Abstract
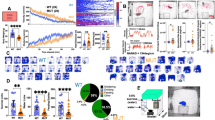
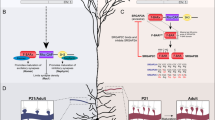
Rab3a is the most abundant Rab (ras-associated binding) protein in the brain and has a regulatory role in synaptic vesicle trafficking1. Mice with a targeted loss-of-function mutation in Rab3a have defects in Ca2+-dependent synaptic transmission: the number of vesicles released in response to an action potential is greater than in wildtype mice, resulting in greater synaptic depression2,3 and the abolishment of CA3 mossy-fiber long term potentiation4. The effect of these changes on behavior is unknown. In a screen for mouse mutants with abnormal rest–activity and sleep patterns, we identified a semidominant mutation, called earlybird, that shortens the circadian period of locomotor activity. Sequence analysis of Rab3a identified a point mutation in the conserved amino acid (Asp77Gly) within the GTP-binding domain of this protein in earlybird mutants, resulting in significantly reduced levels of Rab3a protein. Phenotypic assessment of earlybird mice and a null allele of Rab3a revealed anomalies in circadian period and sleep homeostasis, providing evidence that Rab3a-mediated synaptic transmission is involved in these behaviors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sudhof, T.C. The synaptic vesicle cycle: a cascade of protein–protein interactions. Nature 375, 645–653 (1995).
Geppert, M. et al. The role of Rab3A in neurotransmitter release. Nature 369, 493–497 (1994).
Geppert, M., Goda, Y., Stevens, C.F. & Sudhof, T.C. The small GTP-binding protein Rab3A regulates a late step in synaptic vesicle fusion. Nature 387, 810–814 (1997).
Castillo, P.E. et al. Rab3A is essential for mossy fibre long-term potentiation in the hippocampus. Nature 388, 590–593 (1997).
King, D.P. & Takahashi, J.S. Molecular genetics of circadian rhythms in mammals. Annu. Rev. Neurosci. 23, 713–742 (2000).
Borbely, A.A. A two process model of sleep regulation. Hum. Neurobiol. 1, 195–204 (1982).
Klein, D.C., Moore, R.Y. & Reppert, S.M. Suprachiasmatic Nucleus: The Mind's Clock (Oxford University Press, 1991).
van der Horst, G.T. et al. Mammalian Cry1 and Cry2 are essential for maintenance of circadian rhythms. Nature 398, 627–630 (1999).
Zheng, B. et al. The mPer2 gene encodes a functional component of the mammalian circadian clock. Nature 400, 169–173 (1999).
Bunger, M.K. et al. Mop3 is an essential component of the master circadian pacemaker in mammals. Cell 103, 1009–1017 (2000).
Vitaterna, M.H. et al. Mutagenesis and mapping of a mouse gene, Clock, essential for circadian behavior. Science 264, 719–725 (1994).
King, D.P. et al. Positional cloning of the mouse circadian clock gene. Cell 89, 641–653 (1997).
Nolan, P.M., Kapfhamer, D. & Bucan, M. Random mutagenesis screen for dominant behavioral mutations in mice. Methods 13, 379–395 (1997).
Dumas, J.J., Zhu, Z., Connolly, J.L. & Lambright, D.G. Structural basis of activation and GTP hydrolysis in Rab proteins. Structure Fold Des. 7, 413–423 (1999).
Ostermeier, C. & Brunger, A.T. Structural basis of Rab effector specificity: crystal structure of the small G protein Rab3A complexed with the effector domain of rabphilin-3A. Cell 96, 363–374 (1999).
Bourne, H.R., Sanders, D.A. & McCormick, F. The GTPase superfamily: conserved structure and molecular mechanism. Nature 349, 117–127 (1991).
John, J. et al. Kinetic and structural analysis of the Mg2+-binding site of the guanine nucleotide-binding protein p21H-ras. J. Biol. Chem. 268, 923–929 (1993).
Jung, V. et al. Two types of RAS mutants that dominantly interfere with activators of RAS. Mol. Cell Biol. 14, 3707–3718 (1994).
Panda, S. et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109, 307–320 (2002).
Pennartz, C., de Jeu, M., Bos, N., Schaap, J. & Geurtsen, A. Diurnal modulation of pacemaker potentials and calcium current in the mammalian circadian clock. Nature 416, 286–290 (2002).
Tobler, I. et al. Altered circadian activity rhythms and sleep in mice devoid of prion protein. Nature 380, 639–642 (1996).
Naylor, E. et al. The circadian clock mutation alters sleep homeostasis in the mouse. J. Neurosci. 20, 8138–8143 (2000).
Franken, P., Lopez-Molina, L., Marcacci, L., Schibler, U. & Tafti, M. The transcription factor DBP affects circadian sleep consolidation and rhythmic EEG activity. J. Neurosci. 20, 617–625 (2000).
D'Adamo, P. et al. Mutations in GDI1 are responsible for X-linked non-specific mental retardation. Nature Genet. 19, 134–139 (1998).
Nolan, P.M., Houpt, T.A. & Bucan, M. Chemical mutagenesis and screening for mouse mutations with an altered rest–activity pattern. in Molecular Regulation of Arousal States (ed. Lydic, R.) 143–156 (CRC, 1998).
Veasey, S.C. et al. An automated system for recording and analysis of sleep in mice. Sleep 23, 1025–1040 (2000).
Bennington, J.H., Kodali, S.K. & Heller, H.C. Scoring transitions to REM-sleep in rats based on the EEG phenomena of pre-REM-sleep: an improved analysis of sleep structure. Sleep 17, 28–36 (1994).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Acknowledgements
We thank G. Pickard for his contribution in the early stages of this project; A. Dixon-White for technical assistance; R. Buono, A. deBrunier, J. Eberwine and K. Miyashiro for assistance with immunohistochemistry, western analysis and in situ analysis; L. Maltais for assistance with gene nomenclature; and S. Poethig, M. Chou, T. Abel and G. Leach for comments on the manuscript. This work was supported by grants from the US National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Kapfhamer, D., Valladares, O., Sun, Y. et al. Mutations in Rab3a alter circadian period and homeostatic response to sleep loss in the mouse. Nat Genet 32, 290–295 (2002). https://doi.org/10.1038/ng991
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng991
This article is cited by
-
ADAR-mediated RNA editing suppresses sleep by acting as a brake on glutamatergic synaptic plasticity
Nature Communications (2016)
-
Mammalian sleep genetics
neurogenetics (2012)
-
Small GTPases of the Rab family in the brain of Bombyx mori
Histochemistry and Cell Biology (2010)
-
Rab GTPases as coordinators of vesicle traffic
Nature Reviews Molecular Cell Biology (2009)
-
Search for informative polymorphisms in candidate genes: clock genes and circadian behaviour in blue tits
Genetica (2009)