Abstract
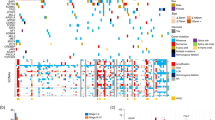
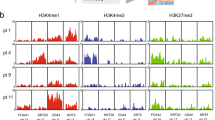
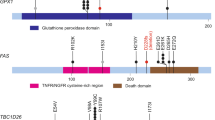
Transitional cell carcinoma (TCC) is the most common type of bladder cancer. Here we sequenced the exomes of nine individuals with TCC and screened all the somatically mutated genes in a prevalence set of 88 additional individuals with TCC with different tumor stages and grades. In our study, we discovered a variety of genes previously unknown to be mutated in TCC. Notably, we identified genetic aberrations of the chromatin remodeling genes (UTX, MLL-MLL3, CREBBP-EP300, NCOR1, ARID1A and CHD6) in 59% of our 97 subjects with TCC. Of these genes, we showed UTX to be altered substantially more frequently in tumors of low stages and grades, highlighting its potential role in the classification and diagnosis of bladder cancer. Our results provide an overview of the genetic basis of TCC and suggest that aberration of chromatin regulation might be a hallmark of bladder cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Parkin, D.M., Bray, F., Ferlay, J. & Pisani, P. Global cancer statistics, 2002. CA Cancer J. Clin. 55, 74–108 (2005).
Wu, X.R. Urothelial tumorigenesis: a tale of divergent pathways. Nat. Rev. Cancer 5, 713–725 (2005).
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007).
Cordon-Cardo, C. et al. p53 mutations in human bladder cancer: genotypic versus phenotypic patterns. Int. J. Cancer 56, 347–353 (1994).
Cairns, P., Proctor, A.J. & Knowles, M.A. Loss of heterozygosity at the RB locus is frequent and correlates with muscle invasion in bladder carcinoma. Oncogene 6, 2305–2309 (1991).
Jebar, A.H. et al. FGFR3 and Ras gene mutations are mutually exclusive genetic events in urothelial cell carcinoma. Oncogene 24, 5218–5225 (2005).
Cappellen, D. et al. Frequent activating mutations of FGFR3 in human bladder and cervix carcinomas. Nat. Genet. 23, 18–20 (1999).
Guenther, M.G., Barak, O. & Lazar, M.A. The SMRT and N-CoR corepressors are activating cofactors for histone deacetylase 3. Mol. Cell. Biol. 21, 6091–6101 (2001).
Lutz, T., Stoger, R. & Nieto, A. CHD6 is a DNA-dependent ATPase and localizes at nuclear sites of mRNA synthesis. FEBS Lett. 580, 5851–5857 (2006).
van Haaften, G. et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nat. Genet. 41, 521–523 (2009).
Wiegand, K.C. et al. ARID1A mutations in endometriosis-associated ovarian carcinomas. N. Engl. J. Med. 363, 1532–1543 (2010).
Dalgliesh, G.L. et al. Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 463, 360–363 (2010).
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011).
Chi, P., Allis, C.D. & Wang, G.G. Covalent histone modifications–miswritten, misinterpreted and mis-erased in human cancers. Nat. Rev. Cancer 10, 457–469 (2010).
Vogelstein, B. & Kinzler, K.W. Cancer genes and the pathways they control. Nat. Med. 10, 789–799 (2004).
Mullighan, C.G. et al. CREBBP mutations in relapsed acute lymphoblastic leukaemia. Nature 471, 235–239 (2011).
Pasqualucci, L. et al. Inactivating mutations of acetyltransferase genes in B-cell lymphoma. Nature 471, 189–195 (2011).
Karamouzis, M.V., Konstantinopoulos, P.A. & Papavassiliou, A.G. Roles of CREB-binding protein (CBP)/p300 in respiratory epithelium tumorigenesis. Cell Res. 17, 324–332 (2007).
Daser, A. & Rabbitts, T.H. The versatile mixed lineage leukaemia gene MLL and its many associations in leukaemogenesis. Semin. Cancer Biol. 15, 175–188 (2005).
Parsons, D.W. et al. The genetic landscape of the childhood cancer medulloblastoma. Science 331, 435–439 (2011).
Sjöblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006).
Bakkar, A.A. et al. FGFR3 and TP53 gene mutations define two distinct pathways in urothelial cell carcinoma of the bladder. Cancer Res. 63, 8108–8112 (2003).
Jones, P.A. & Baylin, S.B. The epigenomics of cancer. Cell 128, 683–692 (2007).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Koboldt, D.C. et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics 25, 2283–2285 (2009).
Yi, X. et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science 329, 75–78 (2010).
Getz, G. et al. Comment on “The consensus coding sequences of human breast and colorectal cancers”. Science 317, 1500 (2007).
Ding, L. et al. Somatic mutations affect key pathways in lung adenocarcinoma. Nature 455, 1069–1075 (2008).
Acknowledgements
This work was supported by grants from the National High Technology Research and Development Program of China (863 Program, 2006AA02A302 to H.Y. 2009AA022707 to X.Z.), the Promotion Program for Shenzhen Key Laboratory, Shenzhen, China (CXB200903090055A and CXB201005250016A to Z.C.), and Research Fund for the Doctoral Program of Higher Education of China (20100001110100).
Author information
Authors and Affiliations
Contributions
Jun Wang, Z.C., Jian Wang, H.Y., S.L. and Y.G. managed the project. A.T., X. Li, L.Z., Z. Li, F.Z., X. Zhao, C. Liang, C. Liu, Y.W., L.S., Z.J., Jing Chen, S. Wu, Z.Z., R. Yang, J. Zhao, C.X., Z.G., J.Y., H. Zhang and W.Y. prepared the samples. X.H., R.W., P.H., H.J., J.L. and X. Zhang performed the sequencing. Y.G., G.G., Y.H., S.G., C.C., M.H., W.J., R. Ye, Z. Liu, S. Wan, H. Zheng, K.K., M.L.N. and Y.L. performed the bioinformatic analysis. Y.H., X.H., Jinnong Chen, S.Y., X. Liu, D.F. and J. Zou performed the validation of somatic mutations. G.G. and Y.H. wrote the paper. Y.G., Jun Wang, Z.C., X.H., Y.L., D.T. and X.S. revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 and Supplementary Tables 1–8. (PDF 442 kb)
Rights and permissions
About this article
Cite this article
Gui, Y., Guo, G., Huang, Y. et al. Frequent mutations of chromatin remodeling genes in transitional cell carcinoma of the bladder. Nat Genet 43, 875–878 (2011). https://doi.org/10.1038/ng.907
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.907
This article is cited by
-
The emerging roles of histone demethylases in cancers
Cancer and Metastasis Reviews (2024)
-
Experimental analysis of bladder cancer-associated mutations in EP300 identifies EP300-R1627W as a driver mutation
Molecular Medicine (2023)
-
Ipsilateral synchronous papillary renal neoplasm with reverse polarity and urothelial carcinoma in a renal transplant recipient: a rare case report with molecular analysis and literature review
Diagnostic Pathology (2023)
-
The impact of mutational clonality in predicting the response to immune checkpoint inhibitors in advanced urothelial cancer
Scientific Reports (2023)
-
Identification of necroptosis-related long non-coding RNAs prognostic signature and the crucial lncRNA in bladder cancer
Journal of Cancer Research and Clinical Oncology (2023)