Abstract
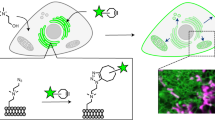
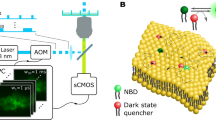
Membrane lipids are dynamic molecules that play important roles in cell signalling and regulation, but an in situ imaging method for quantitatively tracking lipids in living cells is lacking at present. Here, we report a new chemical method of quantitative lipid imaging using sensors engineered by labelling proteins with an environmentally sensitive fluorophore. A prototype sensor for phosphatidylinositol-4,5-bisphosphate (PtdIns(4,5)P2)—a key signalling lipid in diverse cellular processes—was generated by covalently attaching a single 2-dimethylamino-6-acyl-naphthalene group to the N-terminal α-helix of the engineered epsin1 ENTH domain, a protein that selectively binds PtdIns(4,5)P2. The sensor allows robust and sensitive in situ quantitative imaging in mammalian cells, providing new insight into the spatiotemporal dynamics and fluctuation of this key signalling lipid. Application of the sensor to immune cells reveals the presence of a local threshold PtdIns(4,5)P2 concentration required for triggering phagocytosis. This sensor strategy is generally applicable to in situ quantification of other cellular lipids.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hannun, Y. A. & Obeid, L. M. Principles of bioactive lipid signalling: lessons from sphingolipids. Nat. Rev. Mol. Cell. Biol. 9, 139–150 (2008).
Cho, W. Building signaling complexes at the membrane. Sci STKE 2006, pe7 (2006).
Wymann, M. P. & Schneiter, R. Lipid signalling in disease. Nat. Rev. Mol. Cell. Biol. 9, 162–176 (2008).
van Meer, G., Voelker, D. R. & Feigenson, G. W. Membrane lipids: where they are and how they behave. Nat. Rev. Mol. Cell. Biol. 9, 112–124 (2008).
Cho, W. & Stahelin, R. V. Membrane–protein interactions in cell signaling and membrane trafficking. Annu. Rev. Biophys. Biomol. Struct. 34, 119–151 (2005).
Lemmon, M. A. Membrane recognition by phospholipid-binding domains. Nat. Rev. Mol. Cell. Biol. 9, 99–111 (2008).
Varnai, P. & Balla, T. Live cell imaging of phosphoinositide dynamics with fluorescent protein domains. Biochim. Biophys. Acta 1761, 957–967 (2006).
Downes, C. P., Gray, A. & Lucocq, J. M. Probing phosphoinositide functions in signaling and membrane trafficking. Trends Cell. Biol. 15, 259–268 (2005).
Irvine, R. Inositol lipids: to PHix or not to PHix? Curr. Biol. 14, R308–310 (2004).
Han, X. & Gross, R. W. Global analyses of cellular lipidomes directly from crude extracts of biological samples by ESI mass spectrometry: a bridge to lipidomics. J. Lipid Res. 44, 1071–1079 (2003).
van Meer, G. Cellular lipidomics. EMBO J. 24, 3159–3165 (2005).
Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, 651–657 (2006).
McLaughlin, S. & Murray, D. Plasma membrane phosphoinositide organization by protein electrostatics. Nature 438, 605–611 (2005).
McLaughlin, S., Wang, J., Gambhir, A. & Murray, D. PIP(2) and proteins: interactions, organization, and information flow. Annu. Rev. Biophys. Biomol. Struct. 31, 151–175 (2002).
van Rheenen, J., Achame, E. M., Janssen, H., Calafat, J. & Jalink, K. PIP2 signaling in lipid domains: a critical re-evaluation. EMBO J. 24, 1664–1673 (2005).
Hilgemann, D. W. Local PIP(2) signals: when, where, and how? Pflugers Arch. 455, 55–67 (2007).
Botelho, R. J. et al. Localized biphasic changes in phosphatidylinositol-4,5-bisphosphate at sites of phagocytosis. J. Cell. Biol. 151, 1353–1368 (2000).
Garrenton, L. S., Stefan, C. J., McMurray, M. A., Emr, S. D. & Thorner, J. Pheromone-induced anisotropy in yeast plasma membrane phosphatidylinositol-4,5-bisphosphate distribution is required for MAPK signaling. Proc. Natl Acad. Sci. USA 107, 11805–11810 (2010).
Itoh, T. et al. Role of the ENTH domain in phosphatidylinositol-4,5-bisphosphate binding and endocytosis. Science 291, 1047–1051 (2001).
Ford, M. G. et al. Curvature of clathrin-coated pits driven by epsin. Nature 419, 361–366 (2002).
Stahelin, R. V. et al. Contrasting membrane interaction mechanisms of AP180 N-terminal homology (ANTH) and epsin N-terminal homology (ENTH) domains. J. Biol. Chem. 278, 28993–28999 (2003).
Weber, G. & Farris, F. J. Synthesis and spectral properties of a hydrophobic fluorescent probe: 6-propionyl-2-(dimethylamino)naphthalene. Biochemistry 18, 3075–3078 (1979).
Terebiznik, M. R. et al. Elimination of host cell PtdIns(4,5)P(2) by bacterial SigD promotes membrane fission during invasion by Salmonella. Nature Cell Biol. 4, 766–773 (2002).
Ruderman, N. B., Kapeller, R., White, M. F. & Cantley, L. C. Activation of phosphatidylinositol 3-kinase by insulin. Proc. Natl Acad. Sci. USA 87, 1411–1415 (1990).
Rhee, S. G. Regulation of phosphoinositide-specific phospholipase C. Annu. Rev. Biochem. 70, 281–312 (2001).
Inoue, T., Heo, W. D., Grimley, J. S., Wandless, T. J. & Meyer, T. An inducible translocation strategy to rapidly activate and inhibit small GTPase signaling pathways. Nat. Methods 2, 415–418 (2005).
Varnai, P., Thyagarajan, B., Rohacs, T. & Balla, T. Rapidly inducible changes in phosphatidylinositol 4,5-bisphosphate levels influence multiple regulatory functions of the lipid in intact living cells. J. Cell. Biol. 175, 377–382 (2006).
Swanson, J. A. Shaping cups into phagosomes and macropinosomes. Nat. Rev. Mol. Cell. Biol. 9, 639–649 (2008).
Grinstein, S. Imaging signal transduction during phagocytosis: phospholipids, surface charge, and electrostatic interactions. Am. J. Physiol. Cell. Physiol. 299, C876–C881 (2010).
Stahelin, R. V. et al. Mechanism of diacylglycerol-induced membrane targeting and activation of protein kinase Cδ. J. Biol. Chem. 279, 29501–29512 (2004).
Manna, D. et al. Differential roles of phosphatidylserine, PtdIns(4,5)P2, and PtdIns(3,4,5)P3 in plasma membrane targeting of C2 domains. Molecular dynamics simulation, membrane binding, and cell translocation studies of the PKCα C2 domain. J. Biol. Chem. 283, 26047–26058 (2008).
Grynkiewicz, G., Poenie, M. & Tsien, R. Y. A new generation of Ca2+ indicators with greatly improved fluorescence properties. J. Biol. Chem. 260, 3440–3450 (1985).
Acknowledgements
This study was supported by the National Institutes of Health (grant no. GM68849). It was also supported in part by the Chicago Biomedical Consortium with support from the Searl Funds at the Chicago Community Trust. The authors thank I. Kim, S.-Y. Kim, S. Bhattacharjee, Y. Kanaho and L.-W. Gong for technical assistance in phagocytosis and membrane trafficking experiments, and T. Balla for the generous gift of the PtdIns(4,5)-depletion system.
Author information
Authors and Affiliations
Contributions
W.C. and Y.Y. conceived the lipid sensor strategy. W.C. supervised the project and Y.Y. designed and prepared the sensor. P.J.L. and S.K. performed all microscopy imaging and image analysis. Y.Y. and P.J.L. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 1570 kb)
Supplementary information
Supplementary Movie S1 (AVI 1026 kb)
Rights and permissions
About this article
Cite this article
Yoon, Y., Lee, P., Kurilova, S. et al. In situ quantitative imaging of cellular lipids using molecular sensors. Nature Chem 3, 868–874 (2011). https://doi.org/10.1038/nchem.1163
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1163
This article is cited by
-
Selective detection of phospholipids using molecularly imprinted fluorescent sensory core-shell particles
Scientific Reports (2020)
-
Investigation of the phosphatidylserine binding properties of the lipid biosensor, Lactadherin C2 (LactC2), in different membrane environments
Journal of Bioenergetics and Biomembranes (2018)
-
Orthogonal lipid sensors identify transbilayer asymmetry of plasma membrane cholesterol
Nature Chemical Biology (2017)
-
Required hydrophobicity of fluorescent reporters for phosphatidylinositol family of lipid enzymes
Analytical and Bioanalytical Chemistry (2017)
-
Cholesterol modulates cell signaling and protein networking by specifically interacting with PDZ domain-containing scaffold proteins
Nature Communications (2012)