Abstract
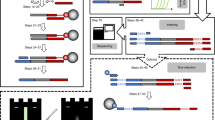
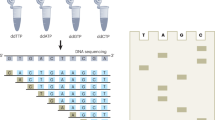
We have developed a high speed instrument for automated DNA sequence analysis. The apparatus employs laser excitation and a cooled CCD detector for the parallel detection of up to 18 sets of four fluorescently labeled DNA sequencing reactions during their electrophoretic separation in ultrathin (50–100 μm) denaturing polyacrylamide gels. Four hundred and fifty bases of sequence information is obtained from 100 ng of M13 template DNA in less than one hour, corresponding to an overall instrument throughput of over 8000 bases/hr.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Hood, L.E., Hunkapiller, M.W. and Smith, L.M. 1987. Automated DNA sequencing and analysis of the human genome. Genomics 1: 201–212.
Drossman, H., Luckey, J.A., Kostichka, A.J., D'Cunha, J. and Smith, L.M. 1990. High speed separations of DNA sequencing reactions by capillary electrophoresis. Anal. Chem. 62: 900–903.
Guttman, A., Cohen, A.S., Heiger, D.N. and Karger, B.L. 1990. Analytical and micropreparative ultrahigh resolution of oligonucleotides by polyacrylamide gel high-performance capillary electrophoresis. Anal. Chem. 62: 137–141.
Karger, B.L. 1990. High-performance capillary electrophoresis. Nature 339: 641–642.
Luckey, J.A., Drossman, H., Kostichka, A.J., Mead, D.A., D'Cunha, J., Norris, T.B. and Smith, L.M. 1990. High speed DNA sequencing by capillary electrophoresis. Nuc. Acids Res. 18: 4417–4421.
Swerdlow, H. and Gesteland, R. 1990. Capillary gel electrophoresis for rapid, high resolution DNA sequencing. Nuc. Acids Res. 18: 1415–1419.
Connell, C., Fung, S., Heiner, C., Bridgham, J., Chakerian, V., Heron, E., Jones, B., Menchen, S., Mordan, W., Raff, M., Recknor, M., Smith, L., Springer, J., Woo, S. and Hunkapiller, M. 1987. Automated DNA sequence analysis. Biotechniques 5: 342–348.
Smith, L.M., Sanders, J.Z., Kaiser, R.J., Hughes, P., Dodd, C., Connell, C.R., Heiner, C., Kent, S.B.H. and Hood, L.E. 1986. Fluorescence detection in automated DNA sequence analysis. Nature 321: 674–679.
Smith, L.M., Fung, S., Hunkapiller, M., Hunkapiller, T. and Hood, L.E. 1985. The synthesis of oligonucleotides containing an aliphatic amino group at the 5′ terminus: Synthesis of fluorescent DNA primers for use in DNA sequence analysis. Nuc. Acids Res. 13: 2399–2412.
Brumley, R.L. and Smith, L.M. Rapid DNA sequencing by horizontal ultrathin gel electrophoresis. 1991. Nuc. Acids Res. 19: 4121–4126.
Linderman, J.J., Harris, L.J., Slakey, L.L. and Gross, D.J. 1990. Charge-coupled device imaging of rapid calcium transients in cultured arterial smooth muscle cells. Cell Calcium 11: 131–144.
Kambara, H., Nagai, K. and Hayasaka, S. 1991. Real time automated simultaneous double-stranded DNA sequencing using two-color fluorophore labeling. Bio/Technology 9: 648–651.
Epperson, P.M. and Denton, M.B. 1989. Binning spectral images in a charge-coupled device. Anal. Chem. 61: 1513–1519.
Smith, L.M., Kaiser, R.J. and Hood, L.E. 1987. The synthesis and use of fluorescent oligonucleotides in DNA sequence analysis. Methods in Enzymol. 155: 260–301.
Smith, L.M. 1989. DNA sequence analysis: past, present, and future. Amer. Biotech. Lab. 7: 10–25.
Mathies, R.A., Peck, K. and Stryer, L. 1990. Optimization of high-sensitivity fluorescence detection. Anal. Chem. 62: 1786–1791.
Ansorge, W., Sproat, B., Stegemann, J., Schwager, C. and Zenke, M. 1987. Automated DNA sequencing: ultrasensitive detection of fluorescent bands during electrophoresis. Nuc. Acids Res. 15: 4593–4602.
Stegemann, J., Schwager, C., Erfle, H., Hewitt, N., Voss, H., Zimmerman, J. and Ansorge, W. 1991. High speed on-line DNA sequencing on ultrathin slab gels. Nuc. Acids Res. 19: 675–676.
Lumpkin, O.J., Dejardin, P. and Zimm, B.H. 1985. Theory of gel electrophoresis of DNA. Biopolymers 24: 1573–1593.
Hervet, H. and Bean, C.P. 1987. Electrophoretic mobility of lambda phage HindIII and HaeIII DNA fragments in agarose gels: a detailed study. Biopolymers 26: 727–742.
Holmes, D.L. and Stellwagen, N.C. 1990. The electric field dependence of DNA mobilities in agarose gels: a reinvestigation. Electrophoresis 11: 5–15.
Ulanovsky, L., Drouin, G. and Gilbert, W. 1990. DNA trapping electrophoresis. Nature 343: 190–192.
Kambara, H., Nishikawa, T., Katayama, Y. and Yamaguchi, T. 1988. Optimization of parameters in a DNA sequenator using fluorescence detection. Bio/Technology 6: 816–821.
Mead, D.A., McClary, J.A., Luckey, J.A., Kosticka, A.J., Witney, F.R. and Smith, L.M. 1991. Bst DNA polymerase permits rapid sequence analysis from nanogram amounts of template. Biotechniques 11: 76–87.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Kostichka, A., Marchbanks, M., Brumley, R. et al. High Speed Automated DNA Sequencing in Ultrathin Slab Gels. Nat Biotechnol 10, 78–81 (1992). https://doi.org/10.1038/nbt0192-78
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0192-78