Abstract
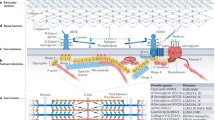
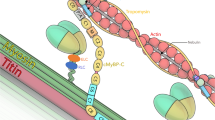
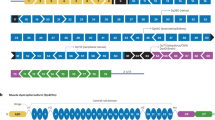
Spontaneous and engineered mouse mutants have facilitated our understanding of the pathogenesis of muscular dystrophy and they provide models for the development of therapeutic approaches1. The mouse myodystrophy (myd) mutation produces an autosomal recessive, neuromuscular phenotype2. Homozygotes have an abnormal gait, show abnormal posturing when suspended by the tail and are smaller than littermate controls. Serum creatine kinase is elevated and muscle histology is typical of a progressive myopathy with focal areas of acute necrosis and clusters of regenerating fibers3. Additional aspects of the phenotype include sensorineural deafness, reduced lifespan and decreased reproductive fitness2,3. The myd mutation maps to mouse chromosome 8 at approximately 33 centimorgans (cM) (refs. 2, 4–7). Here we show that the gene mutated in myd encodes a glycosyltransferase, Large. The human homolog of this gene (LARGE) maps to chromosome 22q. In myd, an intragenic deletion of exons 4–7 causes a frameshift in the resultant mRNA and a premature termination codon before the first of the two catalytic domains. On immunoblots, a monoclonal antibody to α-dystroglycan (a component of the dystrophin-associated glycoprotein complex) shows reduced binding in myd, which we attribute to altered glycosylation of this protein. We speculate that abnormal post-translational modification of α-dystroglycan may contribute to the myd phenotype.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Campbell, K.P. & Allamand, V. Animal models for muscular dystrophy: valuable tools for the development of therapies. Hum. Mol. Genet. 9, 2459–2467 (2000).
Lane, P.W., Beamer, T.C. & Myers, D.D. Myodystrophy, a new myopathy on chromosome 8 of the mouse. J. Hered. 67, 135–138 (1976).
Mathews, K.D., et al. Phenotypic and pathological evaluation of the myd mouse, a candidate model for facioscapulohumeral dystrophy. J. Neuropathol. Exp. Neurol. 54, 601–606 (1995).
Mathews, K.D., Mills, K.A., Bailey, H., Schelper, R. & Murray, J. Mouse myodystrophy (myd) mutation: refined mapping in an interval flanked by homology with distal human 4q. Muscle Nerve S2, S98–S102 (1995).
Mills, K.A., et al. Genetic mapping near the myd locus on mouse chromosome 8. Mamm. Genome 6, 278–280 (1995).
Grewal, P.K., et al. The mouse homolog of FRG1, a candidate gene for FSHD, maps proximal to the myodystrophy mutation on chromosome 8. Mamm. Genome 8, 394–398 (1997).
Grewal, P.K., Carim Todd, L., Maarel, S.V.D., Frants, R.R. & Hewitt, J.E. FRG1, a gene in the FSH muscular dystrophy region on human chromosome 4q35, is highly conserved in vertebrates and invertebrates. Gene 216, 13–19 (1998).
Mathews, K.D. & Mills, K.A. The molecular genetics of human facioscapulohumeral muscular dystrophy and the myodystrophy mouse. Curr. Opin. Neurol. 9, 394–399 (1996).
Grewal, P.K., Bolland, D.J., Carim Todd, L. & Hewitt, J.E. High resolution mapping of mouse chromosome 8 identifies an evolutionary chromosomal breakpoint. Mamm. Genome 9, 603–607 (1998).
Grewal, P.K., et al. Cloning of the murine unconventional myosin gene Myo9b and identification of alternative splicing. Gene 240, 389–398 (1999).
Kim, J., et al. Homology-driven assembly of a sequence-ready mouse BAC contig map spanning regions related to the 46Mb gene-rich euchromatic segments of human chromosome 19. Genomics (in press).
Peyrard, M., et al. The human LARGE gene from 22q12.3-q13.1 is a new distinct member of the glycosyltransferase gene family. Proc. Natl. Acad. Sci. USA 96, 598–603 (1999).
Breton, C. & Imberty, A. Structure/function studies of glycosyltransferases. Curr. Opin. Struct. Biol. 9, 563–571 (1999).
Heinrichs, D.E., Yethon, J.A. & Whitfield, C. Molecular basis for structural diversity in the core region of the lipopolysaccharides of Escherichia coli and Salmonella enterica. Mol. Microbiol. 30, 221–232 (1998).
Sasaki, K., et al. Expression cloning of cDNA encoding a human β-1, 3- N -acetylglucosaminyltransferase that is essential for poly-N-acetyllactosamine synthesis. Proc. Natl. Acad. Sci. USA 94, 14292–14299 (1997).
Durbeej, M., Henry, M.D. & Campbell, K.P. Dystroglycan in development and disease. Curr. Opin. Cell Biol. 10, 594–601 (1998).
Herrmann, R., et al. Dissociation of the dystroglycan complex in caveolin-3-deficient limb girdle muscular dystrophy. Hum. Mol. Genet. 9, 2335–2340 (2000).
Ervasti, J.M. & Campbell, K.P. Membrane organization of the dystrophin-glycoprotein complex. Cell 66, 1121–1131 (1991).
Ibraghimov-Beskrovnaya, O., et al. Primary structure of dystrophin-associated glycoproteins linking dystrophin to the extracellular matrix. Nature 355, 696–702 (1992).
Nutting, D.F., MacPike, A.D. & Meier, H. The calcium content of various tissues from myodystrophic and dystrophic mice. J. Hered. 71, 15–18 (1980).
Côté, P.D., Moukhles, H., Lindebaum, M. & Carbonetto, S. Chimaeric mice deficient in dystroglycans develop muscular dystrophy and have disrupted myoneural synapses. Nature Genet. 23, 338–342 (1999).
Williamson, R.A., et al. Dystroglycan is essential for early embryonic development: disruption of Reichart's membrane in Dag1-null mice. Hum. Mol. Genet. 6, 831–841 (1997).
Kobayashi, K., et al. An ancient retrotransposal insertion causes Fukuyama-type congenital muscular dystrophy. Nature 394, 388–392 (1998).
Aravind, L. & Koonin, E.V. The fukutin protein family—predicted enzymes modifying cell-surface molecules. Curr. Biol. 9, R836–R837 (1999).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Herrmann, R., et al. Dissociation of the dystroglycan complex in caveolin-3-deficient limb girdle muscular dystrophy. Hum. Mol. Genet. 9, 2335–2340 (2000).
Gee, S.H., et al. Laminin-binding protein 120 from brain is closely related to the dystrophin-associated glycoprotein, dystroglycan, and binds with high affinity to the major heparin binding domain of laminin. J. Biol. Chem. 268, 14972–14980 (1993).
Yamada, H., Shimizu, T., Tanaka, T., Campbell, K.P. & Matsumura, K. Dystroglycan is a binding protein of laminin and merosin in peripheral nerve. FEBS Lett. 352, 49–53 (1994).
Acknowledgements
We thank M. Hamshere and P. Scotting for helpful discussions and insightful comments on the manuscript, S. Kröger for generously providing antibody and M. Bucan for access to RPCI BAC library filters. We acknowledge the Wellcome Trust for a travel grant to P.K.G. This work was also supported by the Austrian Verein zur Erforschung der Muskelerkrankungen bei Kindern (P.J.H.) and by grant SFB Nr. 006-F613 from the Austrian Science Research Foundation (R.E.B.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Grewal, P., Holzfeind, P., Bittner, R. et al. Mutant glycosyltransferase and altered glycosylation of α-dystroglycan in the myodystrophy mouse. Nat Genet 28, 151–154 (2001). https://doi.org/10.1038/88865
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/88865
This article is cited by
-
The role of protein glycosylation in muscle diseases
Molecular Biology Reports (2022)
-
LARGE expression in different types of muscular dystrophies other than dystroglycanopathy
BMC Neurology (2018)
-
Biochemical and Functional Interplay Between Ion Channels and the Components of the Dystrophin-Associated Glycoprotein Complex
The Journal of Membrane Biology (2018)
-
Four-week rapamycin treatment improves muscular dystrophy in a fukutin-deficient mouse model of dystroglycanopathy
Skeletal Muscle (2016)
-
Structural basis of laminin binding to the LARGE glycans on dystroglycan
Nature Chemical Biology (2016)