Abstract
RNA maturation relies on various exonucleases to remove nucleotides successively from the 5′ or 3′ end of nucleic acids. However, little is known regarding the molecular basis for substrate and cleavage preference of exonucleases. Our biochemical and structural analyses on RNase T–DNA complexes show that the RNase T dimer has an ideal architecture for binding a duplex with a short 3′ overhang to produce a digestion product of a duplex with a 2-nucleotide (nt) or 1-nt 3′ overhang, depending on the composition of the last base pair in the duplex. A 'C-filter' in RNase T screens out the nucleic acids with 3′-terminal cytosines for hydrolysis by inducing a disruptive conformational change at the active site. Our results reveal the general principles and the working mechanism for the final trimming step made by RNase T in the maturation of stable RNA and pave the way for the understanding of other DEDD family exonucleases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Zuo, Y. & Deutscher, M.P. Exoribonuclease superfamilies: structural analysis and phylogenetic distribution. Nucleic Acids Res. 29, 1017–1026 (2001).
Steitz, T.A. & Steitz, J.A. A general two-metal-ion mechanism for catalytic RNA. Proc. Natl. Acad. Sci. USA 90, 6498–6502 (1993).
Hamdan, S., Carr, P.D., Brown, S.E., Ollis, D.L. & Dixon, N.E. Structural basis for proofreading during replication of the Escherichia coli chromosome. Structure 10, 535–546 (2002).
Briggs, M.W., Burkard, K.T. & Butler, J.S. Rrp6p, the yeast homologue of the human PM-Scl 100-kDa autoantigen, is essential for efficient 5.8 S rRNA 3′ end formation. J. Biol. Chem. 273, 13255–13263 (1998).
Hsiao, Y.Y. et al. Crystal structure of CRN-4: implications for domain function in apoptotic DNA degradation. Mol. Cell. Biol. 29, 448–457 (2009).
Wu, M. et al. Structural insight into poly(A) binding and catalytic mechanism of human PARN. EMBO J. 24, 4082–4093 (2005).
Kavanagh, D. et al. New roles for the major human 3′-5′ exonuclease TREX1 in human disease. Cell Cycle 7, 1718–1725 (2008).
Crow, Y.J. & Rehwinkel, J. Aicardi-Goutieres syndrome and related phenotypes: linking nucleic acid metabolism with autoimmunity. Hum. Mol. Genet. 18, R130–R136 (2009).
Deutscher, M.P., Marlor, C.W. & Zaniewski, R. Ribonuclease T: new exoribonuclease possibly involved in end-turnover of tRNA. Proc. Natl. Acad. Sci. USA 81, 4290–4293 (1984).
Li, Z. & Deutscher, M.P. The tRNA processing enzyme RNase T is essential for maturation of 5S RNA. Proc. Natl. Acad. Sci. USA 92, 6883–6886 (1995).
Li, Z., Pandit, S. & Deutscher, M.P. 3′ exoribonucleolytic trimming is a common feature of the maturation of small, stable RNAs in Escherichia coli. Proc. Natl. Acad. Sci. USA 95, 2856–2861 (1998).
Li, Z., Pandit, S. & Deutscher, M.P. Maturation of 23S ribosomal RNA requires the exoribonuclease RNase T. RNA 5, 139–146 (1999).
Li, Z. & Deutscher, M.P. Maturation pathways for E. coli tRNA precursors: a random multienzyme process in vivo. Cell 86, 503–512 (1996).
Kelly, K.O. & Deutscher, M.P. The presence of only one of five exoribonucleases is sufficient to support the growth of Escherichia coli. J. Bacteriol. 174, 6682–6684 (1992).
Viswanathan, M., Lanjuin, A. & Lovett, S.T. Identification of RNase T as a high-copy suppressor of the UV sensitivity associated with single-strand DNA exonuclease deficiency in Escherichia coli. Genetics 151, 929–934 (1999).
Misra, T.K. & Apirion, D. RNase E, an RNA processing enzyme from Escherichia coli. J. Biol. Chem. 254, 11154–11159 (1979).
Allas, U., Liiv, A. & Remme, J. Functional interaction between RNase III and the Escherichia coli ribosome. BMC Mol. Biol. 4, 8 (2003).
Li, Z. & Deutscher, M.P. RNase E plays an essential role in the maturation of Escherichia coli tRNA precursors. RNA 8, 97–109 (2002).
Li, Z. & Deutscher, M.P. The role of individual exoribonucleases in processing at the 3′ end of Escherichia coli tRNA precursors. J. Biol. Chem. 269, 6064–6071 (1994).
Zuo, Y. & Deutscher, M.P. Mechanism of action of RNase T. I. Identification of residues required for catalysis, substrate binding, and dimerization. J. Biol. Chem. 277, 50155–50159 (2002).
Zuo, Y. & Deutscher, M.P. Mechanism of action of RNase T. II. A structural and functional model of the enzyme. J. Biol. Chem. 277, 50160–50164 (2002).
Zuo, Y. et al. Crystal structure of RNase T, an exoribonuclease involved in tRNA maturation and end turnover. Structure 15, 417–428 (2007).
Zuo, Y. & Deutscher, M.P. The DNase activity of RNase T and its application to DNA cloning. Nucleic Acids Res. 27, 4077–4082 (1999).
Viswanathan, M., Dower, K.W. & Lovett, S.T. Identification of a potent DNase activity associated with RNase T of Escherichia coli. J. Biol. Chem. 273, 35126–35131 (1998).
Zuo, Y. & Deutscher, M.P. The physiological role of RNase T can be explained by its unusual substrate specificity. J. Biol. Chem. 277, 29654–29661 (2002).
Zuo, Y., Wang, Y. & Malhotra, A. Crystal structure of Escherichia coli RNase D, an exoribonuclease involved in structured RNA processing. Structure 13, 973–984 (2005).
Chin, K.H., Yang, C.Y., Chou, C.C., Wang, A.H. & Chou, S.H. The crystal structure of XC847 from Xanthomonas campestris: a 3′-5′ oligoribonuclease of DnaQ fold family with a novel opposingly shifted helix. Proteins 65, 1036–1040 (2006).
de Silva, U. et al. The crystal structure of TREX1 explains the 3′ nucleotide specificity and reveals a polyproline II helix for protein partnering. J. Biol. Chem. 282, 10537–10543 (2007).
Jones, S., Daley, D.T., Luscombe, N.M., Berman, H.M. & Thornton, J.M. Protein-RNA interactions: a structural analysis. Nucleic Acids Res. 29, 943–954 (2001).
Luscombe, N.M., Laskowski, R.A. & Thornton, J.M. Amino acid-base interactions: a three-dimensional analysis of protein-DNA interactions at an atomic level. Nucleic Acids Res. 29, 2860–2874 (2001).
Ellis, J.J., Broom, M. & Jones, S. Protein–RNA Interactions: Structural analysis and functional classes. Proteins. 66, 903–911 (2007).
Lavery, R., Moakher, M., Maddocks, J.H., Petkeviciute, D. & Zakrzewska, K. Conformational analysis of nucleic acids revisited: Curves+. Nucleic Acids Res. 37, 5917–5929 (2009).
Cormack, R.S. & Mackie, G.A. Structural requirements for the processing of Escherichia coli 5S ribosomal RNA by RNase E in vitro. J. Mol. Biol. 228, 1078–1090 (1992).
Ehretsmann, C.P., Carpousis, A.J. & Krisch, H.M. Specificity of Escherichia coli endoribonuclease RNase E: in vivo and in vitro analysis of mutants in a bacteriophage T4 mRNA processing site. Genes Dev. 6, 149–159 (1992).
Deutscher, M.P. Ribonucleases, tRNA nucleotidyltransferase, and the 3′ processing of tRNA. Prog. Nucleic Acid Res. Mol. Biol. 39, 209–240 (1990).
Dupasquier, M., Kim, S., Halkidis, K., Gamper, H. & Hou, Y.M. tRNA integrity is a prerequisite for rapid CCA addition: implication for quality control. J. Mol. Biol. 379, 579–588 (2008).
Hartmann, R.K., Gossringer, M., Spath, B., Fischer, S. & Marchfelder, A. The making of tRNAs and more—RNase P and tRNase Z. Prog. Mol. Biol. Transl. Sci. 85, 319–368 (2009).
Guerrier-Takada, C. & Altman, S. Catalytic activity of an RNA molecule prepared by transcription in vitro. Science 223, 285–286 (1984).
Guerrier-Takada, C. & Altman, S. M1 RNA with large terminal deletions retains its catalytic activity. Cell 45, 177–183 (1986).
Lehtinen, D.A., Harvey, S., Mulcahy, M.J., Hollis, T. & Perrino, F.W. The TREX1 double-stranded DNA degradation activity is defective in dominant mutations associated with autoimmune disease. J. Biol. Chem. 283, 31649–31656 (2008).
Padmanabha, K.P. & Deutscher, M.P. RNase T affects Escherichia coli growth and recovery from metabolic stress. J. Bacteriol. 173, 1376–1381 (1991).
Perry, J.J.P. et al. WRN exonuclease structure and molecular mechanism imply an editing role in DNA end processing. Nat. Struct. Mol. Biol. 13, 414–422 (2006).
Yang, Y.G., Lindahl, T. & Barnes, D.E. Trex1 exonuclease degrades ssDNA to prevent chronic checkpoint activation and autoimmune disease. Cell 131, 873–886 (2007).
Lee-Kirsch, M.A. et al. Mutations in the gene encoding the 3′-5′ DNA exonuclease TREX1 are associated with systemic lupus erythematosus. Nat. Genet. 39, 1065–1067 (2007).
Riggs, A.D., Bourgeois, S., Newby, R.F. & Cohn, M. DNA binding of the lac repressor. J. Mol. Biol. 34, 365–368 (1968).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006–2008 (2006).
Acknowledgements
This work was supported by research grants from Academia Sinica and the National Science Council, Taiwan. Portions of this research were carried out at the National Synchrotron Radiation Research Center (BL-13B1 and BL-13C1), a national user facility supported by the National Science Council of Taiwan. The Synchrotron Radiation Protein Crystallography Facility is supported by the National Research Program for Genomic Medicine. We thank S. Lin-Chao (Academia Sinica) for providing the E. coli knockout strains used in this study.
Author information
Authors and Affiliations
Contributions
Y.-Y.H. and H.S.Y. designed experiments and analyzed data. Y.-Y.H., C.-C.Y., C.L.L., J.L.J.L. and Y.D. carried out biochemical and structural analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–3 (PDF 4579 kb)
Supplementary Movie 1
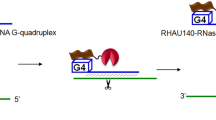
The crystal structure of RNase T bound to a preferred ssDNA with a 3′-end G. The ssDNA is bound at a deep narrow groove near the DEDD active site (PDB ID: 3NGY and 3NH1). Four aromatic residues make π-π stacking interactions with the two 3′-terminal bases. Two Mg2+ ions are bound at the DEDD active site in an active conformation. (MOV 8750 kb)
Supplementary Movie 2
RNase T bound to a non-preferred ssDNA with a 3′-end C induces disruptive conformational changes at the active site. The side chain of Glu73 is rotated to make H-bonds with the 3′-terminal C and the aromatic side chains of Phe29 and Phe27 are shifted to new positions to stack with the 3′-terminal C in the RNase T–DNA2 complex (PDB ID: 3NGZ). As a result, the scissile phosphate in the nonpreferred complex moves up away from the active site and only one Mg2+ ion is bound at the active site in an inactive conformation. (MOV 12732 kb)
Rights and permissions
About this article
Cite this article
Hsiao, YY., Yang, CC., Lin, C. et al. Structural basis for RNA trimming by RNase T in stable RNA 3′-end maturation. Nat Chem Biol 7, 236–243 (2011). https://doi.org/10.1038/nchembio.524
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.524
This article is cited by
-
Combining crystallographic and quantum chemical data to understand DNA-protein π-interactions in nature
Structural Chemistry (2017)
-
The bicoid mRNA localization factor Exuperantia is an RNA-binding pseudonuclease
Nature Structural & Molecular Biology (2016)