Abstract
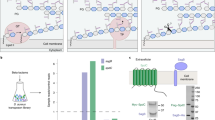
Sacculus is a peptidoglycan (PG) matrix that protects bacteria from osmotic lysis. In Gram-positive organisms, the sacculus is densely functionalized with glycopolymers important for survival, but the way in which assembly occurs is not known. In Staphylococcus aureus, three LCP (LytR-CpsA-Psr) family members have been implicated in attaching the major glycopolymer wall teichoic acid (WTA) to PG, but ligase activity has not been demonstrated for these or any other LCP proteins. Using WTA and PG substrates produced chemoenzymatically, we show that all three proteins can transfer WTA precursors to nascent PGs, establishing that LCP proteins are PG–glycopolymer ligases. Although all S. aureus LCP proteins have the capacity to attach WTA to PG, we show that their cellular functions are not redundant. Strains lacking lcpA have phenotypes similar to those of WTA-null strains, indicating that this is the most important WTA ligase. This work provides a foundation for studying how LCP enzymes participate in cell wall assembly.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Silhavy, T.J., Kahne, D. & Walker, S. The bacterial cell envelope. Cold Spring Harb. Perspect. Biol. 2, a000414 (2010).
Vollmer, W., Blanot, D. & de Pedro, M.A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167 (2008).
Rajagopal, M. & Walker, S. Envelope structures of Gram-positive bacteria. Curr. Top. Microbiol. Immunol. https://dx.doi.org/10.1007/82_2015_5021 (2016).
Neuhaus, F.C. & Baddiley, J. A continuum of anionic charge: structures and functions of d-alanyl-teichoic acids in Gram-positive bacteria. Microbiol. Mol. Biol. Rev. 67, 686–723 (2003).
Weidenmaier, C. & Peschel, A. Teichoic acids and related cell-wall glycopolymers in Gram-positive physiology and host interactions. Nat. Rev. Microbiol. 6, 276–287 (2008).
Atilano, M.L. et al. Teichoic acids are temporal and spatial regulators of peptidoglycan cross-linking in Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 107, 18991–18996 (2010).
Campbell, J. et al. Synthetic lethal compound combinations reveal a fundamental connection between wall teichoic acid and peptidoglycan biosyntheses in Staphylococcus aureus. ACS Chem. Biol. 6, 106–116 (2011).
Weidenmaier, C. et al. Role of teichoic acids in Staphylococcus aureus nasal colonization, a major risk factor in nosocomial infections. Nat. Med. 10, 243–245 (2004).
Brown, S., Santa Maria, J.P. Jr. & Walker, S. Wall teichoic acids of gram-positive bacteria. Annu. Rev. Microbiol. 67, 313–336 (2013).
Farha, M.A. et al. Inhibition of WTA synthesis blocks the cooperative action of PBPs and sensitizes MRSA to β-lactams. ACS Chem. Biol. 8, 226–233 (2013).
Brown, S. et al. Methicillin resistance in Staphylococcus aureus requires glycosylated wall teichoic acids. Proc. Natl. Acad. Sci. USA 109, 18909–18914 (2012).
Sham, L.T. et al. Bacterial cell wall. MurJ is the flippase of lipid-linked precursors for peptidoglycan biogenesis. Science 345, 220–222 (2014).
Ruiz, N. Bioinformatics identification of MurJ (MviN) as the peptidoglycan lipid II flippase in Escherichia coli. Proc. Natl. Acad. Sci. USA 105, 15553–15557 (2008).
Mohammadi, T. et al. Identification of FtsW as a transporter of lipid-linked cell wall precursors across the membrane. EMBO J. 30, 1425–1432 (2011).
Yokoyama, K., Miyashita, T., Araki, Y. & Ito, E. Structure and functions of linkage unit intermediates in the biosynthesis of ribitol teichoic acids in Staphylococcus aureus H and Bacillus subtilis W23. Eur. J. Biochem. 161, 479–489 (1986).
Bera, A. et al. Influence of wall teichoic acid on lysozyme resistance in Staphylococcus aureus. J. Bacteriol. 189, 280–283 (2007).
Schlag, M. et al. Role of staphylococcal wall teichoic acid in targeting the major autolysin Atl. Mol. Microbiol. 75, 864–873 (2010).
Kawai, Y. et al. A widespread family of bacterial cell wall assembly proteins. EMBO J. 30, 4931–4941 (2011).
Dengler, V. et al. Deletion of hypothetical wall teichoic acid ligases in Staphylococcus aureus activates the cell wall stress response. FEMS Microbiol. Lett. 333, 109–120 (2012).
Chan, Y.G., Frankel, M.B., Dengler, V., Schneewind, O. & Missiakas, D. Staphylococcus aureus mutants lacking the LytR-CpsA-Psr family of enzymes release cell wall teichoic acids into the extracellular medium. J. Bacteriol. 195, 4650–4659 (2013).
Chan, Y.G., Kim, H.K., Schneewind, O. & Missiakas, D. The capsular polysaccharide of Staphylococcus aureus is attached to peptidoglycan by the LytR-CpsA-Psr (LCP) family of enzymes. J. Biol. Chem. 289, 15680–15690 (2014).
Liszewski Zilla, M., Chan, Y.G., Lunderberg, J.M., Schneewind, O. & Missiakas, D. LytR-CpsA-Psr enzymes as determinants of Bacillus anthracis secondary cell wall polysaccharide assembly. J. Bacteriol. 197, 343–353 (2015).
Harrison, J. et al. Lcp1 Is a phosphotransferase responsible for ligating arabinogalactan to peptidoglycan in Mycobacterium tuberculosis. MBio 7, e00972–16 (2016).
Over, B. et al. LytR-CpsA-Psr proteins in Staphylococcus aureus display partial functional redundancy and the deletion of all three severely impairs septum placement and cell separation. FEMS Microbiol. Lett. 320, 142–151 (2011).
Brown, S., Meredith, T., Swoboda, J. & Walker, S. Staphylococcus aureus and Bacillus subtilis W23 make polyribitol wall teichoic acids using different enzymatic pathways. Chem. Biol. 17, 1101–1110 (2010).
Brown, S., Zhang, Y.H. & Walker, S. A revised pathway proposed for Staphylococcus aureus wall teichoic acid biosynthesis based on in vitro reconstitution of the intracellular steps. Chem. Biol. 15, 12–21 (2008).
Gale, R.T., Sewell, E.W., Garrett, T.A. & Brown, E.D. Reconstituting poly(glycerol phosphate) wall teichoic acid biosynthesis in vitro using authentic substrates. Chem. Sci. 5, 3823–3830 (2014).
Lee, W. et al. The mechanism of action of lysobactin. J. Am. Chem. Soc. 138, 100–103 (2016).
Ginsberg, C., Zhang, Y.H., Yuan, Y. & Walker, S. In vitro reconstitution of two essential steps in wall teichoic acid biosynthesis. ACS Chem. Biol. 1, 25–28 (2006).
Badurina, D.S., Zolli-Juran, M. & Brown, E.D. CTP: glycerol 3-phosphate cytidylyltransferase (TarD) from Staphylococcus aureus catalyzes the cytidylyl transfer via an ordered Bi-Bi reaction mechanism with micromolar K(m) values. Biochim. Biophys. Acta 1646, 196–206 (2003).
Ye, X.Y. et al. Better substrates for bacterial transglycosylases. J. Am. Chem. Soc. 123, 3155–3156 (2001).
Lebar, M.D. et al. Forming crosslinked peptidoglycan from synthetic gram-negative Lipid II. J. Am. Chem. Soc. 135, 4632–4635 (2013).
Rebets, Y. et al. Moenomycin resistance mutations in Staphylococcus aureus reduce peptidoglycan chain length and cause aberrant cell division. ACS Chem. Biol. 9, 459–467 (2014).
Tipper, D.J., Strominger, J.L. & Ensign, J.C. Structure of the cell wall of Staphylococcus aureus, strain Copenhagen. VII. Mode of action of the bacteriolytic peptidase from Myxobacter and the isolation of intact cell wall polysaccharides. Biochemistry 6, 906–920 (1967).
Perlstein, D.L., Zhang, Y., Wang, T.S., Kahne, D.E. & Walker, S. The direction of glycan chain elongation by peptidoglycan glycosyltransferases. J. Am. Chem. Soc. 129, 12674–12675 (2007).
Eberhardt, A. et al. Attachment of capsular polysaccharide to the cell wall in Streptococcus pneumoniae. Microb. Drug Resist. 18, 240–255 (2012).
Pasquina, L. et al. A synthetic lethal approach for compound and target identification in Staphylococcus aureus. Nat. Chem. Biol. 12, 40–45 (2016).
Percy, M.G. & Gründling, A. Lipoteichoic acid synthesis and function in Gram-positive bacteria. Annu. Rev. Microbiol. 68, 81–100 (2014).
Peschel, A., Vuong, C., Otto, M. & Götz, F. The d-alanine residues of Staphylococcus aureus teichoic acids alter the susceptibility to vancomycin and the activity of autolytic enzymes. Antimicrob. Agents Chemother. 44, 2845–2847 (2000).
Santa Maria, J.P. Jr. et al. Compound–gene interaction mapping reveals distinct roles for Staphylococcus aureus teichoic acids. Proc. Natl. Acad. Sci. USA 111, 12510–12515 (2014).
Suzuki, T. et al. Wall teichoic acid protects Staphylococcus aureus from inhibition by Congo red and other dyes. J. Antimicrob. Chemother. 67, 2143–2151 (2012).
Hübscher, J. et al. MsrR contributes to cell surface characteristics and virulence in Staphylococcus aureus. FEMS Microbiol. Lett. 295, 251–260 (2009).
Hartman, B.J. & Tomasz, A. Low-affinity penicillin-binding protein associated with beta-lactam resistance in Staphylococcus aureus. J. Bacteriol. 158, 513–516 (1984).
Matsuhashi, M. et al. Molecular cloning of the gene of a penicillin-binding protein supposed to cause high resistance to beta-lactam antibiotics in Staphylococcus aureus. J. Bacteriol. 167, 975–980 (1986).
Huang, C.Y. et al. Crystal structure of Staphylococcus aureus transglycosylase in complex with a lipid II analog and elucidation of peptidoglycan synthesis mechanism. Proc. Natl. Acad. Sci. USA 109, 6496–6501 (2012).
Kaito, C. & Sekimizu, K. Colony spreading in Staphylococcus aureus. J. Bacteriol. 189, 2553–2557 (2007).
Kato, F. & Sugai, M. A simple method of markerless gene deletion in Staphylococcus aureus. J. Microbiol. Methods 87, 76–81 (2011).
Chen, L. et al. Intrinsic lipid preferences and kinetic mechanism of Escherichia coli MurG. Biochemistry 41, 6824–6833 (2002).
Tsukamoto, H. & Kahne, D. N-methylimidazolium chloride-catalyzed pyrophosphate formation: application to the synthesis of Lipid I and NDP-sugar donors. Bioorg. Med. Chem. Lett. 21, 5050–5053 (2011).
Perlstein, D.L., Wang, T.S., Doud, E.H., Kahne, D. & Walker, S. The role of the substrate lipid in processive glycan polymerization by the peptidoglycan glycosyltransferases. J. Am. Chem. Soc. 132, 48–49 (2010).
Barrett, D. et al. Analysis of glycan polymers produced by peptidoglycan glycosyltransferases. J. Biol. Chem. 282, 31964–31971 (2007).
Wang, T.S. et al. Primer preactivation of peptidoglycan polymerases. J. Am. Chem. Soc. 133, 8528–8530 (2011).
Santiago, M. et al. A new platform for ultra-high density Staphylococcus aureus transposon libraries. BMC Genomics 16, 252 (2015).
Goecks, J., Nekrutenko, A. & Taylor, J. Galaxy Team Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 11, R86 (2010).
Blankenberg, D. et al. Galaxy: a web-based genome analysis tool for experimentalists. Curr. Protoc. Mol. Biol. 89, 19.10:19.10.1–19.10.21 (2010).
Giardine, B. et al. Galaxy: a platform for interactive large-scale genome analysis. Genome Res. 15, 1451–1455 (2005).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Acknowledgements
We would like to thank V. Haunreiter (Department of Molecular and Cellular Biology, Harvard University) for her generous gift of the strains J126, JH100, RH53, RH72, PS60, PS109, and PS47 (refs. 19, 42), S. Trauger and the Bauer Core Facilities for assistance with MS, and the T. Pang (Harvard Medical School, Department of Microbiology and Immunology) and the Bernhardt lab for their gift of the plasmid pTP63. This research was supported by the NIH (P01AI083214 and R01 AI099144 to S.W.; R01 GM066174 to D.K.; R01 GM076710 to D.K. and S.W.).
Author information
Authors and Affiliations
Contributions
S.W., D.K., K.S., and L.M.M. designed experiments. K.S. prepared wall teichoic acid substrates, and peptidoglycan substrates were prepared with help from Y.Q.; L.M.M. performed the Tn-seq experiments; K.S. and L.M.M. prepared the spot dilution assays and the MIC experiments; K.S. and L.M.M. made strains used in this study except those received from others as noted (Supplementary Table 5); K.S. cloned, expressed, and purified all LCP proteins and performed reconstitution experiments; S.W. designed and supervised the project; S.W., D.K., and K.S. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–10 and Supplementary Tables 1–6. (PDF 1433 kb)
Rights and permissions
About this article
Cite this article
Schaefer, K., Matano, L., Qiao, Y. et al. In vitro reconstitution demonstrates the cell wall ligase activity of LCP proteins. Nat Chem Biol 13, 396–401 (2017). https://doi.org/10.1038/nchembio.2302
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2302
This article is cited by
-
The divisome but not the elongasome organizes capsule synthesis in Streptococcus pneumoniae
Nature Communications (2023)
-
PplD is a de-N-acetylase of the cell wall linkage unit of streptococcal rhamnopolysaccharides
Nature Communications (2022)
-
Staphylococcus aureus cell growth and division are regulated by an amidase that trims peptides from uncrosslinked peptidoglycan
Nature Microbiology (2020)
-
Structure and reconstitution of a hydrolase complex that may release peptidoglycan from the membrane after polymerization
Nature Microbiology (2020)
-
Coordination of capsule assembly and cell wall biosynthesis in Staphylococcus aureus
Nature Communications (2019)