Abstract
Membrane association with mother centriole (M-centriole) distal appendages is critical for ciliogenesis initiation. How the Rab GTPase Rab11–Rab8 cascade functions in early ciliary membrane assembly is unknown. Here, we show that the membrane shaping proteins EHD1 and EHD3, in association with the Rab11–Rab8 cascade, function in early ciliogenesis. EHD1 and EHD3 localize to preciliary membranes and the ciliary pocket. EHD-dependent membrane tubulation is essential for ciliary vesicle formation from smaller distal appendage vesicles (DAVs). Importantly, this step functions in M-centriole to basal body transformation and recruitment of transition zone proteins and IFT20. SNAP29, a SNARE membrane fusion regulator and EHD1-binding protein, is also required for DAV-mediated ciliary vesicle assembly. Interestingly, only after ciliary vesicle assembly is Rab8 activated for ciliary growth. Our studies uncover molecular mechanisms informing a previously uncharacterized ciliogenesis step, whereby EHD1 and EHD3 reorganize the M-centriole and associated DAVs before coordinated ciliary membrane and axoneme growth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
12 March 2015
In the version of this Article originally published Fig. 3g was incorrectly labelled. The top right panel should have been labelled 'EHD1' in red and the entire of 3g should have been labelled 'Rab8a+8b siRNA' in black. The image is corrected in all online versions of the Article.
References
Goetz, S. C. & Anderson, K. V. The primary cilium: a signalling centre during vertebrate development. Nat. Rev. Genet. 11, 331–344 (2010).
Hildebrandt, F., Benzing, T. & Katsanis, N. Ciliopathies. N. Engl. J. Med. 364, 1533–1543 (2011).
Graser, S. et al. Cep164, a novel centriole appendage protein required for primary cilium formation. J. Cell Biol. 179, 321–330 (2007).
Ishikawa, H., Kubo, A., Tsukita, S. & Tsukita, S. Odf2-deficient mother centrioles lack distal/subdistal appendages and the ability to generate primary cilia. Nat. Cell Biol. 7, 517–524 (2005).
Schmidt, K. N. et al. Cep164 mediates vesicular docking to the mother centriole during early steps of ciliogenesis. J. Cell Biol. 199, 1083–1101 (2012).
Joo, K. et al. CCDC41 is required for ciliary vesicle docking to the mother centriole. Proc. Natl Acad. Sci. USA 110, 5987–5992 (2013).
Sillibourne, J. E. et al. Primary ciliogenesis requires the distal appendage component Cep123. Biol. Open 2, 535–545 (2013).
Tanos, B. E. et al. Centriole distal appendages promote membrane docking, leading to cilia initiation. Genes Dev. 27, 163–168 (2013).
Kobayashi, T., Kim, S., Lin, Y. C., Inoue, T. & Dynlacht, B. D. The CP110-interacting proteins Talpid3 and Cep290 play overlapping and distinct roles in cilia assembly. J. Cell Biol. 204, 215–229 (2014).
Sorokin, S. Centrioles and the formation of rudimentary cilia by fibroblasts and smooth muscle cells. J. Cell Biol. 15, 363–377 (1962).
Ye, X., Zeng, H., Ning, G., Reiter, J. F. & Liu, A. C2cd3 is critical for centriolar distal appendage assembly and ciliary vesicle docking in mammals. Proc. Natl Acad. Sci. USA 111, 2164–2169 (2014).
Nachury, M. V. et al. A core complex of BBS proteins cooperates with the GTPase Rab8 to promote ciliary membrane biogenesis. Cell 129, 1201–1213 (2007).
Yoshimura, S., Egerer, J., Fuchs, E., Haas, A. K. & Barr, F. A. Functional dissection of Rab GTPases involved in primary cilium formation. J. Cell Biol. 178, 363–369 (2007).
Knodler, A. et al. Coordination of Rab8 and Rab11 in primary ciliogenesis. Proc. Natl Acad. Sci. USA 107, 6346–6351 (2010).
Westlake, C. J. et al. Primary cilia membrane assembly is initiated by Rab11 and transport protein particle II (TRAPPII) complex-dependent trafficking of Rabin8 to the centrosome. Proc. Natl Acad. Sci. USA 108, 2759–2764 (2011).
Sato, T. et al. Rab8a and Rab8b are essential for multiple apical transport pathways but insufficient for ciliogenesis. J. Cell Sci. 127, 422–431 (2014).
Bryant, D. M. et al. A molecular network for de novo generation of the apical surface and lumen. Nat. Cell Biol. 12, 1035–1045 (2010).
Sorokin, S. P. Reconstructions of centriole formation and ciliogenesis in mammalian lungs. J. Cell Sci. 3, 207–230 (1968).
Zhang, J., Naslavsky, N. & Caplan, S. Rabs and EHDs: alternate modes for traffic control. Biosci. Rep. 32, 17–23 (2012).
Naslavsky, N. & Caplan, S. EHD proteins: key conductors of endocytic transport. Trends Cell Biol. 21, 122–131 (2011).
Naslavsky, N., Rahajeng, J., Sharma, M., Jovic, M. & Caplan, S. Interactions between EHD proteins and Rab11-FIP2: a role for EHD3 in early endosomal transport. Mol. Biol. Cell 17, 163–177 (2006).
Roland, J. T., Kenworthy, A. K., Peranen, J., Caplan, S. & Goldenring, J. R. Myosin Vb interacts with Rab8a on a tubular network containing EHD1 and EHD3. Mol. Biol. Cell 18, 2828–2837 (2007).
Sharma, M., Giridharan, S. S., Rahajeng, J., Naslavsky, N. & Caplan, S. MICAL-L1 links EHD1 to tubular recycling endosomes and regulates receptor recycling. Mol. Biol. Cell 20, 5181–5194 (2009).
Cai, B. et al. Differential roles of C-terminal Eps15 homology domain proteins as vesiculators and tubulators of recycling endosomes. J. Biol. Chem. 288, 30172–30180 (2013).
Giridharan, S. S., Cai, B., Vitale, N., Naslavsky, N. & Caplan, S. Cooperation of MICAL-L1, syndapin2, and phosphatidic acid in tubular recycling endosome biogenesis. Mol. Biol. Cell 24, 1776–1790 (2013).
Rotem-Yehudar, R., Galperin, E. & Horowitz, M. Association of insulin-like growth factor 1 receptor with EHD1 and SNAP29. J. Biol. Chem. 276, 33054–33060 (2001).
Shiba, D. et al. Localization of Inv in a distinctive intraciliary compartment requires the C-terminal ninein-homolog-containing region. J. Cell Sci. 122, 44–54 (2009).
Molla-Herman, A. et al. The ciliary pocket: an endocytic membrane domain at the base of primary and motile cilia. J. Cell Sci. 123, 1785–1795 (2010).
Sonnen, K. F., Schermelleh, L., Leonhardt, H. & Nigg, E. A. 3D-structured illumination microscopy provides novel insight into architecture of human centrosomes. Biol. Open 1, 965–976 (2012).
Sedmak, T. & Wolfrum, U. Intraflagellar transport proteins in ciliogenesis of photoreceptor cells. Biol. Cell 103, 449–466 (2011).
Follit, J. A., Tuft, R. A., Fogarty, K. E. & Pazour, G. J. The intraflagellar transport protein IFT20 is associated with the Golgi complex and is required for cilia assembly. Mol. Biol. Cell 17, 3781–3792 (2006).
Spektor, A., Tsang, W. Y., Khoo, D. & Dynlacht, B. D. Cep97 and CP110 suppress a cilia assembly program. Cell 130, 678–690 (2007).
Naslavsky, N., Boehm, M., Backlund, P. S. Jr & Caplan, S. Rabenosyn-5 and EHD1 interact and sequentially regulate protein recycling to the plasma membrane. Mol. Biol. Cell 15, 2410–2422 (2004).
Naslavsky, N., Rahajeng, J., Chenavas, S., Sorgen, P. L. & Caplan, S. EHD1 and Eps15 interact with phosphatidylinositols via their Eps15 homology domains. J. Biol. Chem. 282, 16612–16622 (2007).
Rainey, M. A. et al. The endocytic recycling regulator EHD1 is essential for spermatogenesis and male fertility in mice. BMC Dev. Biol. 10, 37 (2010).
George, M. et al. Renal thrombotic microangiopathy in mice with combined deletion of endocytic recycling regulators EHD3 and EHD4. PLoS ONE 6, e17838 (2011).
Galperin, E. et al. EHD3: a protein that resides in recycling tubular and vesicular membrane structures and interacts with EHD1. Traffic 3, 575–589 (2002).
De Beer, T. et al. Molecular mechanism of NPF recognition by EH domains. Nat. Struct. Biol. 7, 1018–1022 (2000).
George, M. et al. Shared as well as distinct roles of EHD proteins revealed by biochemical and functional comparisons in mammalian cells and C. elegans. BMC Cell Biol. 8, 3 (2007).
Hehnly, H., Chen, C. T., Powers, C. M., Liu, H. L. & Doxsey, S. The centrosome regulates the Rab11- dependent recycling endosome pathway at appendages of the mother centriole. Curr. Biol. 22, 1944–1950 (2012).
Sprecher, E. et al. A mutation in SNAP29, coding for a SNARE protein involved in intracellular trafficking, causes a novel neurocutaneous syndrome characterized by cerebral dysgenesis, neuropathy, ichthyosis, and palmoplantar keratoderma. Am. J. Hum. Genet. 77, 242–251 (2005).
Fuchs-Telem, D. et al. CEDNIK syndrome results from loss-of-function mutations in SNAP29. Br. J. Dermatol. 164, 610–616 (2011).
Tsang, W. Y. et al. CP110 suppresses primary cilia formation through its interaction with CEP290, a protein deficient in human ciliary disease. Dev. Cell 15, 187–197 (2008).
Li, J. et al. USP33 regulates centrosome biogenesis via deubiquitination of the centriolar protein CP110. Nature 495, 255–259 (2013).
Cajanek, L. & Nigg, E. A. Cep164 triggers ciliogenesis by recruiting Tau tubulin kinase 2 to the mother centriole. Proc. Natl Acad. Sci. USA 111, E2841–E2850 (2014).
Goetz, S. C., Liem, K. F. Jr & Anderson, K. V. The spinocerebellar ataxia-associated gene Tau tubulin kinase 2 controls the initiation of ciliogenesis. Cell 151, 847–858 (2012).
Clement, C. A. et al. TGF-beta signaling is associated with endocytosis at the pocket region of the primary cilium. Cell Rep. 3, 1806–1814 (2013).
Klinger, M. et al. The novel centriolar satellite protein SSX2IP targets Cep290 to the ciliary transition zone. Mol. Biol. Cell 25, 495–507 (2014).
Cheeseman, I. M. & Desai, A. A combined approach for the localization and tandem affinity purification of protein complexes from metazoans. Sci. STKE 2005, pl1 (2005).
Gray, D. C. et al. pHUSH: a single vector system for conditional gene expression. BMC Biotechnol. 7, 61 (2007).
Murone, M., Rosenthal, A. & de Sauvage, F. J. Sonic hedgehog signaling by the patched-smoothened receptor complex. Curr. Biol. 9, 76–84 (1999).
Wright, K. J. et al. An ARL3-UNC119-RP2 GTPase cycle targets myristoylated NPHP3 to the primary cilium. Genes Dev. 25, 2347–2360 (2011).
Jaiswal, B. S. et al. Combined targeting of BRAF and CRAF or BRAF and PI3K effector pathways is required for efficacy in NRAS mutant tumors. PLoS ONE 4, e5717 (2009).
Insinna, C., Pathak, N., Perkins, B., Drummond, I. & Besharse, J. C. The homodimeric kinesin, Kif17, is essential for vertebrate photoreceptor sensory outer segment development. Dev. Biol. 316, 160–170 (2008).
Malicki, J., Avanesov, A., Li, J., Yuan, S. & Sun, Z. Analysis of cilia structure and function in zebrafish. Methods Cell Biol. 101, 39–74 (2011).
Lu, Q. et al. Chromatin-bound NLS proteins recruit membrane vesicles and nucleoporins for nuclear envelope assembly via importin-α/β. Cell Res. 22, 1562–1575 (2012).
Omori, Y. et al. Elipsa is an early determinant of ciliogenesis that links the IFT particle to membrane-associated small GTPase Rab8. Nat. Cell Biol. 10, 437–444 (2008).
Thisse, C. & Thisse, B. High-resolution in situ hybridization to whole-mount zebrafish embryos. Nat. Protoc. 3, 59–69 (2008).
Acknowledgements
We thank K. Nagashima and A. Kamata for help with IEM and EM, C. Lamont for data analysis, S. Lockett for help with SIM imaging, S. Specht for help with cell culture, A. Peden for SNAP29 antibodies, J. Goldenring for Rab11 antibodies, Z. Sun for the transgenic line and S. Burgess for assistance raising zebrafish embryos. We are grateful to J. Donaldson for critical reading of this manuscript. This research was supported by the Intramural Research Program of the National Institutes of Health, National Cancer Institute, Center for Cancer Research. This project has also been funded in part with federal funds from the Frederick National Laboratory for Cancer Research, National Institutes of Health, under contract HHSN261200800001E.
Author information
Authors and Affiliations
Contributions
Q.L. and C.I. carried out most experiments with help from C.J.W. (RNAi, fluorescence imaging and PCR with reverse transcription), C.O. (SIM imaging), P.A.P., S.L., I.O.D. and Y-S.H. (in situ expression in zebrafish), U.B. (EM and CLEM), V.W. (immunoblotting), J.S. (zebrafish MO experiments) and A.C. (cell line generation and RNAi rescue experiments). S.C. and J.R. provided reagents. S.C., C.O., J.L-S., and P.K.J. discussed the results and commented on the manuscript; C.J.W. and C.I. wrote the paper with suggestions from Q.L. and C.O. C.J.W., Q.L. and C.I. conceived and designed the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
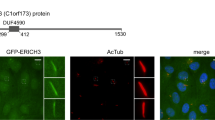
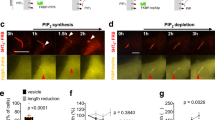
Supplementary Figure 1 EHD1 and EHD3 functions in ciliogenesis and localizes in the ciliary pocket.
a. EHD transcripts were reduced by > 90% as determined by qRT-PCR using two different siRNA oligonucleotides (#1 and #2) for each EHD family member. Averages of three technical replicates from n = 1 experiment are shown. b,c. Quantification of ciliation in RPE cells treated with siRNA as described in Fig 1b. b. Mean ± SEM from n = 3 independent experiments (> 300 cells per treatment) is shown. Two tailed t-test analysis compared with siControl. c. Pooled averages across a single experiment. d. Representative images of RPE cells quantified in Fig 1c. Cells were treated for 6h with siEHD1#1, transiently transfected with siRNA resistant (Res) GFP-EHD proteins or GFP and after 48h cells were serum starved for 24h. Cells were stained with Actub and pericentrin (PCNT) antibodies and imaged by epifluorescence microscopy. e. Quantification of IFT20 fluorescence intensity at the M-centriole in cells treated as described in (d). IFT20 localization was determined as described in Fig 5g. Mean ± SEM from n = 3 independent experiments is shown (total cells counted across all experiments: GFP, 34 cells; GFP-EHD1, 35 cells, GFP-EHD3, 52 cells). Two tailed t-test analysis compared with GFP. f. Quantification of RPGRIP1L fluorescence intensity at the transition zone in cells treated as described in (d). RPGRIP1L transition zone (TZ) localization was determined as described in Fig. 5b. Mean ± SEM from n = 3 independent experiments is shown (total cells counted across all experiments: GFP, 28 cells; GFP-EHD1, 36 cells, GFP-EHD3, 37 cells). Two-tailed t-test analysis compared with GFP. g. Representative image of RPE cells transiently expressing GFP-EHD3 and Smo-tRFP, serum starved for 24h, stained with anti-EHD1 and imaged by epifluorescence microscopy. Scale bar: 2.5 μm. h. Representative image of RPE cells stably expressing GFP-Rab8a, serum starved for 24h and stained with anti-EHD1 and anti-Actub antibodies. Scale bar: 5 μm. i. Representative images of IMCD3 cells transiently expressing GFP-EHD1 (top panel) or GFP-EHD3 (bottom panels), serum starved for 24h and stained with anti-Actub antibody and DAPI and imaged by epifluorescence microscopy. Scale bar: 5 μm. j. Representative images from correlative light and electron microscope (CLEM) imaging of RPE GFP-EHD1 cells transiently expressing tRFP-Rab8a. Following fixation cells were imaged by spinning disk confocal imaging and processed for electron microscopy. Region of interest with cells from fluorescence imaging were identified in electron microscopy sections. The same cilium detected by electron microscopy and fluorescence microscopy (dotted box) is shown on the right panels. n: nuclei. Scale bars: 2 μm. k,l. Representative SIM images of a RPE GFP-EHD1 cells transiently expressing tRFP-Rab8a (k) and tdTomato-Inversin (tdTom-Inversin) (l) for 72h with serum starvation for 24h. Scale bar: 500 nm. ∗ P < 0.05,∗∗∗ P < 0.0001. Statistics source data for Supplemental Fig 1b,e,f can be found in Supplementary Table 2.
Supplementary Figure 2 Expression of Ehd1 and Ehd3 in zebrafish.
a. Western blot analysis of lysates from uninjected embryos or embryos injected with ehd1a + b (left panel) or ehd3 (right panel) MO with and without human EHD1 or EHD3 mRNA. Knockdown efficiency and rescue protein expression levels from western blots band densities were quantified and shown in bar graphs. Note that in these representative rescue experiments, embryos co-injected with ehd1MO and hEHD3 mRNA had < 20% Ehd1 protein left and overexpressed EHD3 at > 15 fold. Similarly, embryos co-injected with ehd3MO and hEHD1 mRNA had < 10% Ehd3 protein left and overexpressed EHD1 at > 8 fold. Un-cropped images are shown in Supplemental Fig 6. b. Quantification of kinocilia number in neuromasts of ehd1 and ehd3 morphants. Note that ehd3 morphants present a partial neuromast defect with an average of 2.5 ± 1.8% kinocilia per neuromast versus 0.9 ± 1.3% in ehd1 morphants. Averages ± SD are pooled data from 3 independent experiments from control n = 19 neuromasts, ehd1MO n = 15 neuromasts, and ehd3 MO n = 16 neuromasts. Two tailed t-test analysis, ∗∗∗ P < 0.0001. Statistics source data for Supplemental Fig. 2b can be found in Supplementary Table 2. c. Standard whole mount in situ hybridizations performed at 48hpf and 72hpf with ehd1 and ehd3 probes designed for the 3’UTR regions. The ehd1 probe was predicted to recognize both ehd1a and ehd1b transcripts. ehd1 mRNA expression was detected prominently in the eye and otic vesicles (OV) at 48hr and 72hpf whereas ehd3 mRNA was mainly detected in the eye at 48 and 72hpf. Insets show 48 hpf expression in OV region. d. Sections of retinae from 4dpf control or MO injected embryos stained with anti-Ehd1 or anti-Ehd3 antibodies and DAPI. Uninjected and MO injected retinae sections were stained at the same time demonstrating specific expression of Ehd1 and Ehd3 in the retinae. Scale bar: 10 μm. e. Representatives images of otic vesicles in MO injected embryos at 3dpf stained with anti-Actub and anti-Ehd1 or anti-Ehd3 antibodies as described in (d). Scale bar: 10 μm.
Supplementary Figure 3 GFP-EHD1 and tRFP-Rab8 localizations in RPE cells following serum starvation.
Full cell images of RPE GFP-EHD1 cells transiently expressing tRFP-Rab8a after serum starvation shown in Fig 3c. Cropped regions were shown in square boxes and the nuclear region is marked by blue dotted lines. Scale bar: 10 μm.
Supplementary Figure 4 Representative electron micrographs of M-centrioles from RPE cells and zebrafish photoreceptors.
a. Additional representative TEM images of RPE cells treated with siControl in presence of serum, or siEHD1 (upper panel) or siRab8 (lower panel) after 24h starvation as shown in Fig 4a and quantified in Fig 4b. Scale bar: 250 nm. b. Additional representative TEM images of zebrafish photoreceptor cells treated with ehd1 + 3 MO and Rab8 MO as described in Fig 4g and quantified in Fig 4h. Scale bar: 250 nm.
Supplementary Figure 5 B9D2-GFP is recruited to the M-centriole after the ciliary membrane receptor 5-HT6-tRFP in RPE cells following serum starvation.
RPE B9D2-GFP cells transiently expressing 5HT6-tRFP, serum starved for 1h and imaged live by spinning disk confocal microscopy. Images show single xy-planes from z-stacks. Scale bar: 2 μm.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1577 kb)
Supplementary Table 1
Supplementary Information (XLS 29 kb)
Supplementary Table 2
Supplementary Information (XLSX 23 kb)
GFP-EHD1 localizes to developing ciliary membranes prior to tRFP-Rab8a.
Time-lapse video of a RPE cell expressing GFP-EHD1 and tRFP-Rab8a. Spinning disk confocal images were taken every 5 min after 1hr starvation. (MOV 1351 kb)
GFP-EHD1 localizes to the developing ciliary membrane with Smo-tRFP.
Time-lapse video of a RPE cell expressing GFP-EHD1 and Smo-tRFP. Spinning disk confocal images were taken every 10 min after 1hr starvation. (MOV 440 kb)
GFP-Rab8 is recruited to the developing ciliary membrane after Smo-tRFP.
Time-lapse video of a RPE cell expressing GFP-Rab8 and Smo-tRFP. Spinning disk confocal images were taken every 10 min after 1hr starvation. (MOV 125 kb)
IFT20-GFP is recruited after pre-ciliary membrane accumulation of Smo-tRFP.
Time-lapse series showing a RPE cell expressing IFT20-GFP and Smo-tRFP. Spinning disk confocal images were captured every 10 min after 1hr starvation. (MOV 262 kb)
IFT20-tRFP localizes to developing cilia prior to GFP-Rab8a recruitment.
Time-lapse video of a RPE cell expressing GFP-Rab8 and IFT20-tRFP. Spinning disk confocal images were captured every 20 min after 1hr starvation. (MOV 310 kb)
Rights and permissions
About this article
Cite this article
Lu, Q., Insinna, C., Ott, C. et al. Early steps in primary cilium assembly require EHD1/EHD3-dependent ciliary vesicle formation. Nat Cell Biol 17, 228–240 (2015). https://doi.org/10.1038/ncb3109
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3109
This article is cited by
-
Appearing and disappearing acts of cilia
Journal of Biosciences (2023)
-
Potassium Channel KCNH1 Activating Variants Cause Altered Functional and Morphological Ciliogenesis
Molecular Neurobiology (2022)
-
FERARI is required for Rab11-dependent endocytic recycling
Nature Cell Biology (2020)
-
Freeing the brake: Proliferation needs primary cilium to disassemble
Journal of Biosciences (2020)
-
Rho GTPases in cancer radiotherapy and metastasis
Cancer and Metastasis Reviews (2020)