Abstract
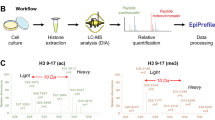
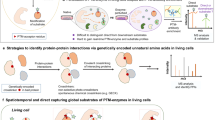
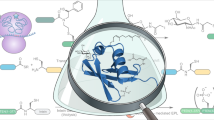
Post-translational modifications (PTMs) have key roles in regulating protein-protein interactions in living cells. However, it remains a challenge to identify these PTM-mediated interactions. Here we develop a new lysine-based photo-reactive amino acid, termed photo-lysine. We demonstrate that photo-lysine, which is readily incorporated into proteins by native mammalian translation machinery, can be used to capture and identify proteins that recognize lysine PTMs, including 'readers' and 'erasers' of histone modifications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Walsh, C.T. Posttranslational Modifications of Proteins: Expanding Nature's Inventory (Roberts and Co. Publishers, Greenwood Village, Colorado, USA, 2006).
Pham, N.D., Parker, R.B. & Kohler, J.J. Curr. Opin. Chem. Biol. 17, 90–101 (2013).
Suchanek, M., Radzikowska, A. & Thiele, C. Nat. Methods 2, 261–267 (2005).
Lan, F. et al. Nature 448, 718–722 (2007).
Musrati, R.A., Kollárová, M., Mernik, N. & Mikulásová, D. Gen. Physiol. Biophys. 17, 193–210 (1998).
Nemoto, T., Ohara-Nemoto, Y., Ota, M., Takagi, T. & Yokoyama, K. Eur. J. Biochem. 233, 1–8 (1995).
Bukau, B. & Horwich, A.L. Cell 92, 351–366 (1998).
Walker, J.R., Corpina, R.A. & Goldberg, J. Nature 412, 607–614 (2001).
Roth, S.Y., Denu, J.M. & Allis, C.D. Annu. Rev. Biochem. 70, 81–120 (2001).
Wang, H. et al. Mol. Cell 8, 1207–1217 (2001).
Nishioka, K. et al. Genes Dev. 16, 479–489 (2002).
Houtkooper, R.H., Pirinen, E. & Auwerx, J. Nat. Rev. Mol. Cell Biol. 13, 225–238 (2012).
Li, Y. et al. Cell 159, 558–571 (2014).
Tippmann, E.M., Liu, W., Summerer, D., Mack, A.V. & Schultz, P.G. ChemBioChem 8, 2210–2214 (2007).
Chou, C.J., Uprety, R., Davis, L., Chin, J.W. & Deiters, A. Chem. Sci. (Camb.) 2, 480–483 (2011).
Zhang, M. et al. Nat. Chem. Biol. 7, 671–677 (2011).
Muir, T.W. Annu. Rev. Biochem. 72, 249–289 (2003).
Chin, J.W. Annu. Rev. Biochem. 83, 379–408 (2014).
Yang, Y.Y., Grammel, M., Raghavan, A.S., Charron, G. & Hang, H.C. Chem. Biol. 17, 1212–1222 (2010).
Du, J. et al. Science 334, 806–809 (2011).
Hubbard, B.P. et al. Science 339, 1216–1219 (2013).
Bao, X. et al. eLife 3, e02999 (2014).
Yang, T.P., Liu, Z. & Li, X.D. Chem. Sci. (Camb.) 6, 1011–1017 (2015).
Filippakopoulos, P. et al. Cell 149, 214–231 (2012).
Couture, J.F., Collazo, E., Hauk, G. & Trievel, R.C. Nat. Struct. Mol. Biol. 13, 140–146 (2006).
Shechter, D., Dormann, H.L., Allis, C.D. & Hake, S.B. Nat. Protoc. 2, 1445–1457 (2007).
Goldberg, D. et al. J. Proteome Res. 6, 3995–4005 (2007).
Acknowledgements
We acknowledge support from the Hong Kong Research Grants Council General Research Fund (GRF 17303114, GRF 17127915), Early Career Scheme (ECS) (HKU 709813P ) and Collaborative Research Fund (CRF C7037-14G). We acknowledge the University of Hong Kong for an e-SRT on Integrative Biology and grants from the Seed Funding Program (201411159101, 201409160027, 201311159007 and 201309176090). We thank X. Li for discussion. We thank E. Verdin (University of California, San Francisco), Q. Hao (The University of Hong Kong), H. Li (Tsinghua University), R.C. Trievel (University of Michigan), K. Guan (Fudan University) and H. Sun (City University of Hong Kong) for providing the plasmids.
Author information
Authors and Affiliations
Contributions
T.Y., X.-M.L. and X.D.L. designed the experiments and wrote the paper. T.Y. synthesized the small-molecule compounds and peptides and performed all of the photo-lysine–based cross-linking experiments in vitro. X.-M.L. performed all of the photo-lysine–based cross-linking experiments in living cells and prepared the MS sample. X.B. performed the ITC experiments. X.B. and Y.M.E.F. conducted the MS-based proteomics experiments. X.-M.L., X.B. and X.D.L. analyzed the MS/MS data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–14 and Supplementary Notes 1–4. (PDF 12956 kb)
Supplementary Table 1
The estimated incorporation rates of photo-Lys at the identified sites. (XLSX 14 kb)
Supplementary Table 2
The estimated incorporation rates of photo-Lys at the identified histone sites. (XLSX 11 kb)
Supplementary Table 3
Summary of the identified histone- and chromatin-binding proteins. (XLSX 17 kb)
Supplementary Data Set 1
Proteins quantified in HeLa S3 cells with/without photo-Lys labeling. (XLSX 222 kb)
Supplementary Data Set 2
Proteins quantified in SILAC experiment to identify histone-/chromatin-binding proteins (XLSX 193 kb)
Rights and permissions
About this article
Cite this article
Yang, T., Li, XM., Bao, X. et al. Photo-lysine captures proteins that bind lysine post-translational modifications. Nat Chem Biol 12, 70–72 (2016). https://doi.org/10.1038/nchembio.1990
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1990
This article is cited by
-
Photo-ANA enables profiling of host–bacteria protein interactions during infection
Nature Chemical Biology (2023)
-
Identification of photocrosslinking peptide ligands by mRNA display
Communications Chemistry (2023)
-
Examining sterically demanding lysine analogs for histone lysine methyltransferase catalysis
Scientific Reports (2020)
-
Contemporary approaches to site-selective protein modification
Nature Reviews Chemistry (2019)
-
The First MS-Cleavable, Photo-Thiol-Reactive Cross-Linker for Protein Structural Studies
Journal of the American Society for Mass Spectrometry (2019)