Abstract
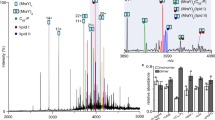
Peptidoglycan is an essential crosslinked polymer that surrounds bacteria and protects them from osmotic lysis. β-lactam antibiotics target the final stages of peptidoglycan biosynthesis by inhibiting the transpeptidases that crosslink glycan strands to complete cell wall assembly. Characterization of transpeptidases and their inhibition by β-lactams have been hampered by lack of access to a suitable substrate. We describe a general approach to accumulate Lipid II in bacteria and to obtain large quantities of this cell wall precursor. We demonstrate the utility of this strategy by isolating Staphylococcus aureus Lipid II and reconstituting the synthesis of crosslinked peptidoglycan by the essential penicillin-binding protein 2 (PBP2), which catalyzes both glycan polymerization and transpeptidation. We also show that we can compare the potencies of different β-lactams by directly monitoring transpeptidase inhibition. The methods reported here will enable a better understanding of cell wall biosynthesis and facilitate studies of next-generation transpeptidase inhibitors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Walsh, C.T. & Wencewicz, T. Antibiotics: Challenges, Mechanisms, Opportunities (ASM press, 2016).
Centers for Disease Control and Prevention Office of Infectious Disease. Antibiotic resistance threat in the United States, 2013 (US Department of Health and Human Services, 2013).
Chung, B.C. et al. Crystal structure of MraY, an essential membrane enzyme for bacterial cell wall synthesis. Science 341, 1012–1016 (2013).
Men, H., Park, P., Ge, M. & Walker, S. Substrate synthesis and activity assay for MurG. J. Am. Chem. Soc. 120, 2484–2485 (1998).
Ha, S. et al. The kinetic characterization of Escherichia coli MurG using synthetic substrate analogues. J. Am. Chem. Soc. 121, 8415–8426 (1999).
Siewert, G. & Strominger, J.L. Biosynthesis of the peptidoglycan of bacterial cell walls. XI. Formation of the isoglutamine amide group in the cell walls of Staphylococcus aureus. J. Biol. Chem. 243, 783–790 (1968).
Münch, D. et al. Identification and in vitro analysis of the GatD/MurT enzyme-complex catalyzing lipid II amidation in Staphylococcus aureus. PLoS Pathog. 8, e1002509 (2012).
Matsuhashi, M., Dietrich, C.P. & Strominger, J.L. Incorporation of glycine into the cell wall glycopeptide in Staphylococcus aureus: role of sRNA and lipid intermediates. Proc. Natl. Acad. Sci. USA 54, 587–594 (1965).
Bumsted, R.M., Dahl, J.L., Söll, D. & Strominger, J.L. Biosynthesis of the peptidoglycan of bacterial cell walls. X. Further study of the glycyl transfer ribonucleic acids active in peptidoglycan synthesis in Staphylococcus aureus. J. Biol. Chem. 243, 779–782 (1968).
Schneider, T. et al. In vitro assembly of a complete, pentaglycine interpeptide bridge containing cell wall precursor (lipid II-Gly5) of Staphylococcus aureus. Mol. Microbiol. 53, 675–685 (2004).
Ruiz, N. Bioinformatics identification of MurJ (MviN) as the peptidoglycan lipid II flippase in Escherichia coli. Proc. Natl. Acad. Sci. USA 105, 15553–15557 (2008).
Sham, L.T. et al. Bacterial cell wall. MurJ is the flippase of lipid-linked precursors for peptidoglycan biogenesis. Science 345, 220–222 (2014).
Ye, X.Y. et al. Better substrates for bacterial transglycosylases. J. Am. Chem. Soc. 123, 3155–3156 (2001).
Lebar, M.D. et al. Forming cross-linked peptidoglycan from synthetic gram-negative Lipid II. J. Am. Chem. Soc. 135, 4632–4635 (2013).
Born, P., Breukink, E. & Vollmer, W. In vitro synthesis of cross-linked murein and its attachment to sacculi by PBP1A from Escherichia coli. J. Biol. Chem. 281, 26985–26993 (2006).
Waxman, D.J. & Strominger, J.L. Penicillin-binding proteins and the mechanism of action of beta-lactam antibiotics. Annu. Rev. Biochem. 52, 825–869 (1983).
Tipper, D.J. & Strominger, J.L. Mechanism of action of penicillins: a proposal based on their structural similarity to acyl-D-alanyl-D-alanine. Proc. Natl. Acad. Sci. USA 54, 1133–1141 (1965).
Lo, M.-C. et al. A new mechanism of action proposed for ramoplanin. J. Am. Chem. Soc. 122, 3540–3541 (2000).
Schwartz, B., Markwalder, J.A. & Wang, Y. Lipid II: total synthesis of the bacterial cell wall precursor and utilization as a substrate for glycosyltransfer and transpeptidation by penicillin binding protein (PBP) 1b of Escherichia coli. J. Am. Chem. Soc. 123, 11638–11643 (2001).
VanNieuwenhze, M.S. et al. The first total synthesis of lipid II: the final monomeric intermediate in bacterial cell wall biosynthesis. J. Am. Chem. Soc. 124, 3656–3660 (2002).
Breukink, E. et al. Lipid II is an intrinsic component of the pore induced by nisin in bacterial membranes. J. Biol. Chem. 278, 19898–19903 (2003).
Tsukamoto, H. & Kahne, D. N-methylimidazolium chloride-catalyzed pyrophosphate formation: application to the synthesis of Lipid I and NDP-sugar donors. Bioorg. Med. Chem. Lett. 21, 5050–5053 (2011).
Bugg, T.D., Braddick, D., Dowson, C.G. & Roper, D.I. Bacterial cell wall assembly: still an attractive antibacterial target. Trends Biotechnol. 29, 167–173 (2011).
Patin, D. et al. Colicin M hydrolyses branched lipids II from Gram-positive bacteria. Biochimie 94, 985–990 (2012).
Zapun, A. et al. In vitro reconstitution of peptidoglycan assembly from the Gram-positive pathogen Streptococcus pneumoniae. ACS Chem. Biol. 8, 2688–2696 (2013).
Lebar, M.D. et al. Reconstitution of peptidoglycan cross-linking leads to improved fluorescent probes of cell wall synthesis. J. Am. Chem. Soc. 136, 10874–10877 (2014).
Huang, L.Y. et al. Enzymatic synthesis of lipid II and analogues. Angew. Chem. Int. Ed. Engl. 53, 8060–8065 (2014).
Guan, Z., Breazeale, S.D. & Raetz, C.R.H. Extraction and identification by mass spectrometry of undecaprenyl diphosphate-MurNAc-pentapeptide-GlcNAc from Escherichia coli. Anal. Biochem. 345, 336–339 (2005).
Qiao, Y. et al. Detection of lipid-linked peptidoglycan precursors by exploiting an unexpected transpeptidase reaction. J. Am. Chem. Soc. 136, 14678–14681 (2014).
Lee, W. et al. The mechanism of action of lysobactin. J. Am. Chem. Soc. 138, 100–103 (2016).
McPherson, D.C. & Popham, D.L. Peptidoglycan synthesis in the absence of class A penicillin-binding proteins in Bacillus subtilis. J. Bacteriol. 185, 1423–1431 (2003).
Silhavy, T.J., Kahne, D. & Walker, S. The bacterial cell envelope. Cold Spring Harb. Perspect. Biol. 2, a000414 (2010).
Butler, E.K., Davis, R.M., Bari, V., Nicholson, P.A. & Ruiz, N. Structure-function analysis of MurJ reveals a solvent-exposed cavity containing residues essential for peptidoglycan biogenesis in Escherichia coli. J. Bacteriol. 195, 4639–4649 (2013).
Higashi, Y., Strominger, J.L. & Sweeley, C.C. Biosynthesis of the peptidoglycan of bacterial cell walls. XXI. Isolation of free C55-isoprenoid alcohol and of lipid intermediates in peptidoglycan synthesis from Staphylococcus aureus. J. Biol. Chem. 245, 3697–3702 (1970).
Park, J.T. Uridine-5′-pyrophosphate derivatives. II. Isolation from Staphylococcus aureus. J. Biol. Chem. 194, 877–884 (1952).
Kohlrausch, U. & Höltje, J.V. Analysis of murein and murein precursors during antibiotic-induced lysis of Escherichia coli. J. Bacteriol. 173, 3425–3431 (1991).
Anderson, J.S., Matsuhashi, M., Haskin, M.A. & Strominger, J.L. Biosythesis of the peptidoglycan of bacterial cell walls. II. Phospholipid carriers in the reaction sequence. J. Biol. Chem. 242, 3180–3190 (1967).
Vollmer, W., Blanot, D. & de Pedro, M.A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167 (2008).
Sieber, P. & Riniker, B. Protection of carboxamide functions by the trityl residue. Application to peptide synthesis. Tetrahedron Lett. 32, 739–742 (1991).
Hoyland, C.N. et al. Structure of the LdcB LD-carboxypeptidase reveals the molecular basis of peptidoglycan recognition. Structure 22, 949–960 (2014).
Lovering, A.L., Safadi, S.S. & Strynadka, N.C.J. Structural perspective of peptidoglycan biosynthesis and assembly. Annu. Rev. Biochem. 81, 451–478 (2012).
Lupoli, T.J. et al. Transpeptidase-mediated incorporation of D-amino acids into bacterial peptidoglycan. J. Am. Chem. Soc. 133, 10748–10751 (2011).
Kuru, E. et al. In Situ probing of newly synthesized peptidoglycan in live bacteria with fluorescent D-amino acids. Angew. Chem. Int. Ed. Engl. 51, 12519–12523 (2012).
Helassa, N., Vollmer, W., Breukink, E., Vernet, T. & Zapun, A. The membrane anchor of penicillin-binding protein PBP2a from Streptococcus pneumoniae influences peptidoglycan chain length. FEBS J. 279, 2071–2081 (2012).
Thumm, G. & Götz, F. Studies on prolysostaphin processing and characterization of the lysostaphin immunity factor (Lif) of Staphylococcus simulans biovar staphylolyticus. Mol. Microbiol. 23, 1251–1265 (1997).
Georgopapadakou, N.H. & Liu, F.Y. Binding of beta-lactam antibiotics to penicillin-binding proteins of Staphylococcus aureus and Streptococcus faecalis: relation to antibacterial activity. Antimicrob. Agents Chemother. 18, 834–836 (1980).
Chambers, H.F. & Miick, C. Characterization of penicillin-binding protein 2 of Staphylococcus aureus: deacylation reaction and identification of two penicillin-binding peptides. Antimicrob. Agents Chemother. 36, 656–661 (1992).
Łeski, T.A. & Tomasz, A. Role of penicillin-binding protein 2 (PBP2) in the antibiotic susceptibility and cell wall cross-linking of Staphylococcus aureus: evidence for the cooperative functioning of PBP2, PBP4, and PBP2A. J. Bacteriol. 187, 1815–1824 (2005).
Barreteau, H. et al. Quantitative high-performance liquid chromatography analysis of the pool levels of undecaprenyl phosphate and its derivatives in bacterial membranes. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 877, 213–220 (2009).
Hartman, B. & Tomasz, A. Altered penicillin-binding proteins in methicillin-resistant strains of Staphylococcus aureus. Antimicrob. Agents Chemother. 19, 726–735 (1981).
van Heijenoort, Y., Gómez, M., Derrien, M., Ayala, J. & van Heijenoort, J. Membrane intermediates in the peptidoglycan metabolism of Escherichia coli: possible roles of PBP 1b and PBP 3. J. Bacteriol. 174, 3549–3557 (1992).
Anderson, J.S., Meadow, P.M., Haskin, M.A. & Strominger, J.L. Biosynthesis of the peptidoglycan of bacterial cell walls. I. Utilization of uridine diphosphate acetylmuramyl pentapeptide and uridine diphosphate acetylglucosamine for peptidoglycan synthesis by particulate enzymes from Staphylococcus aureus and Micrococcus lysodeikticus. Arch. Biochem. Biophys. 116, 487–515 (1966).
Oku, Y., Kurokawa, K., Ichihashi, N. & Sekimizu, K. Characterization of the Staphylococcus aureus mprF gene, involved in lysinylation of phosphatidylglycerol. Microbiology 150, 45–51 (2004).
Kühner, D., Stahl, M., Demircioglu, D.D. & Bertsche, U. From cells to muropeptide structures in 24 h: peptidoglycan mapping by UPLC-MS. Sci. Rep. 4, 7494 (2014).
Acknowledgements
The authors thank J.X. Wang at Harvard Small Molecule Mass Spectrometry Facility for assistance in running LC–MS and LC–MS/MS and interpretation of MS/MS spectra. This work was funded by US National Institutes of Health grants R01 GM100951 to N.R., R01 GM076710 to S.W. and D.K., R01 GM066174 to D.K., and a Singapore A*STAR NSS (PhD) scholarship to Y.Q.
Author information
Authors and Affiliations
Contributions
Y.Q. and V.S. developed methods to isolate and quantify Lipid II from Staphylococcus aureus, Escherichia coli and Bacillus subtilis, based in part on studies conducted by F.R. and K.S.; Y.Q. and V.S. purified S. aureus PBP2; Y.Q. performed studies on PBP2 transpeptidase activity and characterized β-lactam inhibition; N.R. constructed the E. coli MurJA29C strain; S.W. and D.K. designed and supervised the project. The manuscript and figures were prepared by Y.Q., V.S., S.W., and D.K. with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Table 1 and Supplementary Figures 1–11. (PDF 12719 kb)
Source data
Rights and permissions
About this article
Cite this article
Qiao, Y., Srisuknimit, V., Rubino, F. et al. Lipid II overproduction allows direct assay of transpeptidase inhibition by β-lactams. Nat Chem Biol 13, 793–798 (2017). https://doi.org/10.1038/nchembio.2388
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2388
This article is cited by
-
Allosteric activation of cell wall synthesis during bacterial growth
Nature Communications (2023)
-
Modulation of MRSA virulence gene expression by the wall teichoic acid enzyme TarO
Nature Communications (2023)
-
Serum apolipoprotein A-I potentiates the therapeutic efficacy of lysocin E against Staphylococcus aureus
Nature Communications (2021)
-
Staphylococcus aureus cell growth and division are regulated by an amidase that trims peptides from uncrosslinked peptidoglycan
Nature Microbiology (2020)
-
Structure and reconstitution of a hydrolase complex that may release peptidoglycan from the membrane after polymerization
Nature Microbiology (2020)