (2015).
Choulika, A., Perrin, A., Dujon, B. & Nicolas, J.F. Induction of homologous recombination in mammalian chromosomes by using the I-SceI system of Saccharomyces cerevisiae. Mol. Cell. Biol. 15, 1968–1973 (1995).
Brenneman, M., Gimble, F.S. & Wilson, J.H. Stimulation of intrachromosomal homologous recombination in human cells by electroporation with site-specific endonucleases. Proc. Natl. Acad. Sci. USA 93, 3608–3612 (1996).
Donoho, G., Jasin, M. & Berg, P. Analysis of gene targeting and intrachromosomal homologous recombination stimulated by genomic double-strand breaks in mouse embryonic stem cells. Mol. Cell. Biol. 18, 4070–4078 (1998).
Smih, F., Rouet, P., Romanienko, P.J. & Jasin, M. Double-strand breaks at the target locus stimulate gene targeting in embryonic stem cells. Nucleic Acids Res. 23, 5012–5019 (1995).
Taghian, D.G. & Nickoloff, J.A. Chromosomal double-strand breaks induce gene conversion at high frequency in mammalian cells. Mol. Cell. Biol. 17, 6386–6393 (1997).
Lu, J. et al. Human homologues of yeast helicase. Nature 383, 678–679 (1996).
Ira, G., Malkova, A., Liberi, G., Foiani, M. & Haber, J.E. Srs2 and Sgs1-Top3 suppress crossovers during double-strand break repair in yeast. Cell 115, 401–411 (2003).
Miyajima, A. et al. Different domains of Sgs1 are required for mitotic and meiotic functions. Genes Genet. Syst. 75, 319–326 (2000).
Mullen, J.R., Kaliraman, V. & Brill, S.J. Bipartite structure of the SGS1 DNA helicase in Saccharomyces cerevisiae. Genetics 154, 1101–1114 (2000).
Weinstein, J. & Rothstein, R. The genetic consequences of ablating helicase activity and the Top3 interaction domain of Sgs1. DNA Repair (Amst.) 7, 558–571 (2008).
Boddy, M.N. et al. Damage tolerance protein Mus81 associates with the FHA1 domain of checkpoint kinase Cds1. Mol. Cell. Biol. 20, 8758–8766 (2000).
Mullen, J.R., Kaliraman, V., Ibrahim, S.S. & Brill, S.J. Requirement for three novel protein complexes in the absence of the Sgs1 DNA helicase in Saccharomyces cerevisiae. Genetics 157, 103–118 (2001).
Chu, W.K. & Hickson, I.D. RecQ helicases: multifunctional genome caretakers. Nat. Rev. Cancer 9, 644–654 (2009).
Gangloff, S., McDonald, J.P., Bendixen, C., Arthur, L. & Rothstein, R. The yeast type I topoisomerase Top3 interacts with Sgs1, a DNA helicase homolog: a potential eukaryotic reverse gyrase. Mol. Cell. Biol. 14, 8391–8398 (1994).
Bennett, R.J., Sharp, J.A. & Wang, J.C. Purification and characterization of the Sgs1 DNA helicase activity of Saccharomyces cerevisiae. J. Biol. Chem. 273, 9644–9650 (1998).
Ui, A. et al. The ability of Sgs1 to interact with DNA topoisomerase III is essential for damage-induced recombination. DNA Repair (Amst.) 4, 191–201 (2005).
Kennedy, J.A., Daughdrill, G.W. & Schmidt, K.H. A transient α-helical molecular recognition element in the disordered N-terminus of the Sgs1 helicase is critical for chromosome stability and binding of Top3/Rmi1. Nucleic Acids Res. 41, 10215–10227 (2013).
Bennett, R.J., Noirot-Gros, M.F. & Wang, J.C. Interaction between yeast sgs1 helicase and DNA topoisomerase III. J. Biol. Chem. 275, 26898–26905 (2000).
Dunø, M., Thomsen, B., Westergaard, O., Krejci, L. & Bendixen, C. Genetic analysis of the Saccharomyces cerevisiae Sgs1 helicase defines an essential function for the Sgs1-Top3 complex in the absence of SRS2 or TOP1. Mol. Gen. Genet. 264, 89–97 (2000).
Fricke, W.M., Kaliraman, V. & Brill, S.J. Mapping the DNA topoisomerase III binding domain of the Sgs1 DNA helicase. J. Biol. Chem. 276, 8848–8855 (2001).
Ui, A. et al. The N-terminal region of Sgs1, which interacts with Top3, is required for complementation of MMS sensitivity and suppression of hyper-recombination in sgs1 disruptants. Mol. Genet. Genomics 265, 837–850 (2001).
Onodera, R. et al. Functional and physical interaction between Sgs1 and Top3 and Sgs1-independent function of Top3 in DNA recombination repair. Genes Genet. Syst. 77, 11–21 (2002).
Cejka, P., Plank, J.L., Bachrati, C.Z., Hickson, I.D. & Kowalczykowski, S.C. Rmi1 stimulates decatenation of double Holliday junctions during dissolution by Sgs1-Top3. Nat. Struct. Mol. Biol. 17, 1377–1382 (2010).
Kastenmayer, J.P. et al. Functional genomics of genes with small open reading frames (sORFs) in S. cerevisiae. Genome Res. 16, 365–373 (2006).
Balakrishnan, R. et al. YeastMine-an integrated data warehouse for Saccharomyces cerevisiae data as a multipurpose tool-kit. Database. (Oxford) 2012, bar062 (2012).
Chen, X. & Zhang, J. The genomic landscape of position effects on protein expression level and noise in yeast. Cell Syst. 2, 347–354 (2016).
Horwitz, A.A. et al. Efficient multiplexed integration of synergistic alleles and metabolic pathways in yeasts via CRISPR-Cas. Cell Syst. 1, 88–96 (2015).
Munoz, D.M. et al. CRISPR screens provide a comprehensive assessment of cancer vulnerabilities but generate false-positive his for highly amplified genomic regions. Cancer Discov. 6, 900–913 (2016).
Shi, J. et al. Discovery of cancer drug targets by CRISPR-Cas9 screening of protein domains. Nat. Biotechnol. 33, 661–667 (2015).
Michel, A.H. et al. Functional mapping of yeast genomes by saturated transposition. eLife 6, e23570 (2017).
Fowler, D.M. et al. High-resolution mapping of protein sequence-function relationships. Nat. Methods 7, 741–746 (2010).
Fowler, D.M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11, 801–807 (2014).
Starita, L.M. et al. Variant interpretation: functional assays to the rescue. Am. J. Hum. Genet. 101, 315–325 (2017).
Weile, J. et al. A framework for exhaustively mapping functional missense variants. Mol. Syst. Biol. 13, 957 (2017).
Kleinstiver, B.P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Christianson, T.W., Sikorski, R.S., Dante, M., Shero, J.H. & Hieter, P. Multifunctional yeast high-copy-number shuttle vectors. Gene 110, 119–122 (1992).
Kusano, K., Berres, M.E. & Engels, W.R. Evolution of the RECQ family of helicases: a drosophila homolog, Dmblm, is similar to the human bloom syndrome gene. Genetics 151, 1027–1039 (1999).
Yao, W. et al. The INO80 complex requires the Arp5-les6 subcomplex for chromatin remodeling and metabolic regulation. Mol. Cell. Biol. 36, 979–991 (2016).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Rost, B., Yachdav, G. & Liu, J. The PredictProtein server. Nucleic Acids Res. 32, W321–6 (2004).
Kaas, C.S., Kristensen, C., Betenbaugh, M.J. & Andersen, M.R. Sequencing the CHO DXB11 genome reveals regional variations in genomic stability and haploidy. BMC Genomics 16, 160 (2015).
Paix, A. et al. Scalable and versatile genome editing using linear DNAs with microhomology to Cas9 Sites in Caenorhabditis elegans. Genetics 198, 1347–1356 (2014).
Acknowledgements
G.M.C. was supported by NIH grants RM1 HG008525 and P50 HG005550. A.C. was funded by the National Cancer Institute grant no. 5T32CA009216-34. J.J.C. was funded by the Defense Threat Reduction Agency grant HDTRA1-14-1-0006, the Paul G. Allen Frontiers Group. Y.Y. was supported by the Damon Runyon Research Foundation grant DRG-2248-16.
Author information
Authors and Affiliations
Contributions
X.G. and A.C. conceived the idea, led the study, and designed all experiments. A.C. and R.C. with input from J.E.D. demonstrated the initial feasibility of the guide+donor approach. X.G. performed majority of the experiments, including the oligo library design, library construction and analysis, with significant technical contribution from A.T. Y.C. provided expertise in statistical analysis. Y.Y. performed the whole genome sequencing experiment for off-target analysis. C.K. generated the RNA-seq data for the BY4741 yeast strain, provided the FPKM values and analyzed the whole genome data from yeast isolates modified by guide+donor for off-target effects. S.L.G. assisted with oligo library design. E.K. provided insight with regard to library construction methods and analysis. M.S. provided technical expertise with regard to methods to increase guide+donor efficiency. J.J.C. and G.M.C. oversaw the study. X.G. and A.C. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
G.M.C. is the founder and holds leadership positions in many companies (http://arep.med.harvard.edu/gmc/tech.html). X.G., A.C., M.S., and E.K. have filed a patent application (US Patent Application 62/348,438) relating to this work.
Integrated supplementary information
Supplementary Figure 1 Enhanced transformation efficiency with engineered guide+donor approach.
a Schematic representations of unmodified guide+donor configuration (circular) and our engineered version (linearized) along with their corresponding transformation efficiencies. SNR52 promoter, N20, donor, and URA3 marker are labeled. Imaged plates are representative of 34 guide+donor yeast transformations in b targeting 34 different genomic sites performed with both configurations. b Scatterplot comparing transformation efficiencies of the selected 34 guide+donor plasmids between the unmodified and the engineered approaches in yeast strains expressing (blue) and not expressing Cas9 (orange). Genomic target corresponds to each guide+donor is indicated. Each guide+donor yeast transformation was performed twice. Relevant genotype of yeast strain background is exhibited on the x-axis. Average fold change in transformation efficiency is indicated on the y-axis and was calculated based on the number of transformants obtained from the engineered linearized plasmid setup divided by the number of transformants obtained from the circular plasmid transformation.
Supplementary Figure 2 Guide+donor editing efficiency of SNPs at various distances from the cut site.
Frequency of correct SNP edits are displayed on the y-axis with varying distances from the Cas9-generated cut site indicated on the x-axis. Position of cut site is marked by the middle vertical line (red). Number of samples genotyped are indicated.
Supplementary Figure 3 Examining the influence of homology lengths on transformation efficiency of SNPs further away from the PAM.
a Schematic representation of a region of ADE2 centered around an N20-targeting site with SNPs (blue), SNPs distances, various homology lengths, cut site position, and wildtype and mutated PAMs (red) indicated. b Transformation efficiency of ADE2 guide+donors with varying donor homology lengths and SNPs distances from PAM as indicated on the legend. Transformation efficiency was calculated by the number of colonies obtained in the presence of Cas9 divided by the number of colonies in the absence of Cas9. Average transformation efficiency is displayed on the y-axis (n=2 independent cultures).
Supplementary Figure 4 Testing different design parameters for efficient programmed genomic edits at the ADE2 locus.
a Frequency of correct programmed deletions are indicated on the y-axis with increasing sizes of deletions labeled on the x-axis. b Frequency of correct edits on the y-axis with replacement of a 61bp endogenous sequence region by increasing sizes of a linker sequence as denoted on the x-axis.
Supplementary Figure 5 Whole genome sequencing of guide+donor-edited yeast strains to determine off-target effects.
Whole genome sequence alignment of strains transformed with guide+donors targeting (a) ADE2 to delete a w61bp region, (b) SGS1 to delete a 60bp fragment, or (c) 3bp ATG start site.
Supplementary Figure 6 Construction and functional interrogation of guide+donor library targeting C-terminus of SGS1 with controls targeting ARS214.
a Graphical representation of programmed edits in ARS214 (grey) and SGS1 (blue) targeting C-terminus generated by guide+donor constructs followed by phenotypic characterization in b and c. Asterisk, dotted box, and solid dash denote substitution, deletion, and replacement of an amino acid stretch with a linker sequence, respectively. Figures not drawn to scale. b Dot plot showing fold change of guide+donor constructs targeting ARS214 and SGS1 in response to HU condition. Types of genetic modifications are labeled on the x-axis. Depletion is represented in log2 scale on the y-axis and compared between ARS214 and SGS1. Yeast library transformation was performed twice. c Dot plot showing depletion of ARS214 and SGS1 guide+donor constructs in mms4Δ genetic background. Genetic modifications and their corresponding abundance in log2 scale are indicated on x- and y-axes, respectively. Two independent yeast library transformations were transformed.
Supplementary Figure 7 Phenotypic testing of individual guide+donor generated sgs1 protein variants in response to HU.
Cells were grown to log phase. 3μl of each undiluted and 5-fold serially diluted culture were spotted onto control SC-URA plate or SC-URA plate containing 40mM HU. All plates were incubated at 30ºC for 48 hours and photographed.
Supplementary Figure 8 Phenotypic validation of individual mutants from sgs1 amino acid library screen.
Wildtype and selected sensitive and non-sensitive hits from HU library screen are indicated. 3μl of each undiluted and 5-fold serially diluted culture were spotted onto control SC-URA plate or SC-URA plate containing 40mM HU. All plates were incubated at 30ºC for 48 hours and photographed.
Supplementary Figure 9 Replicate analysis of smORF library screens in response to various test conditions.
Replicate analysis of log2 fold changes between biological replicates of two independent yeast transformations for each smORF library screen in Figure 4 (a-d). Pearson correlation coefficient is indicated.
Supplementary Figure 10 Phenotypic validation of smORF mutants from heat and fluconazole screens.
Wildtype and selected sensitive and non-sensitive hits from smORF library screens are indicated. For the heat screen validation (a), cells were grown to log phase. 3μl of each undiluted and 5-fold serially diluted culture were spotted onto SC-URA plates that were incubated at the indicated temperatures for 48 hours and photographed. For the fluconazole screen validation (b), cells were cultured in a similar manner followed by spotting 5μl of each undiluted and 5-fold serially diluted culture onto SC-URA plates with and without fluconazole. Plates were incubated at 30ºC for 48 hours and photographed.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 (PDF 953 kb)
Supplementary Tables
Supplementary tables1–7 (PDF 273 kb)
Supplementary Data 1
Log2-fold changes in abundance of guide+donor members in SGS1 tiling deletion screen (XLSX 60 kb)
Supplementary Data 2
Log2-fold changes in abundance of guide+donor members in Sgs1 conserved residue amino acid substitution screen (XLSX 49 kb)
Supplementary Data 3
Log2-fold changes in abundance of guide+donor members in smORF screen (XLSX 404 kb)
Supplementary Data 4
Analysis of amino acid length, gene expression, prediction of secondary structure formation and level of conservation in humans between smORFs and ORFs (XLSX 45 kb)
Rights and permissions
About this article
Cite this article
Guo, X., Chavez, A., Tung, A. et al. High-throughput creation and functional profiling of DNA sequence variant libraries using CRISPR–Cas9 in yeast. Nat Biotechnol 36, 540–546 (2018). https://doi.org/10.1038/nbt.4147
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4147
This article is cited by
-
Exploring microproteins from various model organisms using the mip-mining database
BMC Genomics (2023)
-
CRISPR/Cas9-mediated genome editing in diploid and tetraploid potatoes
Acta Physiologiae Plantarum (2023)
-
High-content CRISPR screening
Nature Reviews Methods Primers (2022)
-
Applications of CRISPR/Cas gene-editing technology in yeast and fungi
Archives of Microbiology (2022)
-
CRISPR/Cas-mediated knockdown of vacuolar invertase gene expression lowers the cold-induced sweetening in potatoes
Planta (2022)