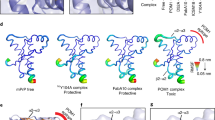
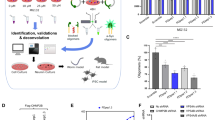
Abstract
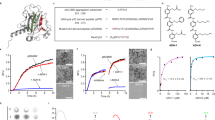
Safely eradicating prions, amyloids and preamyloid oligomers may ameliorate several fatal neurodegenerative disorders. Yet whether small-molecule drugs can directly antagonize the entire spectrum of distinct amyloid structures or 'strains' that underlie distinct disease states is unclear. Here, we investigated this issue using the yeast prion protein Sup35. We have established how epigallocatechin-3-gallate (EGCG) blocks synthetic Sup35 prionogenesis, eliminates preformed Sup35 prions and disrupts inter- and intramolecular prion contacts. Unexpectedly, these direct activities were strain selective, altered the repertoire of accessible infectious forms and facilitated emergence of a new prion strain that configured original, EGCG-resistant intermolecular contacts. In vivo, EGCG cured and prevented induction of susceptible, but not resistant strains, and elicited switching from susceptible to resistant forms. Importantly, 4,5-bis-(4-methoxyanilino)phthalimide directly antagonized EGCG-resistant prions and synergized with EGCG to eliminate diverse Sup35 prion strains. Thus, synergistic small-molecule combinations that directly eradicate complete strain repertoires likely hold considerable therapeutic potential.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Nelson, R. & Eisenberg, D. Structural models of amyloid-like fibrils. Adv. Protein Chem. 73, 235–282 (2006).
Skovronsky, D.M., Lee, V.M.-Y. & Trojanowski, J.Q. Neurodegenerative diseases: new concepts of pathogenesis and their therapeutic implications. Annu. Rev. Pathol. 1, 151–170 (2006).
Prusiner, S.B. Prions. Proc. Natl. Acad. Sci. USA 95, 13363–13383 (1998).
Roberts, B.E. & Shorter, J. Escaping amyloid fate. Nat. Struct. Mol. Biol. 15, 544–546 (2008).
Wells, J.A. & McClendon, C.L. Reaching for high-hanging fruit in drug discovery at protein-protein interfaces. Nature 450, 1001–1009 (2007).
Ehrnhoefer, D.E. et al. Redirecting aggregation pathways: small molecule-mediated conversion of amyloidogenic polypeptides in unstructured, off-pathway oligomers. Nat. Struct. Mol. Biol. 15, 558–566 (2008).
Ehrnhoefer, D.E. et al. Green tea (-)-epigallocatechin-gallate modulates early events in huntingtin misfolding and reduces toxicity in Huntington's disease models. Hum. Mol. Genet. 15, 2743–2751 (2006).
Gestwicki, J.E., Crabtree, G.R. & Graef, I.A. Harnessing chaperones to generate small-molecule inhibitors of amyloid beta aggregation. Science 306, 865–869 (2004).
Hammarstrom, P., Wiseman, R.L., Powers, E.T. & Kelly, J.W. Prevention of transthyretin amyloid disease by changing protein misfolding energetics. Science 299, 713–716 (2003).
Li, J., Zhu, M., Rajamani, S., Uversky, V.N. & Fink, A.L. Rifampicin inhibits alpha-synuclein fibrillation and disaggregates fibrils. Chem. Biol. 11, 1513–1521 (2004).
Wang, H. et al. Direct and selective elimination of specific prions and amyloids by 4,5-dianilinophthalimide and analogs. Proc. Natl. Acad. Sci. USA 105, 7159–7164 (2008).
Legname, G. et al. Synthetic mammalian prions. Science 305, 673–676 (2004).
Petkova, A.T. et al. Self-propagating, molecular-level polymorphism in Alzheimer's beta-amyloid fibrils. Science 307, 262–265 (2005).
Safar, J. et al. Eight prion strains have PrPSc molecules with different conformations. Nat. Med. 4, 1157–1165 (1998).
Tanaka, M., Chien, P., Naber, N., Cooke, R. & Weissman, J.S. Conformational variations in an infectious protein determine prion strain differences. Nature 428, 323–328 (2004).
Tessier, P.M. & Lindquist, S. Unraveling infectious structures, strain variants and species barriers for the yeast prion [PSI+]. Nat. Struct. Mol. Biol. 16, 598–605 (2009).
Tribouillard, D. et al. Antiprion drugs as chemical tools to uncover mechanisms of prion propagation. Prion 1, 48–52 (2007).
Aguzzi, A. Staining, straining and restraining prions. Nat. Neurosci. 11, 1239–1240 (2008).
Castilla, J. et al. Crossing the species barrier by PrPSc replication in vitro generates unique infectious prions. Cell 134, 757–768 (2008).
Kocisko, D.A. et al. New inhibitors of scrapie-associated prion protein formation in a library of 2000 drugs and natural products. J. Virol. 77, 10288–10294 (2003).
Trevitt, C.R. & Collinge, J. A systematic review of prion therapeutics in experimental models. Brain 129, 2241–2265 (2006).
Kocisko, D.A. et al. Comparison of protease-resistant prion protein inhibitors in cell cultures infected with two strains of mouse and sheep scrapie. Neurosci. Lett. 388, 106–111 (2005).
Balch, W.E., Morimoto, R.I., Dillin, A. & Kelly, J.W. Adapting proteostasis for disease intervention. Science 319, 916–919 (2008).
Mu, T.W. et al. Chemical and biological approaches synergize to ameliorate protein-folding diseases. Cell 134, 769–781 (2008).
Shorter, J. & Lindquist, S. Prions as adaptive conduits of memory and inheritance. Nat. Rev. Genet. 6, 435–450 (2005).
Krishnan, R. & Lindquist, S.L. Structural insights into a yeast prion illuminate nucleation and strain diversity. Nature 435, 765–772 (2005).
Tessier, P.M. & Lindquist, S. Prion recognition elements govern nucleation, strain specificity and species barriers. Nature 447, 556–561 (2007).
Scheibel, T., Bloom, J. & Lindquist, S.L. The elongation of yeast prion fibers involves separable steps of association and conversion. Proc. Natl. Acad. Sci. USA 101, 2287–2292 (2004).
Scheibel, T. & Lindquist, S.L. The role of conformational flexibility in prion propagation and maintenance for Sup35p. Nat. Struct. Biol. 8, 958–962 (2001).
Serio, T.R. et al. Nucleated conformational conversion and the replication of conformational information by a prion determinant. Science 289, 1317–1321 (2000).
Shorter, J. & Lindquist, S. Hsp104 catalyzes formation and elimination of self-replicating Sup35 prion conformers. Science 304, 1793–1797 (2004).
Mukhopadhyay, S., Krishnan, R., Lemke, E.A., Lindquist, S. & Deniz, A.A. A natively unfolded yeast prion monomer adopts an ensemble of collapsed and rapidly fluctuating structures. Proc. Natl. Acad. Sci. USA 104, 2649–2654 (2007).
Shorter, J. & Lindquist, S. Destruction or potentiation of different prions catalyzed by similar Hsp104 remodeling activities. Mol. Cell 23, 425–438 (2006).
Scheibel, T., Kowal, A.S., Bloom, J.D. & Lindquist, S.L. Bidirectional amyloid fiber growth for a yeast prion determinant. Curr. Biol. 11, 366–369 (2001).
Toyama, B.H., Kelly, M.J., Gross, J.D. & Weissman, J.S. The structural basis of yeast prion strain variants. Nature 449, 233–237 (2007).
Derkatch, I.L., Chernoff, Y.O., Kushnirov, V.V., Inge-Vechtomov, S.G. & Liebman, S.W. Genesis and variability of [PSI+] prion factors in Saccharomyces cerevisiae. Genetics 144, 1375–1386 (1996).
Tanaka, M., Collins, S.R., Toyama, B.H. & Weissman, J.S. The physical basis of how prion conformations determine strain phenotypes. Nature 442, 585–589 (2006).
Sawaya, M.R. et al. Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Nature 447, 453–457 (2007).
Shewmaker, F., Wickner, R.B. & Tycko, R. Amyloid of the prion domain of Sup35p has an in-register parallel beta-sheet structure. Proc. Natl. Acad. Sci. USA 103, 19754–19759 (2006).
Diaz-Avalos, R., King, C.Y., Wall, J., Simon, M. & Caspar, D.L. Strain-specific morphologies of yeast prion amyloid fibrils. Proc. Natl. Acad. Sci. USA 102, 10165–10170 (2005).
Kayed, R. et al. Common structure of soluble amyloid oligomers implies common mechanism of pathogenesis. Science 300, 486–489 (2003).
O'Nuallain, B. & Wetzel, R. Conformational Abs recognizing a generic amyloid fibril epitope. Proc. Natl. Acad. Sci. USA 99, 1485–1490 (2002).
Patino, M.M., Liu, J.J., Glover, J.R. & Lindquist, S. Support for the prion hypothesis for inheritance of a phenotypic trait in yeast. Science 273, 622–626 (1996).
Kryndushkin, D.S., Alexandrov, I.M., Ter-Avanesyan, M.D. & Kushnirov, V.V. Yeast [PSI+] prion aggregates are formed by small Sup35 polymers fragmented by Hsp104. J. Biol. Chem. 278, 49636–49643 (2003).
Kochneva-Pervukhova, N.V. et al. [PSI+] prion generation in yeast: characterization of the 'strain' difference. Yeast 18, 489–497 (2001).
Staszewski, S. et al. Efavirenz plus zidovudine and lamivudine, efavirenz plus indinavir, and indinavir plus zidovudine and lamivudine in the treatment of HIV-1 infection in adults. N. Engl. J. Med. 341, 1865–1873 (1999).
Burgess, M.R., Skaggs, B.J., Shah, N.P., Lee, F.Y. & Sawyers, C.L. Comparative analysis of two clinically active BCR-ABL kinase inhibitors reveals the role of conformation-specific binding in resistance. Proc. Natl. Acad. Sci. USA 102, 3395–3400 (2005).
Spilman, P. et al. A gamma-secretase inhibitor and quinacrine reduce prions and prevent dendritic degeneration in murine brains. Proc. Natl. Acad. Sci. USA 105, 10595–10600 (2008).
Hennessy, E.J. & Buchwald, S.L. Synthesis of 4,5-dianilinophthalimide and related analogues for potential treatment of Alzheimer's disease via palladium-catalyzed amination. J. Org. Chem. 70, 7371–7375 (2005).
Trinks, U. et al. Dianilinophthalimides: potent and selective, ATP-competitive inhibitors of the EGF-receptor protein tyrosine kinase. J. Med. Chem. 37, 1015–1027 (1994).
Acknowledgements
We thank E. Hennessy (Massachusetts Institute of Technology), S. Buchwald (Massachusetts Institute of Technology), R. Krishnan (Whitehead Institute for Biomedical Research), S. Lindquist (Whitehead Institute for Biomedical Research), J. Weissman (University of California San Francisco), R. Wetzel (University of Pittsburgh School of Medicine) and C. Glabe (University of California Irvine) for generous provision of reagents; J. Chan for preliminary in vivo experiments; and M. Lemmon, A. Gitler, S.B. Cullinan, N. Bonini and S.W. Englander for comments on the manuscript. This work was supported by US National Institutes of Health (NIH) training grant 2T32GM008275-21 (E.A.S), an NIH Director's New Innovator Award (DP2OD002177), an Ellison Medical Foundation New Scholar in Aging Award, an American Heart Association Scientist Development Grant, and University of Pennsylvania Institute on Aging and Alzheimer's Disease Core Center pilots (J.S.).
Author information
Authors and Affiliations
Contributions
B.E.R., M.L.D., H.W., C.C., N.P.L., E.A.S. and J.S. designed experiments, contributed key reagents, performed experiments and interpreted data. M.N.K. contributed key reagents. M.L.D. and J.S. wrote the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 2104 kb)
Rights and permissions
About this article
Cite this article
Roberts, B., Duennwald, M., Wang, H. et al. A synergistic small-molecule combination directly eradicates diverse prion strain structures. Nat Chem Biol 5, 936–946 (2009). https://doi.org/10.1038/nchembio.246
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.246
This article is cited by
-
Structure-based discovery of small molecules that disaggregate Alzheimer’s disease tissue derived tau fibrils in vitro
Nature Communications (2022)
-
Antibiofilm activity of flavonoids on staphylococcal biofilms through targeting BAP amyloids
Scientific Reports (2020)
-
Molecular Targets and Therapeutic Strategies in Spinocerebellar Ataxia Type 7
Neurotherapeutics (2019)
-
Recent perspectives on the molecular basis of biofilm formation by Pseudomonas aeruginosa and approaches for treatment and biofilm dispersal
Folia Microbiologica (2018)
-
The role of amyloidogenic protein oligomerization in neurodegenerative disease
Journal of Molecular Medicine (2013)