Abstract
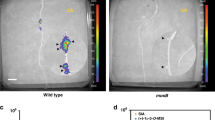
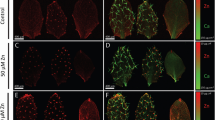
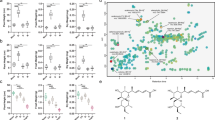
Plant roots release a range of enzymes capable of degrading chemical compounds in their immediate vicinity1,2. We present a system of phytoremediation ex planta based on the overexpression of one such enzyme, a secretory laccase. Laccases catalyze the oxidation of a broad range of phenolic compounds3, including polychlorinated phenols such as 2,4,6-trichlorophenol (TCP), that are among the most hazardous and recalcitrant pollutants in the environment4. We isolated a secretory laccase cDNA of LAC1, which is specifically expressed in the roots of Gossypium arboreum (cotton). Transgenic Arabidopsis thaliana plants overexpressing LAC1 exhibited enhanced resistance to several phenolic allelochemicals and TCP. The secretory laccase activity in these plants was responsible for the conversion of sinapic acid into a mono-lactone type dimer and for the transformation of TCP.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Inderjit & Duke, S.O. Ecophysiological aspects of allelopathy. Planta 217, 529–539 (2003).
Boyajian, G.E. & Carreira, L.H. Phytoremediation: a clean transition from laboratory to marketplace? Nat. Biotechnol. 15, 127–128 (1997).
Mayer, A.M. & Staples, R.C. Laccase: new functions for an old enzyme. Phytochemistry 60, 551–565 (2002).
Gupta, S.S. et al. Rapid total destruction of chlorophenols by activated hydrogen peroxide. Science 296, 326–328 (2002).
Nitta, K., Kataoka, K. & Sakurai, T. Primary structure of a Japanese lacquer tree laccase as a prototype enzyme of multicopper oxidases. J. Inorg. Biochem. 91, 125–131 (2002).
Sterjiades, R., Dean, J.F.D., Gamble, G., Himmelsbach, D.S. & Eriksson, K.-E.L. Extracellular laccases and peroxidases from sycamore maple (Acer pseudoplatanus) cell suspension cultures. Reactions with monolignols and lignin model compounds. Planta 190, 75–87 (1993).
Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Wu, H., Haig, T., Pratley, J., Lemerle, D. & An, M. Biochemical basis for wheat seedling allelopathy on the suppression of annual ryegrass (Lolium rigidum). J. Agric. Food Chem. 50, 4567–4571 (2002).
Chu, W. & Wong, C.C. A disappearance model for the prediction of trichlorophenol ozonation. Chemosphere 51, 289–294 (2003).
Leontievsky, A.A. et al. Transformation of 2,4,6-trichlorophenol by free and immobilized fungal laccase. Appl. Microbiol. Biotechnol. 57, 85–91 (2001).
Ahn, M.Y., Dec, J., Kim, J.E. & Bollag, J.M. Treatment of 2,4-dichlorophenol polluted soil with free and immobilized laccase. J. Environ. Qual. 31, 1509–1515 (2002).
Ullah, M.A., Bedford, C.T. & Evans, C.S. Reactions of pentachlorophenol with laccase from Coriolus versicolor. Appl. Microbiol. Biotechnol. 53, 230–234 (2000).
Niwa, T., Doi, U., Kato, Y. & Osawa, T. Inhibitory mechanism of sinapinic acid against peroxynitrite-mediated tyrosine nitration of protein in vitro. FEBS Lett. 459, 43–46 (1999).
Li, L. & Steffens, J.C. Overexpression of polyphenol oxidase in transgenic tomato plants results in enhanced bacterial disease resistance. Planta 215, 239–247 (2002).
Ranocha, P. et al. Laccase down-regulation causes alterations in phenolic metabolism and cell wall structure in poplar. Plant Physiol. 129, 145–155 (2002).
Chapple, C.C., Vogt, T., Ellis, B.E. & Somerville, C.R. An Arabidopsis mutant defective in the general phenylpropanoid pathway. Plant Cell 4, 1413–1424 (1992).
Lim, E.K. et al. Identification of glucosyltransferase genes involved in sinapate metabolism and lignin synthesis in Arabidopsis. J. Biol. Chem. 276, 4344–4349 (2001).
Bais, H.P., Vepachedu, R., Gilroy, S., Callaway, R.M. & Vivanco, J.M. Allelopathy and exotic plant invasion: from molecules and genes to species interactions. Science 301, 1377–1380 (2003).
Dur´n, N., Rosa, M.A., D'Annibale, A. & Gianfreda, L. Applications of laccases and tyrosinases (phenoloxidases) immobilized on different supports: a review. Enzyme Microb. Technol. 31, 907–931 (2002).
Dhankher, O.P. et al. Engineering tolerance and hyperaccumulation of arsenic in plants by combining arsenate reductase and γ-glutamylcysteine synthetase expression. Nat. Biotechnol. 20, 1140–1145 (2002).
Doty, S.L. et al. Enhanced metabolism of halogenated hydrocarbons in transgenic plants containing mammalian cytochrome P450 2E1. Proc. Natl. Acad. Sci. USA 97, 6287–6291 (2000).
Hannink, N. et al. Phytodetoxification of TNT by transgenic plants expressing a bacterial nitroreductase. Nat. Biotechnol. 19, 1168–1172 (2001).
Song, W.Y. et al. Engineering tolerance and accumulation of lead and cadmium in transgenic plants. Nat. Biotechnol. 21, 914–919 (2003).
Luo, P., Wang, Y.H., Wang, G.D., Essenberg, M. & Chen, X.Y. Molecular cloning and functional identification of (+)-δ-cadinene-8-hydroxylase, a cytochrome P450 monooxygenase (CYP706B1) of cotton sesquiterpene biosynthesis. Plant J. 28, 95–104 (2001).
Tan, X.P. et al. Expression pattern of (+)-δ-cadinene synthase genes and biosynthesis of sesquiterpene aldehydes in plants of Gossypium arboreum L. Planta 210, 644–651 (2000).
Jefferson, R.A. Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol. Biol. Rep. 5, 387–405 (1987).
Min, K.L., Kim, Y.H., Kim, Y.W., Jung, H.S. & Hah, Y.C. Characterization of a novel laccase produced by the wood-rotting fungus Phellinus ribis. Arch. Biochem. Biophys. 392, 279–286 (2001).
Howles, P.A. et al. Overexpression of L-phenylalanine ammonia-lyase in transgenic tobacco plants reveals control points for flux into phenylpropanoid biosynthesis. Plant Physiol. 112, 1617–1624 (1996).
Acknowledgements
We thank J. Chen, W.L. Hu and Y.J. Lai for their help in HPLC-MS and GC-MS analysis. This research was supported by National Natural Sciences Foundation of China (grants 30030020 and 39925005) and by the Chinese Academy of Sciences (grant KSCX2-SW-313).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
GUS staining of the 4-day-old Arabidopsis seedlings expressing 35S::GUS or pLAC1::GUS. (PDF 140 kb)
Supplementary Fig. 2
Resistance of Arabidopsis plants of different transgenic LAC lines to TCP, 3 weeks after the second spraying. See also Figure 3. (PDF 182 kb)
Supplementary Fig. 3
HPLC-MS analysis of soluble phenolics of the 2-week-old WT and LAC 4-2 seedlings of Arabidopsis cultured in 1/2 MS medium (a) or in the medium containing 0.5 mM sinapic acid (b). (PDF 15 kb)
Supplementary Table 1
Root elongation of WT and LAC 4-2 seedlings grown in the agar plate in the presence of TCP (PDF 18 kb)
Supplementary Table 2
Root elongation of WT and LAC seedlings grown in the agar plate in the presence of 20 μM TCP (PDF 16 kb)
Rights and permissions
About this article
Cite this article
Wang, GD., Li, QJ., Luo, B. et al. Ex planta phytoremediation of trichlorophenol and phenolic allelochemicals via an engineered secretory laccase. Nat Biotechnol 22, 893–897 (2004). https://doi.org/10.1038/nbt982
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt982
This article is cited by
-
Site suitability analysis for phytoremediation implementation: A case study of Barjora and Durgapur Industrial areas, West Bengal, India
Environment, Development and Sustainability (2023)
-
A comprehensive overview of cotton genomics, biotechnology and molecular biological studies
Science China Life Sciences (2023)
-
Structural basis for monolignol oxidation by a maize laccase
Nature Plants (2020)
-
Rapid dehydration of grape berries dampens the post-ripening transcriptomic program and the metabolite profile evolution
Horticulture Research (2020)
-
Laccases: structure, function, and potential application in water bioremediation
Microbial Cell Factories (2019)