Abstract
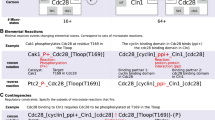
Accurate simulation of intracellular biochemical networks is essential to furthering our understanding of biological system behavior. The number of protein complexes and of chemical interactions among them has traditionally posed significant problems for simulation algorithms. Here we describe an approach to the exact stochastic simulation of biochemical networks that emphasizes the contribution of protein complexes to these systems. This simulation approach starts from a description of monomeric proteins and specifications for binding, unbinding and other reactions. This manageable specification is reasonably intuitive for biologists. Rather than requiring the inclusion of all possible complexes and reactions from the outset, our approach incorporates new complexes and reactions only when needed as the simulation proceeds. As a result, the simulation generates much smaller reaction networks, which can be exported to other simulators for further analysis. We apply this approach to the automatic generation of reaction systems for the study of signal transduction networks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Frenkel, D. & Smit, B. Understanding Molecular Simulation (Academic Press, San Diego, California, 1996).
Pauling, L. The Nature of the Chemical Bond and the Structure of Molecules and Crystals (Cornell University Press, Ithaca, New York, 1960).
Vaidehi, N. & Goddard, W. Atomic-level simulation and modeling of biomacromolecules. in Computational Modeling of Genetic and Biochemical Networks. (eds. Bower, J. & Bolouri, H.) 161–188 (MIT Press, Cambridge, Massachusetts, 2001).
Gillespie, D. A rigorous derivation of the chemical master equation. Physica A 188, 404–425 (1992).
Elowitz, M., Surrete, M., Wolf, P., Stock, J. & Leibler, S. Protein mobility in the cytoplasm of Escherichia coli. J. Bacteriol. 181, 197–203 (1999).
Gillespie, D. Markov processes: an introduction for physical scientists (Academic Press, Boston, Massachusetts, 1992).
Gillespie, D. A general method for numerically simulating the stochastic time evolution of coupled chemical reactions. J. Comp. Phys. 22, 403–434 (1976).
Deuflhard, P. & Bornemann, F. Scientific Computing with Ordinary Differential Equations (Springer-Verlag, New York, 2002).
Elowitz, M. et al. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
McAdams, H. & Arkin, A. Stochastic mechanisms in gene expression. Proc. Natl. Acad. Sci. USA 94, 814–819 (1997).
Rao, C., Wolf, D. & Arkin, A. Control, exploitation and tolerance of intracellular noise. Nature 420, 231–237 (2002).
Mendes, P. Computer simulation of the dynamics of biochemical pathways. PhD thesis, University of Wales Aberystwyth (1994).
Cross, F., Archambault, V., Miler, M. & Klovstad, M. Testing a mathematical model of the yeast cell cycle. Mol. Biol. Cell 13, 52–70 (2002).
Chen, K.C. et al. Kinetic analysis of a molecular model of the budding yeast cell cycle. Mol. Biol. Cell 11, 369–391 (2000).
Bormann, G., Brosens, F. & De Schutter, E. Diffusion. in Computational Modeling of Genetic and Biochemical Networks. (eds. Bower, J. & Bolouri, H.) 189–224 (MIT Press, Cambridge, Massachusetts, 2001).
Gillespie, D. Approximate accelerated stochastic simulation of chemically reacting systems. J. Chem. Phys. 115, 1716–1733 (2001).
Gibson, M. Computational methods for stochastic biological systems. PhD Thesis, California Institute of Technology (2000).
Gibson, M. & Bruck, J. Efficient exact stochastic simulation of chemical systems with many species and many channels. J. Phys. Chem. 104, 1876–1889 (1999).
Morton-Firth, C. Stochastic simulation of cell signalling pathways. PhD thesis, University of Cambridge (1998).
Gillespie, D. & Petzold, L. Improved leap-size selection for accelerated stochastic simulation. J. Chem. Phys. 119, 8229–8234 (2003).
Haseltine, E. & Rawlings, J. Approximate simulation of coupled fast and slow reactions for stochastic chemical kinetics. J. Chem. Phys. 117, 6959–6969 (2002).
Rao, C. & Arkin, A. Stochastic chemical kinetics and the quasi-steady-state assumption: Application to the Gillespie algorithm. J. Chem. Phys. 118, 4999–5010 (2003).
Keane, J., Bradley, C. & Eberling, C. A compiled accelerator for biological cell signaling simulations. ACM SIGDA Int. Symp. Field Program Gate Arrays FPGA 12, 233–241 (2004).
Salwinski, L. & Eisenberg, D. In silico simulation of biological network dynamics. Nat. Biotechnol. 22, 1017–1019 (2004).
Fricke, T. & Wendt, D. The Markoff automaton: a new algorithm for simulating the time-evolution of large stochastic dynamic systems. Int. J. Mod. Phys. 6, 277–306 (1995).
Stiles, J. & Bartol, T. Monte Carlo methods for simulating realistic synaptic microphysiology using MCell. in Computational Neuroscience: Realistic Modeling for Experimentalists. (ed. de Schutter, E.) 87–127 (CRC Press, Boca Raton, Florida, 2000).
Hodges, P., Payne, W. & Garrels, J. The yeast protein database (YPD): a curated proteome database for Saccharomyces cerevisiae. Nucleic Acids Res. 26, 68–72 (1998).
Ptashne, M. A genetic switch: phage λ and higher organisms (Blackwell Scientific Publications, Cambridge, Massachusetts, 1992).
Bray, D. & Lay, S. Computer-based analysis of the binding steps in protein complex formation. Proc. Natl. Acad. Sci. USA 94, 13493–13498 (1997).
Brent, R. Genomic biology. Cell 100, 169–183 (2000).
Endy, D. & Brent, R. Modelling cellular behavior. Nature 409, 391–395 (2001).
Dohlman, H. & Thorner, J. Regulation of G-protein initiated signal transduction in yeast: Paradigms and principles. Annu. Rev. Biochem. 70 (2001).
Press, W., Teukolsky, S., Vetterling, W. & Flannery, B. Numerical Recipes in C, edn. 2 (Cambridge University Press, Cambridge, 1992).
Hucka, M. et al. The systems biology markup language (SBML): A medium for representation and exchange of biochemical network models. Bioinformatics 19, 524–531 (2003).
Acknowledgements
L.L. conceived, designed and programmed Moleculizer. R.B. started and fostered the program of quantitative biological inquiry that enabled the development of Moleculizer and helped frame explanations of the work for biologists. L.L. and R.B. wrote the manuscript and stand as guarantors of its veracity. The work was supported by a grant from the US Defense Advanced Research Projects Agency and by the “Alpha Project” at the Center for Genomic Experimentation and Computation, a National Institutes of Health Center of Excellence in Genomic Science. The Alpha Project is supported by grant P50 HG02370 to R.B. from the National Human Genome Research Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
An intentional combinatorial explosion generating many species of complexes. (PDF 42 kb)
Supplementary Fig. 2
Use of reperesentation–invariant hash (PDF 34 kb)
Supplementary Fig. 3
Major components of the alpha signal transduction pathway in yeast. (PDF 48 kb)
Supplementary Table 1
Performance effects of species and reaction generation on simulation of receptor part of the alpha pathway. (PDF 51 kb)
Supplementary Table 2
Performance effects of species and reaction generation on “full” alpha pathway simulation. (PDF 37 kb)
Supplementary Table 3
Main executables delivered with Moleculizer 1.0. (PDF 39 kb)
Supplementary Table 4
File formats connected with Moleculizer 1.0. (PDF 48 kb)
Rights and permissions
About this article
Cite this article
Lok, L., Brent, R. Automatic generation of cellular reaction networks with Moleculizer 1.0. Nat Biotechnol 23, 131–136 (2005). https://doi.org/10.1038/nbt1054
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1054
This article is cited by
-
Generalizing Gillespie’s Direct Method to Enable Network-Free Simulations
Bulletin of Mathematical Biology (2019)
-
Simulation tools for particle-based reaction-diffusion dynamics in continuous space
BMC Biophysics (2014)
-
Specification, annotation, visualization and simulation of a large rule-based model for ERBB receptor signaling
BMC Systems Biology (2012)
-
Computational modeling of cellular signaling processes embedded into dynamic spatial contexts
Nature Methods (2012)
-
A framework for mapping, visualisation and automatic model creation of signal‐transduction networks
Molecular Systems Biology (2012)