Abstract
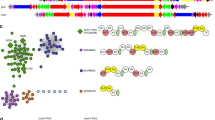
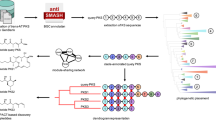
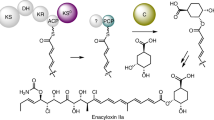
Modular polyketide synthases (PKSs) are giant bacterial enzymes that synthesize many polyketides of therapeutic value. In contrast to PKSs that provide acyltransferase (AT) activities in cis, trans-AT PKSs lack integrated AT domains and exhibit unusual enzymatic features with poorly understood functions in polyketide assembly. This has retarded insight into the assembly of products such as mupirocin, leinamycin and bryostatin 1. We show that trans-AT PKSs evolved in a fundamentally different fashion from cis-AT systems, through horizontal recruitment and assembly of substrate-specific ketosynthase (KS) domains. The insights obtained from analysis of these KS mosaics will facilitate both the discovery of novel polyketides by genome mining, as we demonstrate for the thailandamides of Burkholderia thailandensis, and the extraction of chemical information from short trans-AT PCR products, as we show using metagenomic DNA of marine sponges. Our data also suggest new strategies for dissecting polyketide biosynthetic pathways and engineering polyketide assembly.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Fischbach, M.A. & Walsh, C.T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Chem. Rev. 106, 3468 (2006).
Piel, J. A polyketide synthase-peptide synthetase gene cluster from an uncultured bacterial symbiont of Paederus beetles. Proc. Natl. Acad. Sci. USA 99, 14002–14007 (2002).
Piel, J. et al. Antitumor polyketide biosynthesis by an uncultivated bacterial symbiont of the marine sponge Theonella swinhoei. Proc. Natl. Acad. Sci. USA 101, 16222–16227 (2004).
Mochizuki, S. et al. The large linear plasmid pSLA2-L of Streptomyces rochei has an unusually condensed gene organization for secondary metabolism. Mol. Microbiol. 48, 1501–1510 (2003).
Chen, X.H. et al. Structural and functional characterization of three polyketide synthase gene clusters in Bacillus amyloliquefaciens FZB 42. J. Bacteriol. 188, 4024–4036 (2006).
Sudek, S. et al. Identification of the putative bryostatin polyketide synthase gene cluster from “Candidatus Endobugula sertula”, the uncultivated microbial symbiont of the marine bryozoan Bugula neritina. J. Nat. Prod. 70, 67–74 (2007).
Perlova, O., Gerth, K., Kaiser, O., Hans, A. & Müller, R. Identification and analysis of the chivosazol biosynthetic gene cluster from the myxobacterial model strain Sorangium cellulosum So ce56. J. Biotechnol. 121, 174–191 (2006).
Cheng, Y.Q., Tang, G.L. & Shen, B. Type I polyketide synthase requiring a discrete acyltransferase for polyketide biosynthesis. Proc. Natl. Acad. Sci. USA 100, 3149–3154 (2003).
Schneider, K. et al. Macrolactin is the polyketide biosynthesis product of the pks2 cluster of Bacillus amyloliquefaciens FZB42. J. Nat. Prod. 70, 1417–1423 (2007).
El-Sayed, A.K. et al. Characterization of the mupirocin biosynthesis gene cluster from Pseudomonas fluorescens NCIMB 10586. Chem. Biol. 10, 419–430 (2003).
Simunovic, V. et al. Myxovirescin A biosynthesis is directed by hybrid polyketide synthases/nonribosomal peptide synthetase, 3-hydroxy-3-methylglutaryl-CoA synthases, and trans-acting acyltransferases. ChemBioChem 7, 1206–1220 (2006).
Paitan, Y., Alon, G., Orr, E., Ron, E.Z. & Rosenberg, E. The first gene in the biosynthesis of the polyketide antibiotic TA of Myxococcus xanthus codes for a unique PKS module coupled to a peptide synthetase. J. Mol. Biol. 286, 465–474 (1999).
Partida-Martinez, L.P. & Hertweck, C. A gene cluster encoding rhizoxin biosynthesis in “Burkholderia rhizoxina”, the bacterial endosymbiont of the fungus Rhizopus microsporus. ChemBioChem 8, 41–45 (2007).
Pulsawat, N., Kitani, S. & Nihira, T. Characterization of biosynthetic gene cluster for the production of virginiamycin M, a streptogramin type A antibiotic, in Streptomyces virginiae. Gene 393, 31–42 (2007).
Piel, J., Hui, D., Fusetani, N. & Matsunaga, S. Targeting polyketide synthases with iteratively acting acyltransferases from metagenomes of uncultured bacterial consortia. Environ. Microbiol. 6, 921–927 (2004).
Wenzel, S.C. & Müller, R. Formation of novel secondary metabolites by bacterial multimodular assembly lines: deviations from textbook biosynthetic logic. Curr. Opin. Chem. Biol. 9, 447–458 (2005).
Jenke-Kodama, H., Börner, T. & Dittmann, E. Natural biocombinatorics in the polyketide synthase genes of the actinobacterium Streptomyces avermitilis. PloS Comput. Biol. 2, e132 (2006).
Jenke-Kodama, H., Sandmann, A., Müller, R. & Dittmann, E. Evolutionary implications of bacterial polyketide synthases. Mol. Biol. Evol. 22, 2027–2039 (2005).
Fieseler, L. et al. Widespread occurrence and genomic context of unusually small polyketide synthase genes in microbial consortia associated with marine sponges. Appl. Environ. Microbiol. 73, 2144–2155 (2007).
Huelsenbeck, J.P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Ginolhac, A. et al. Type I polyketide synthases may have evolved through horizontal gene transfer. J. Mol. Evol. 60, 716–725 (2005).
Lopez, J.V. Naturally mosaic operons for secondary metabolite biosynthesis: variability and putative horizontal transfer of discrete catalytic domains of the epothilone polyketide synthase locus. Mol. Genet. Genomics 270, 420–431 (2003).
Davies, C., Heath, R.J., White, S.W. & Rock, C.O. The 1.8 angstrom crystal structure and active-site architecture of beta-ketoacyl-acyl carrier protein synthase III (FabH) from Escherichia coli. Structure 8, 185–195 (2000).
Calderone, C.T., Kowtoniuk, W.E., Kelleher, N.L., Walsh, C.T. & Dorrestein, P.C. Convergence of isoprene and polyketide biosynthetic machinery: Isoprenyl-S-carrier proteins in the pksX pathway of Bacillus subtilis. Proc. Natl. Acad. Sci. USA 103, 8977–8982 (2006).
Albertini, A.M., Caramori, T., Scoffone, F., Scotti, C. & Galizzi, A. Sequence around the 159° region of the Bacillus subtilis genome: the pksX locus spans 33.6 Kb. Microbiology 141, 299–309 (1995).
Patel, P.S. et al. Bacillaene, a novel inhibitor of prokaryotic protein synthesis produced by Bacillus subtilis: production, taxonomy, isolation, physicochemical characterization and biological activity. J. Antibiot. 48, 997–1003 (1995).
Butcher, R.A. et al. The identification of bacillaene, the product of the PksX megacomplex in Bacillus subtilis. Proc. Natl. Acad. Sci. USA 104, 1506–1509 (2007).
Partida-Martinez, L.P. & Hertweck, C. Pathogenic fungus harbours endosymbiotic bacteria for toxin production. Nature 437, 884–888 (2005).
Piel, J., Wen, G., Platzer, M. & Hui, D. Unprecedented diversity of catalytic domains in the first four modules of the putative pederin polyketide synthase. ChemBioChem 5, 93–98 (2004).
Kroken, S., Glass, N.L., Taylor, J.W., Yoder, O.C. & Turgeon, B.G. Phylogenomic analysis of type I polyketide synthase genes in pathogenic and saprobic ascomycetes. Proc. Natl. Acad. Sci. USA 100, 15670–15675 (2003).
Moffitt, M.C. & Neilan, B.A. Evolutionary affiliations within the superfamily of ketosynthases reflect complex pathway associations. J. Mol. Evol. 56, 446–457 (2003).
Hopwood, D.A. Genetic contributions to understanding polyketide synthases. Chem. Rev. 97, 2465–2497 (1997).
Stachelhaus, T., Mootz, H.D. & Marahiel, M.A. The specificity-conferring code of adenylation domains in nonribosomal peptide synthetases. Chem. Biol. 6, 493–505 (1999).
Challis, G.L., Ravel, J. & Townsend, C.A. Predictive, structure-based model of amino acid recognition by nonribosomal peptide synthetase adenylation domains. Chem. Biol. 7, 211–224 (2000).
Tang, Y. & Kim, C.Y. Mathews, II, Cane, D.E. & Khosla, C. The 2.7-angstrom crystal structure of a 194-kDa homodimeric fragment of the 6-deoxyerythronolide B synthase. Proc. Natl. Acad. Sci. USA 103, 11124–11129 (2006).
Fischbach, M.A. & Walsh, C.T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: Logic, machinery, and mechanisms. Chem. Rev. 106, 3468–3496 (2006).
Finking, R. & Marahiel, M.A. Biosynthesis of nonribosomal peptides. Annu. Rev. Microbiol. 58, 453–488 (2004).
Stewart, G.J. & Carlson, C.A. The biology of natural transformation. Annu. Rev. Microbiol. 40, 211–235 (1986).
Pfeifer, B.A., Admiraal, S.J., Gramajo, H., Cane, D.E. & Khosla, C. Biosynthesis of complex polyketides in a metabolically engineered strain of E coli. Science 291, 1790–1792 (2001).
Moldenhauer, J., Chen, X., Borriss, R. & Piel, J. Biosynthesis of the antibiotic bacillaene, the product of a giant polyketide synthase complex of the trans-AT family. Angew. Chem. Int. Edn Engl. 46, 8195–8197 (2007).
Suwa, M. et al. Identification of two polyketide synthase gene clusters on the linear plasmid pSLA2-L in Streptomyces rochei. Gene 246, 123–131 (2000).
Gross, H. Strategies to unravel the function of orphan biosynthesis pathways: recent examples and future prospects. Appl. Microbiol. Biotechnol. 75, 267–277 (2007).
Zazopoulos, E. et al. A genomics-guided approach for discovering and expressing cryptic metabolic pathways. Nat. Biotechnol. 21, 187–190 (2003).
McAlpine, J.B. et al. Microbial Genomics as a guide to drug discovery and structural elucidation: ECO-02301, a novel antifungal agent, as an example. J. Nat. Prod. 68, 493–496 (2005).
Donia, M.S. et al. Natural combinatorial peptide libraries in cyanobacterial symbionts of marine ascidians. Nat. Chem. Biol. 2, 729–735 (2006).
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F. & Higgins, D.G. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997).
Abascal, F., Zardoya, R. & Posada, D. ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21, 2104–2105 (2005).
Whelan, S. & Goldman, N. A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol. Biol. Evol. 18, 691–699 (2001).
Felsenstein, J. PHYLIP - Phylogeny Inference Package version 3.6. Cladistics 5, 164–166 (1989).
Hrvatin, S. & Piel, J. Rapid isolation of rare clones from highly complex DNA libraries by PCR analysis of liquid gel pools. J. Microbiol. Methods 68, 434–436 (2007).
Acknowledgements
We thank Andrea Perner, Heike Heinecke and Franziska Rhein for HRMS and NMR measurements, Daniela Werler for sequencing assistance, Martin Roth and Matthias Steinacker of the BioPilotPlant for assistance in fermentation and extraction, and Shigeki Matsunaga for a specimen of the sponge T. swinhoei. This work was supported by the Deutsche Forschungsgemeinschaft (DFG) (PI 430/1-3 and SFB 624 to J.P. and grants of the priority programs SPP1152: PI 430/5-2, Di 910/1-3 He 3268/2-3 to J.P., E.D. and C.H., respectively).
Author information
Authors and Affiliations
Contributions
J.P. designed research, performed alignments and phylogenetic calculations, analyzed data and wrote the manuscript; H.J.-K. and E.D. designed research and performed phylogenetic analysis; T.H. analyzed sequence data; T.N. and C.G. isolated and analyzed the PKS cluster from the sponge metagenome; S.T. and M.P. performed sequencing, C.H. designed research and analyzed data, K.I. isolated and characterized thailandamides.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Data 1,2; Table 1 (PDF 1533 kb)
Rights and permissions
About this article
Cite this article
Nguyen, T., Ishida, K., Jenke-Kodama, H. et al. Exploiting the mosaic structure of trans-acyltransferase polyketide synthases for natural product discovery and pathway dissection. Nat Biotechnol 26, 225–233 (2008). https://doi.org/10.1038/nbt1379
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1379
This article is cited by
-
Global characterization of biosynthetic gene clusters in non-model eukaryotes using domain architectures
Scientific Reports (2024)
-
Enzymology of assembly line synthesis by modular polyketide synthases
Nature Chemical Biology (2023)
-
Insights into azalomycin F assembly-line contribute to evolution-guided polyketide synthase engineering and identification of intermodular recognition
Nature Communications (2023)
-
Genome-based discovery and total synthesis of janustatins, potent cytotoxins from a plant-associated bacterium
Nature Chemistry (2022)
-
Strategies to access biosynthetic novelty in bacterial genomes for drug discovery
Nature Reviews Drug Discovery (2022)