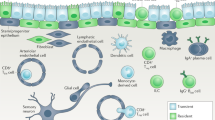
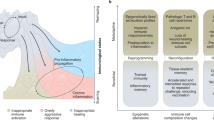
Abstract
Standard definitions of immunological memory are all built on the idea that once infected, animals are protected more efficiently against a second infection. This common view overlooks an unavoidable consequence of the exposure of cells to pathogens, danger signals and environmental agents in general: stimuli change cell properties and activity in a transient yet sustained manner that extends beyond the exposure time and modulates the response of cells of both the innate and adaptive immune systems to secondary stimulation. We suggest that this transient phenomenon represents 'short-term memory' of environmental exposure and discuss the evidence that this is mediated by the persistence of long-lived regulatory molecules, notably a subset of newly deposited chromatin modifications and inducible noncoding RNAs.
Similar content being viewed by others
Main
Memory is the retention of information over a period of time. In the specific case of long-term memory, retrieving the stored information even years after the primary event is an essential component of the process. However, short-term memory (such as memorizing a telephone number just for the time needed before dialing it) is equally relevant in everyday life. As clarified by that example, short-term memory has little to do with the ability to better react to external circumstances on the basis of previous experience. Still, it can affect behavior and maximize performance. To prevent the incremental accumulation and archiving of an excessive amount of information that is relevant for only a short period of time, it is equally important that such short-term memories fade away or be actively removed.
A clinically relevant example of how a transient exposure to a danger signal can affect the ability of a biological system to respond to a secondary encounter with the same signal is endotoxin tolerance, defined as the inability to respond to secondary stimulation with lipopolysaccharide (LPS)1. Indeed, monocytes from humans recovering from sepsis or major trauma are refractory to a subsequent microbial challenge. Such an unresponsive state can be induced within as little as 1 hour of LPS stimulation and may persist for many days2. However, this suppression of the immune system, which is probably needed to prevent the negative consequences of excessive inflammation, may lead to enhanced susceptibility to secondary infections, which in fact represent a major cause of death in those who survive sepsis3,4.
Tolerance to LPS is just one specific example of the more pervasive phenomenon of the conditioning of biological systems by exposure to environmental changes. This conditioning reflects the need to adapt to the external milieu and to maintain such an adapted state after stimulus termination to avoid the adverse consequences of exaggerated, chronic or repeated stimulation. In other words, by 'recording' prior environmental stimulation, cells acquire new properties and maintain them for some time. Thus, the essential features of such phenomena are their persistence beyond the initial stimulation and their decay over time, and in this way they can be considered short-term memories of environmental exposure (Box 1).
In principle, every inducible change that is not rapidly reversed could represent the mechanistic basis for such short-term memory (Fig. 1). We suggest that the mediators of this process are a broad class of stimulus-responsive regulatory molecules that differ from labile second messengers in that they turn over or are eliminated with slow kinetics. For example, a subset of newly deposited chromatin modifications, some inducible regulatory molecules such as microRNAs (miRNAs) and even some activated transcription factors are all characterized by the ability to persist and exert their functions long after termination of the initial stimulus. Such memory states are not epigenetic in the strict sense, in that they do not involve any active transmission through DNA replication and mitosis. Nevertheless, they contribute to modulating, in a sustained but fully reversible way, cellular responses to changing environments.
After the first exposure to an environmental challenge, cells respond by inducing a variety of mediators needed to appropriately react to the stimulus. Most of these mediators are relatively short-lived and are quickly turned off or disposed of after cessation of the stimulus. However, a subset of inducible mediators persists for a variable amount of time after resolution of the response and influences secondary responses to a further challenge.
Histone modification
Not all chromatin modifications are equally suitable mediators of short-term memory. At one end of the spectrum are labile modifications, such as histone acetylation, which are characterized by rapid turnover. Histone acetylation is strongly induced by stimulation because of the ability of stimulus-responsive transcription factors (such as NF-κB and the interferon-regulatory factors) to recruit histone acetyltransferases to their bound genomic sites5,6. However, 'erasure' by histone deacetylases as early as during the course of the stimulation makes it unlikely that this modification is suitable for establishing and maintaining memory. Specifically, at gene promoters, the opposing activities of histone deacetylases and histone acetyltransferases impose turnover times on the order of minutes7. At the other end of the spectrum is DNA methylation, which is the most thermodynamically stable chromatin modification identified so far, and whose deposition occurs slowly during development and is in general poorly responsive to environmental stimuli8. In between those two, other modifications have a range of inducibility and stability, as measured by advanced mass-spectrometry approaches9,10, that makes them potentially suitable for storing information in a sustained but reversible way.
There are two modifications for which a role in mediating sustained effects of transient stimulation has been demonstrated: monomethylation of histone H3 at Lys4 (H3K4me1) and trimethylation of histone H3 at Lys4 (H3K4me3)11,12. H3K4me3 is selectively associated with active or poised gene promoters8,13, and it is deposited after activation of inducible genes with no basal transcription14,15. A role for newly deposited H3K4me3 in macrophage-dependent protection from Candida albicans has been suggested12. Exposure to C. albicans was found to enhance macrophage-dependent protection from reinfection. In peritoneal macrophages, this protective effect was associated with persistence for 1 week of newly deposited H3K4me3 at many genes encoding proinflammatory molecules, as well as at those encoding signal transducers important for the anti–C. albicans response12. It is suggested that the persistence of this mark might be directly linked to memory. A related phenomenon may be represented by the 'bookmarking' by the histone acetyltransferase p300 and RNA polymerase II of the promoters of some early inducible genes after T cell stimulation16. Such 'bookmarking' correlates with faster reactivation of those genes, as well as with their induction in response to stimuli unable to activate them in the absence of prior stimulation. The mechanism responsible for the persistent association of p300 and RNA polymerase II with those promoters is unknown.
H3K4me1 is associated with enhancers17—that is, regulatory regions that act at variable distances (from kilobases to more than one megabase) from their target genes. Enhancers can now be identified on a genome scale on the basis of covalent modifications of histones (H3K4me1 and histone acetylation, notably acetylation of histone H3 at Lys27 (H3K27Ac))18, binding of transcriptional regulators (such as histone acetyltransferases)17,19 and the greater accessibility of the underlying DNA to DNAse I (ref. 20). Data from individual laboratories19,21, as well as the concerted efforts of the ENCODE (Encyclopedia of DNA Elements) consortium22,23, have clarified some basic principles on the use of the available genomic regulatory information. As a general rule, the repertoire of genomic regions that are marked as enhancers and can be coopted for transcriptional regulation in each differentiated cell type is unique and different from that of any other cell type21,24,25. This uniqueness depends on the role of the master transcription factors involved in lineage determination in the selection of the repertoire of enhancer elements active (or poised for activity) in a given lineage26. However, master regulators that specify functionally distinct subsets within a lineage (such as transcription factors that specify T cell subsets; for example, Foxp3 for regulatory T cells, RORγt for the TH17 subset of helper T cells, and T-bet for T helper type 1 (TH1) cells) work within a predetermined landscape generated by transcription factors that act upstream in lineage determination (such as STAT1 and STAT4 for TH1 cells)27,28,29.
Transcription factors activated in differentiated cells in response to stimulation (for example, by microbes) act within the predetermined landscape generated by the master regulators and in general do not change it6,30. However, stimulation of macrophages by a variety of agonists also induces the acquisition of H3K4me1 and acetylated histones at a few thousand regulatory regions that are completely unmarked in naive cells and for this reason have been called 'latent enhancers'11. After termination of the stimulus, a subset of those regions loses acetylated histones while retaining H3K4me1, and after restimulation that subset is able to mediate a faster and stronger transcriptional response (possibly because of the ability of H3K4me1 to promote the recruitment of some histone acetyltransferases)31. Therefore, the epigenome of cells that have been exposed to a given stimulus retains thousands of 'footprints' of the stimulation, which makes these cells transiently different from both unstimulated cells and cells exposed to a different stimulus (Fig. 2).
After the first encounter with a stimulus, cells acutely respond by inducing or silencing various subsets of genes, including those encoding miRNAs. Most of those modifications either are transient and return to a basal state or are very stable and are usually associated with gene silencing. Conversely, some miRNAs and chromatin modifications persist long after the initial stimulus has been removed and may therefore influence the response to a secondary challenge. TF, transcription factor; TSS, transcriptional start site; RISC, RNA-induced silencing complex; 3′UTR, 3′ untranslated region.
An important issue in this context is the nature of the temporal limits of histone-mediated memory effects. Apart from DNA replication and the deposition of newly synthesized histones, and in addition to the active 'erasure' of histone modifications by dedicated enzymes such as histone demethylases32, a central mechanism is mediated by the replication-independent turnover of histones and nucleosomes by specific machineries33. Indeed, at active regulatory regions in particular, two distinct type of events occur: first, the replacement of individual histones of a given nucleosome with histone variants encoded by related but distinct genes (which may have regulatory implications)34; and second, the replacement of the entire histone octamer (the protein core of the nucleosome, around which DNA is wrapped) with a new one by histone chaperones, with the resulting loss of the initial modifications33. Although replication-independent turnover of nucleosomes has been formally demonstrated at genomic regulatory elements in yeast35 the features of this phenomenon in higher eukaryotes are still to be determined.
The functional relevance and mechanism of action of such modifications is still unclear and is to some extent a matter of controversy. The general model is that modified histones act as recognition platforms that enable or stabilize the recruitment of protein complexes that bear specific recognition domains and mediate downstream effects (such as recruitment of the transcriptional and splicing machinery)36. Regardless of the specific mechanisms involved, these results demonstrate that histone methylation has kinetic properties that make it suitable to store information over time, although not necessarily across DNA synthesis and cell division. It is also important to emphasize that histone modifications have an impressive chemical variety37, and the list of modifications that may be prone to cause memory effects because of their intrinsic kinetic properties is probably much larger than can be imagined today.
Three-dimensional chromatin folding and transcriptional memory
The flexibility of the chromatin fiber enables physical interactions between regions located far away from each other in the linear genome. Looping interactions between promoters and distal enhancers involve the binding of RNA polymerase II and sequence-specific transcription factors at both locations38,39 and are sufficient to drive gene expression39. Although the existence of such loops has been known for more than 20 years40, only recently have technological advances (and specifically the development of chromosome conformation capture and its variants)41,42 allowed investigation of chromatin folding in vivo at an unprecedented scale and resolution43,44. A simplified view is that complex eukaryotic genomes (from flies to mammals) are organized in topological domains with an average size of less than 1 megabase in mice and humans45,46,47. Looping interactions between each promoter and its cognate distal regulatory elements probably happen only within the same topological domain. Some studies of yeast have suggested a possible role for such loops in transcriptional memory. After inducible gene activation, the promoters and the 3′ ends of some yeast genes make stable loops that are retained after transcriptional termination and whose persistence correlates with enhanced gene activation after restimulation48. That has led to the intriguing suggestion that such interactions represent 'memory gene loops', although no definitive cause-effect relationship between the two phenomena is available49. The looping landscape of mammalian genes is clearly more complex than that of the much simpler yeast genome, as indicated by recently developed high-resolution techniques50. Nevertheless, the hypothesis that the new formation of gene loops may be involved in the memory of a rapid and inducible transcriptional response is definitely plausible (and is now experimentally testable). Indeed, because of the physical properties of the chromatin fiber, the probability of interaction between two genomic regions is inversely correlated with their physical distance in the linear genome51 and is expected to be extremely low for regulatory elements located far from their target gene. That indicates that some long-range interactions may occur only in a small fraction of the cells at any given time. Therefore, their induction after cells are stimulated and their persistence after the stimulus is terminated may enable the maintenance of contacts that are critical for enabling or maximizing gene activation at restimulation.
DNA methylation and hydroxymethylation
Different from the histone modifications discussed above, DNA methylation is poorly suited to contribute to short-term memory because of its stability and poor reactivity to external stimuli52. However, modifications of 5-methylcytosine (5mC), such as those mediated by the TET ('ten-eleven-translocation') family of dioxygenases53,54, generate chemical species with limited half-lives, which are thus potentially compatible with short-term memory. Specifically, TET enzymes have been shown to catalyze the oxidation of 5mC to 5-hydroxymethylcytosine (5hmC) and to further oxidation products, such as 5-formylcytosine and 5-carboxylcytosine54,55,56. Chemical modifications of such methyl marks can alter transcriptional regulation in many different ways: they might alter the relative affinity of methyl-cytosine–binding proteins, which tether chromatin modifiers to methylated DNA57; they may be recognized by regulatory proteins that specifically bind 5hmC and thus have specific regulatory roles in the context of this modification57,58,59; they may lead to passive demethylation during replication by excluding DNA methyltransferases that maintain symmetric methylation of CpG dinucleotides60; and they may lead to active demethylation by deamination or further oxidation followed by DNA-repair mechanisms55,61,62. DNA modifications are not assumed to be 'on-off switches' that rapidly regulate gene expression. However, indicative of the potential role of such modifications in transcriptional control, gene bodies and transcriptional start sites show enrichment for 5hmC63,64,65,66, and TET proteins contribute to histone modification by interacting with chromatin-modifying enzymes65,67,68. Notably, TET proteins, in particular TET1, can contribute to both activating and repressive effects on gene expression, probably depending on the context and location of binding59,65,69,70. Proteins able to selectively recognize nonmethylated DNA, 5mC, 5hmC, 5-formylcytosine and 5-carboxylcytosine have been identified in mammalian cells71, which hints at a highly specific discrimination of different cytosine-methylation and cytosine-oxidation states.
Because the 5hmC mark can be actively removed from DNA55,61, its inherent relative instability makes it potentially suitable for transient modification of gene expression. Indeed, the concept that 5hmC may provide regulatory functions is suggested by the fact that a simple intermediate of demethylation would be unlikely to persist for hours or days because of the progressive oxidation of 5mC by TET proteins, which leads to 5-formylcytosine or 5-carboxylcytosine that can be excised by thymine-DNA glycosylase. The resultant abasic site is eventually repaired by base-excision repair. In fact, oxidized derivatives of 5mC have been shown to specifically recruit a large number of DNA-repair proteins71, and 5hmC is specifically recognized by chromatin-binding proteins, which may bring about downstream effects71. Given those observations and the growing list of known roles for TET proteins during differentiation, development and even disease63,69,72,73,74,75,76,77,78,79, a situation could be envisioned in which acute stimulation may lead to altered expression or recruitment of those proteins and consequent modification of the 5hmC landscape, ultimately leading to altered expression of specific genes. Such a process would allow a window of time during which 5hmC modification of DNA could contribute to transcriptional control, either by counteracting the repressive effects of 5mC or because of the presence of 5hmC itself61. However, because 5hmC can also be actively removed from the DNA by base-excision repair mechanisms, such a process would have another characteristic of short-term memory—that is, reversibility—and the unmodified cytosine introduced by DNA repair could be remethylated by DNA methyltransferases, thereby ensuring reacquisition of the basal, prestimulus state. Finally, 5hmC-containing DNA has been shown to be demethylated in human cells in a DNA replication–independent manner, which is potentially consistent with a kind of short-term memory that is not necessarily transmitted to daughter cells61,62. In such a hypothetical model, transcription factors could also have an important role in recruiting chromatin-modifying complexes and therefore in determining the sites at which the methylation status will be altered to ensure proper transcriptional regulation80.
Genome-wide mapping and functional analysis of 5hmC has not yet been reported for cells of the immune system, and therefore it will be interesting to learn whether this modification can alter the responses of those cells. However, although a specific role for 5hmC and TET proteins in acute responses has yet to be determined, and although little is known about how TET proteins themselves are transcriptionally regulated81, the dynamic changes in the expression of Tet2 and Tet3 in LPS-stimulated macrophages hint at a possible regulatory role for inducible 5hmC in acute responses (unpublished observations). Moreover, it must be noted that demethylation of discrete regulatory regions after activation of T lymphocytes has been reported. Specifically, after activation of CD4+ T cells, rapid demethylation of the locus encoding interleukin 2 in a DNA replication–independent manner has been observed82. Along the same lines, dynamic methylation at the locus encoding interferon-γ has been reported for memory CD8+ T lymphocytes: although the locus was partially methylated in the resting state, it underwent rapid demethylation within 5 hours of antigenic stimulation83. This process was independent of DNA replication and cell division, and an as-yet-unidentified demethylase activity was proposed. Although the last two examples strictly involve cell activation rather than short-term memory, they suggest that dynamic regulation of the methylation and hydroxymethylation of cytosine at specific loci during acute antigenic stimulation may also contribute to modulating the responses of immune cells, and thus further investigation of their role in the context of memory to a stimulus is warranted.
MicroRNA
Among the regulatory molecules induced by acute stimulation, miRNAs have an essential property that makes them compatible with the maintenance of short-term memory: their stability84. MicroRNAs are regulated at the levels of transcription, biogenesis, stability and decay, all processes that are likely to act in a context- and cell-type-dependent manner85. Relevant to a possible role in propagating the memory of an environmental change, many miRNAs have been shown to be rather stable molecules86. Specifically, in the liver, the miRNA miR-122 has high metabolic stability, regardless of the expression of pre-miR-122, with a half-life estimated to exceed 24 hours87. Even more strikingly, miR-208 has a half-life of over 12 days in mouse heart and is important for the regulation of stress-dependent gene expression and growth of cardiomyocytes88. As exemplified by the fact that the consequences of altered miR-208 expression in the heart become apparent only under stress conditions, the functional implications of miRNA-mediated short-term memory mechanisms are suggested indirectly by numerous observations hinting at a more general role for miRNA in stress responses. Indeed, ablation of many miRNAs is well tolerated in the mouse89, and mutants often develop substantially altered phenotypes only after being subjected to some kind of stress90. These observations suggest that the persistence of inducible miRNAs due to their long half-lives after initial stimulation may enable the maintenance of gene-expression programs that enhance the resistance of cells to repeated exposure, and when such 'memory' miRNAs are absent (such as in animals depleted of the genes encoding the miRNAs), aberrant phenotypes become apparent90.
Most important for a memory effect, miRNAs can be recycled after recognizing their targets and can participate in multiple rounds of targeting91,92. Therefore, after an initial, stimulus-induced burst of transcriptional activation, miRNAs may perpetuate their activity long after the stimulus is removed. Eventually, the miRNA concentration will return to a prestimulus amount through passive dilution as the cells divide and/or through mechanisms of decay and possibly even active secretion93. The last two mechanisms would be especially important for cells that proliferate slowly or not at all, to allow them to eliminate miRNAs that accumulated after acute stimulation. Notably, cell-to-cell transfer of miRNAs as a mean of regulating translation in the recipient cell has been reported for mast cells, dendritic cells and monocytes in the immune system94,95,96. Therefore, secretion of miRNAs may transfer the memory state to neighboring cells not directly exposed to the primary stimulus, thereby transiently modulating the responses of other cells in the microenvironment.
As what eventually matters for the regulation of gene expression is the concentration of a given miRNA relative to that of its target, the process of miRNA-dependent short-term memory might be mediated directly by 'acutely induced' miRNAs and indirectly by miRNAs that are instead actively downregulated after cell activation, which allows a window of time during which intrinsically stable mRNAs can be relieved from suppression by miRNAs and can be expressed. The most likely candidate for regulation of short-term memory in cells of the immune system is miR-146a, one of a few miRNAs that is consistently induced by a variety of stimuli in many cell types97,98,99. Its fundamental role in dampening immune responses has been clearly demonstrated in mice in which the gene encoding it is inactivated100,101. Specifically, ablation of the gene encoding miR-146a leads to hyper-responsiveness to LPS, unrestricted proliferation of myeloid cells and spontaneous development of tumors in secondary lymphoid organs100,101. Indeed, miR-146a acts essentially as a 'brake' on the immune response by directly targeting the signal transducers TRAF6 and IRAK1 and thereby limiting the activation of NF-κB in response to acute stimulation98,99,100. The role of miR-146a in the immune system has been extensively reviewed102,103; here, we just want to suggest the possibility that it could represent a prototypic 'memory' miRNA involved in initiating and maintaining a flexible program of short-term memory triggered by extracellular cues. Potentially all miRNAs that are induced after stimulation and are not actively or passively eliminated (that is, by dilution in proliferating cells) within the time of the challenge may help alter the cellular transcriptome for a window of time large enough to affect secondary responses. Additional miRNAs induced by specific stimuli in cells of the innate immune system include (among others) miR-155 (refs. 98,104,105), miR-150 (ref. 106), miR-221 (ref. 107) and miR-187 (ref. 108). Similar to miR-146a, miR-155 is an important regulator of inflammation, although with an effect opposite to that of miR-146a, as it enhances rather than inhibiting inflammation109. Stimulation of natural killer (NK) T cells with α-galactosylceramide induces miR-150, which regulates cytokine production in both NK cells and NKT cells106,110. Moreover, miR-221 is proposed to contribute to an enhanced response to a secondary challenge in mast cells107, whereas miR-187 has been shown to negatively regulate the production of various cytokines in monocytes108. Indicative of the possible relevance of miRNAs in regulating the stimulus responsiveness of monocytes from patients with sepsis, miR-146a, miR-155, miR-9 and miR-187 have all been shown to be specifically upregulated in peripheral blood mononuclear cells of patients with systemic inflammatory response syndrome108.
As for miRNAs that are actively downregulated after cell activation, the abundance of mature miRNAs can be regulated by a variety of mechanisms, first and foremost being the availability of Argonaute proteins, which are essential for the assembly and function of the miRNA-containing RNA-induced silencing complex. Indeed, expression of these proteins is a limiting factor for miRNA expression, which suggests they have a protective role in preserving the overall concentration of miRNA85,111. Specifically, genetic ablation or forced expression of these proteins results in a lower or higher concentration of mature miRNAs, respectively, an effect probably due to the stabilization of miRNAs mediated by their binding to the Argonaute proteins111,112,113. Although this mechanism probably does not act selectively on specific miRNAs, it could nevertheless contribute to rapid remodeling of the miRNA repertoire during and after cell activation. More specifically for nondividing cells, both endogenous miRNAs and exogenous transfected small interfering RNAs can persist for over 3 weeks in inert complexes containing Argonaute proteins that are reactivated after mitogenic stimulation of quiescent cells114. In fact, that last study provided formal evidence that mature miRNAs are able to bear functional regulatory information for an extended period of time, regardless of the expression of the primary miRNA transcript, to modulate subsequent cellular responses.
Although only a few specific exonucleases that can catalyze the degradation of mature miRNAs in mammals have been identified84,115, a growing number of individual miRNAs have been shown to undergo accelerated decay in specific contexts84. For example, rapid degradation of miRNA seems to be especially active in neurons, which may compensate for their inability to dilute miRNA concentration by cell division116. Specifically, miR-183–miR-96–miR-182, miR-204 and miR-211 are regulated by light in the mouse retina, and their concentrations are halved within ∼90 minutes of exposure to dark116. That state is reversible, as after reexposure to light those same miRNAs return to their maximal concentrations within 30 minutes as a result of transcriptional upregulation116. Interestingly, a circular RNA with high expression in mouse and human brain functions as a miR-7 'sponge', which indicates a possible additional layer of regulation of miRNA function117. Although many questions remain about the exact mechanism and physiological relevance of the accelerated decay of specific miRNAs, the degradation and turnover of these regulatory molecules probably represent important mechanisms in the context of short-term memory that allow the active elimination of memory miRNAs, especially by noncycling cells. Along the same lines, the 3′-to-5′ exoRNase Eri1 has been shown to regulate miRNA turnover in NK cells and T lymphocytes in the mouse118, which indicates that mechanisms of regulating miRNA decay may be also active and physiologically relevant in cells of the innate and adaptive immune systems. Ultimately, the overall abundance of a given miRNA in any particular state of cell activation may be determined by the combination of its transcription, the expression of Argonaute proteins, its dilution among newly transcribed RNAs (especially in fast-dividing cells) and its active decay.
Long noncoding RNA
The possible role of long noncoding RNAs in cellular memory is much less clear. These RNAs vary in length from a few hundred bases to tens of kilobases, and although they are generated from a large number of genomic locations, in most cases they have very low expression (less than one copy per cell) and are expressed in a cell type–specific way119. Although a function has been identified for only a few of these molecules, they may have diverse roles, ranging from regulation of chromosome inactivation to control of gene expression, and they are worth mentioning here as another potential mechanism of inducible transient regulation of gene expression. Several long noncoding RNAs are important during development and are involved in transcriptional repression, and many can bind polycomb protein complexes and other chromatin-modifying complexes120,121,122. However, a role for these RNAs in acute stimulation and response to secondary encounter with a stimulus remains to be established. Here we will just mention a few such RNAs that have been shown to regulate signal transduction or to be regulated by transcription factors, as examples of additional molecules that may regulate short-term memory of stimuli. A long noncoding RNA involved in regulating signal transduction after activation is NRON ('noncoding repressor of NFAT'), which modulates trafficking of the transcription factor NFAT to the nucleus123. Although it is unclear whether NRON expression is modulated by stimulation, it has been found to function as the RNA component of a large cytoplasmic RNA-protein scaffold complex that regulates the subcellular localization and activity of NFAT in response to stimulation123,124. Consistent with that, T cells depleted of NRON have enhanced production of NFAT-dependent cytokines124. Another example of the modulation of gene activity by long noncoding RNAs in response to external stimuli is provided by two such RNAs, lincRNA-p21 and PANDA, which act in the tumor-suppressor p53 pathway to regulate apoptosis and cell-cycle arrest121,125,126.
Other long noncoding RNAs have enhancer-like functions and activate gene expression121,127. One such RNA, NeST, has been shown to induce interferon-γ expression in activated CD8+ T lymphocytes and to regulate susceptibility to viral and bacterial pathogens128. The effect of NeST on interferon-γ was conferred even by transgenic expression from ectopic loci, which indicates that NeST function could be provided in trans. Finally, long noncoding RNAs can also regulate gene expression by functioning as decoys for other RNA species, such as miRNAs129,130. Indeed, the pseudogene PTENP1 can compete with PTEN for binding of miRNAs and thus effectively derepress PTEN expression and modulate cell growth129.
Concluding remarks
We have proposed an initial conceptual and mechanistic framework for the short-term memory of transient environmental stimuli, and specifically immunological stimuli, and have suggested possible involvement of such memory in controlling sustained feed-forward loops and negative feedbacks triggered by stimulation. Although miRNAs and chromatin modifications with an intermediate degree of stability represent obvious mediators of this short-term memory, additional participants may be involved as well—notably, signal-activated transcription factors. For example, antigen-experienced CD8+ T lymphocytes accumulate NFAT into the nucleus within less than a minute of contact with antigen-presenting cells in vivo131. However, after loss of contact with antigen-presenting cells, extrusion of NFAT from the nucleus has much slower kinetics, which allows transcriptional activation of a specific subset of target genes and in fact results in short-term memory of the transient cell-to-cell contact131. The slow kinetics of the export of nuclear NFAT are not due to continuous signaling, and therefore this represents a true example of transcription factor–dependent short-term memory in which the persistence of a transcription factor in the nucleus long after the stimulus is removed induces temporary changes that alter secondary responses to the same environmental challenge. In this conceptual framework, it will be important to reconsider the kinetics of activation and deactivation of signal-responsive transcription factors activated by short-lasting stimuli, which probably represent the most common occurrence in vivo.
To summarize, we propose that the memory of environmental changes mediated by regulatory mechanisms characterized by rapid induction and slow decay is a common and pervasive feature typical of cells of innate and adaptive immunity and even of non-immune cells.
References
Biswas, S.K. & Lopez-Collazo, E. Endotoxin tolerance: new mechanisms, molecules and clinical significance. Trends Immunol. 30, 475–487 (2009).
del Fresno, C. et al. Potent phagocytic activity with impaired antigen presentation identifying lipopolysaccharide-tolerant human monocytes: demonstration in isolated monocytes from cystic fibrosis patients. J. Immunol. 182, 6494–6507 (2009).
Limaye, A.P. et al. Cytomegalovirus reactivation in critically ill immunocompetent patients. J. Am. Med. Assoc. 300, 413–422 (2008).
van der Poll, T. & Opal, S.M. Host-pathogen interactions in sepsis. Lancet Infect. Dis. 8, 32–43 (2008).
Hottiger, M.O., Felzien, L.K. & Nabel, G.J. Modulation of cytokine-induced HIV gene expression by competitive binding of transcription factors to the coactivator p300. EMBO J. 17, 3124–3134 (1998).
Ghisletti, S. et al. Identification and characterization of enhancers controlling the inflammatory gene expression program in macrophages. Immunity 32, 317–328 (2010).
Hazzalin, C.A. & Mahadevan, L.C. Dynamic acetylation of all lysine 4-methylated histone H3 in the mouse nucleus: analysis at c-fos and c-jun. PLoS Biol. 3, e393 (2005).
Bernstein, B.E., Meissner, A. & Lander, E.S. The mammalian epigenome. Cell 128, 669–681 (2007).
Zee, B.M., Levin, R.S., Dimaggio, P.A. & Garcia, B.A. Global turnover of histone post-translational modifications and variants in human cells. Epigenetics Chromatin 3, 22 (2010).
Zee, B.M. et al. In vivo residue-specific histone methylation dynamics. J. Biol. Chem. 285, 3341–3350 (2009).
Ostuni, R. et al. Latent enhancers activated by stimulation in differentiated cells. Cell 152, 157–171 (2013).
Quintin, J. et al. Candida albicans infection affords protection against reinfection via functional reprogramming of monocytes. Cell Host Microbe 12, 223–232 (2012).
Ruthenburg, A.J., Allis, C.D. & Wysocka, J. Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol. Cell 25, 15–30 (2007).
Hargreaves, D.C., Horng, T. & Medzhitov, R. Control of inducible gene expression by signal-dependent transcriptional elongation. Cell 138, 129–145 (2009).
De Santa, F. et al. Jmjd3 contributes to the control of gene expression in LPS-activated macrophages. EMBO J. 28, 3341–3352 (2009).
Byun, J.S. et al. Dynamic bookmarking of primary response genes by p300 and RNA polymerase II complexes. Proc. Natl. Acad. Sci. USA 106, 19286–19291 (2009).
Heintzman, N. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Visel, A. et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009).
Boyle, A.P. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008).
Heintzman, N.D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Natoli, G. Maintaining cell identity through global control of genomic organization. Immunity 33, 12–24 (2010).
Ciofani, M. et al. A validated regulatory network for Th17 cell specification. Cell 151, 289–303 (2012).
Samstein, R.M. et al. Foxp3 exploits a pre-existent enhancer landscape for regulatory T cell lineage specification. Cell 151, 153–166 (2012).
Vahedi, G. et al. STATs shape the active enhancer landscape of T cell populations. Cell 151, 981–993 (2012).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Jeong, K.W. et al. Recognition of enhancer element-specific histone methylation by TIP60 in transcriptional activation. Nat. Struct. Mol. Biol. 18, 1358–1365 (2011).
Cloos, P.A., Christensen, J., Agger, K. & Helin, K. Erasing the methyl mark: histone demethylases at the center of cellular differentiation and disease. Genes Dev. 22, 1115–1140 (2008).
Burgess, R.J. & Zhang, Z. Histone chaperones in nucleosome assembly and human disease. Nat. Struct. Mol. Biol. 20, 14–22 (2013).
Elsaesser, S.J., Goldberg, A.D. & Allis, C.D. New functions for an old variant: no substitute for histone H3.3. Curr. Opin. Genet. Dev. 20, 110–117 (2010).
Dion, M.F. et al. Dynamics of replication-independent histone turnover in budding yeast. Science 315, 1405–1408 (2007).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Tan, M. et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146, 1016–1028 (2011).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Deng, W. et al. Controlling long-range genomic interactions at a native locus by targeted tethering of a looping factor. Cell 149, 1233–1244 (2012).
Cullen, K.E., Kladde, M.P. & Seyfred, M.A. Interaction between transcription regulatory regions of prolactin chromatin. Science 261, 203–206 (1993).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
de Wit, E. & de Laat, W. A decade of 3C technologies: insights into nuclear organization. Genes Dev. 26, 11–24 (2012).
Tanay, A. & Cavalli, G. Chromosomal domains: epigenetic contexts and functional implications of genomic compartmentalization. Curr. Opin. Genet. Dev. 23, 197–203 (2013).
Gibcus, J.H. & Dekker, J. The hierarchy of the 3D genome. Mol. Cell 49, 773–782 (2013).
Dixon, J.R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E.P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Tan-Wong, S.M., Wijayatilake, H.D. & Proudfoot, N.J. Gene loops function to maintain transcriptional memory through interaction with the nuclear pore complex. Genes Dev. 23, 2610–2624 (2009).
Deng, W. & Blobel, G.A. Do chromatin loops provide epigenetic gene expression states? Curr. Opin. Genet. Dev. 20, 548–554 (2010).
van de Werken, H.J. et al. Robust 4C-seq data analysis to screen for regulatory DNA interactions. Nat. Methods 9, 969–972 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
He, Y.F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
Mellén, M., Ayata, P., Dewell, S., Kriaucionis, S. & Heintz, N. MeCP2 binds to 5hmC enriched within active genes and accessible chromatin in the nervous system. Cell 151, 1417–1430 (2012).
Yildirim, O. et al. Mbd3/NURD complex regulates expression of 5-hydroxymethylcytosine marked genes in embryonic stem cells. Cell 147, 1498–1510 (2011).
Wu, H. et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 473, 389–393 (2011).
Valinluck, V. & Sowers, L.C. Endogenous cytosine damage products alter the site selectivity of human DNA maintenance methyltransferase DNMT1. Cancer Res. 67, 946–950 (2007).
Guo, J.U., Su, Y., Zhong, C., Ming, G.L. & Song, H. Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell 145, 423–434 (2011).
Cortellino, S. et al. Thymine DNA glycosylase is essential for active DNA demethylation by linked deamination-base excision repair. Cell 146, 67–79 (2011).
Ficz, G. et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 473, 398–402 (2011).
Pastor, W.A. et al. Genome-wide mapping of 5-hydroxymethylcytosine in embryonic stem cells. Nature 473, 394–397 (2011).
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011).
Xu, Y. et al. Genome-wide regulation of 5hmC, 5mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Mol. Cell 42, 451–464 (2011).
Chen, Q., Chen, Y., Bian, C., Fujiki, R. & Yu, X. TET2 promotes histone O-GlcNAcylation during gene transcription. Nature 493, 561–564 (2013).
Vella, P. et al. Tet proteins connect the O-linked N-acetylglucosamine transferase Ogt to chromatin in embryonic stem cells. Mol. Cell 49, 645–656 (2013).
Dawlaty, M.M. et al. Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9, 166–175 (2011).
Wu, H. et al. Genome-wide analysis of 5-hydroxymethylcytosine distribution reveals its dual function in transcriptional regulation in mouse embryonic stem cells. Genes Dev. 25, 679–684 (2011).
Spruijt, C.G. et al. Dynamic readers for 5-(hydroxy)methylcytosine and Its oxidized derivatives. Cell 152, 1146–1159 (2013).
Dawlaty, M.M. et al. Combined deficiency of tet1 and tet2 causes epigenetic abnormalities but is compatible with postnatal development. Dev. Cell 24, 310–323 (2013).
Ko, M. et al. Ten-Eleven-Translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc. Natl. Acad. Sci. USA 108, 14566–14571 (2011).
Ko, M. et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 468, 839–843 (2010).
Koh, K.P. et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8, 200–213 (2011).
Li, Z. et al. Deletion of Tet2 in mice leads to dysregulated hematopoietic stem cells and subsequent development of myeloid malignancies. Blood 118, 4509–4518 (2011).
Moran-Crusio, K. et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell 20, 11–24 (2011).
Cimmino, L., Abdel-Wahab, O., Levine, R.L. & Aifantis, I. TET family proteins and their role in stem cell differentiation and transformation. Cell Stem Cell 9, 193–204 (2011).
Gu, T.P. et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477, 606–610 (2011).
Smith, Z.D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Kallin, E.M. et al. Tet2 facilitates the derepression of myeloid target genes during CEBPalpha-induced transdifferentiation of pre-B cells. Mol. Cell 48, 266–276 (2012).
Bruniquel, D. & Schwartz, R.H. Selective, stable demethylation of the interleukin-2 gene enhances transcription by an active process. Nat. Immunol. 4, 235–240 (2003).
Kersh, E.N. et al. Rapid demethylation of the IFN-γ gene occurs in memory but not naive CD8 T cells. J. Immunol. 176, 4083–4093 (2006).
Rüegger, S. & Grosshans, H. MicroRNA turnover: when, how, and why. Trends Biochem. Sci. 37, 436–446 (2012).
Kai, Z.S. & Pasquinelli, A.E. MicroRNA assassins: factors that regulate the disappearance of miRNAs. Nat. Struct. Mol. Biol. 17, 5–10 (2010).
Bhattacharyya, S.N., Habermacher, R., Martine, U., Closs, E.I. & Filipowicz, W. Relief of microRNA-mediated translational repression in human cells subjected to stress. Cell 125, 1111–1124 (2006).
Gatfield, D. et al. Integration of microRNA miR-122 in hepatic circadian gene expression. Genes Dev. 23, 1313–1326 (2009).
van Rooij, E. et al. Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 316, 575–579 (2007).
Park, C.Y. et al. A resource for the conditional ablation of microRNAs in the mouse. Cell Rep 1, 385–391 (2012).
Leung, A.K. & Sharp, P.A. MicroRNA functions in stress responses. Mol. Cell 40, 205–215 (2010).
Hutvágner, G. & Zamore, P.D. A microRNA in a multiple-turnover RNAi enzyme complex. Science 297, 2056–2060 (2002).
Baccarini, A. et al. Kinetic analysis reveals the fate of a microRNA following target regulation in mammalian cells. Curr. Biol. 21, 369–376 (2011).
Chen, X., Liang, H., Zhang, J., Zen, K. & Zhang, C.Y. Secreted microRNAs: a new form of intercellular communication. Trends Cell Biol. 22, 125–132 (2012).
Montecalvo, A. et al. Mechanism of transfer of functional microRNAs between mouse dendritic cells via exosomes. Blood 119, 756–766 (2012).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Zhang, Y. et al. Secreted monocytic miR-150 enhances targeted endothelial cell migration. Mol. Cell 39, 133–144 (2010).
Rusca, N. et al. MiR-146a and NF-κB1 regulate mast cell survival and T lymphocyte differentiation. Mol. Cell Biol. 32, 4432–4444 (2012).
Taganov, K.D., Boldin, M.P., Chang, K.J. & Baltimore, D. NF-κB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 103, 12481–12486 (2006).
Yang, L. et al. miR-146a controls the resolution of T cell responses in mice. J. Exp. Med. 209, 1655–1670 (2012).
Boldin, M.P. et al. miR-146a is a significant brake on autoimmunity, myeloproliferation, and cancer in mice. J. Exp. Med. 208, 1189–1201 (2011).
Zhao, J.L. et al. NF-κB dysregulation in microRNA-146a-deficient mice drives the development of myeloid malignancies. Proc. Natl. Acad. Sci. USA 108, 9184–9189 (2011).
Boldin, M.P. & Baltimore, D. MicroRNAs, new effectors and regulators of NF-κB. Immunol. Rev. 246, 205–220 (2012).
Montagner, S., Orlandi, E.M., Merante, S. & Monticelli, S. The role of miRNAs in mast cells and other innate immune cells. Immunol. Rev. (in the press) (2013).
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611 (2007).
Thai, T.H. et al. Regulation of the germinal center response by microRNA-155. Science 316, 604–608 (2007).
Zheng, Q., Zhou, L. & Mi, Q.S. MicroRNA miR-150 is involved in Vα14 invariant NKT cell development and function. J. Immunol. 188, 2118–2126 (2012).
Mayoral, R.J. et al. MiR-221 influences effector functions and actin cytoskeleton in mast cells. PLoS ONE 6, e26133 (2011).
Rossato, M. et al. IL-10-induced microRNA-187 negatively regulates TNF-α, IL-6, and IL-12p40 production in TLR4-stimulated monocytes. Proc. Natl. Acad. Sci. USA 109, E3101–E3110 (2012).
O'Connell, R.M., Chaudhuri, A.A., Rao, D.S. & Baltimore, D. Inositol phosphatase SHIP1 is a primary target of miR-155. Proc. Natl. Acad. Sci. USA 106, 7113–7118 (2009).
Bezman, N.A., Chakraborty, T., Bender, T. & Lanier, L.L. miR-150 regulates the development of NK and iNKT cells. J. Exp. Med. 208, 2717–2731 (2011).
Bronevetsky, Y. et al. T cell activation induces proteasomal degradation of Argonaute and rapid remodeling of the microRNA repertoire. J. Exp. Med. 210, 417–432 (2013).
Diederichs, S. & Haber, D.A. Dual role for argonautes in microRNA processing and posttranscriptional regulation of microRNA expression. Cell 131, 1097–1108 (2007).
O'Carroll, D. et al. A Slicer-independent role for Argonaute 2 in hematopoiesis and the microRNA pathway. Genes Dev. 21, 1999–2004 (2007).
Olejniczak, S.H., La Rocca, G., Gruber, J.J. & Thompson, C.B. Long-lived microRNA-Argonaute complexes in quiescent cells can be activated to regulate mitogenic responses. Proc. Natl. Acad. Sci. USA 110, 157–162 (2013).
Chatterjee, S. & Grosshans, H. Active turnover modulates mature microRNA activity in Caenorhabditis elegans. Nature 461, 546–549 (2009).
Krol, J. et al. Characterizing light-regulated retinal microRNAs reveals rapid turnover as a common property of neuronal microRNAs. Cell 141, 618–631 (2010).
Hansen, T.B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013).
Thomas, M.F. et al. Eri1 regulates microRNA homeostasis and mouse lymphocyte development and antiviral function. Blood 120, 130–142 (2012).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Guttman, M. & Rinn, J.L. Modular regulatory principles of large non-coding RNAs. Nature 482, 339–346 (2012).
Wang, K.C. & Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 43, 904–914 (2011).
Ponting, C.P., Oliver, P.L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009).
Willingham, A.T. et al. A strategy for probing the function of noncoding RNAs finds a repressor of NFAT. Science 309, 1570–1573 (2005).
Sharma, S. et al. Dephosphorylation of the nuclear factor of activated T cells (NFAT) transcription factor is regulated by an RNA-protein scaffold complex. Proc. Natl. Acad. Sci. USA 108, 11381–11386 (2011).
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010).
Hung, T. et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat. Genet. 43, 621–629 (2011).
Ørom, U.A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010).
Gomez, J.A. et al. The NeST long ncRNA controls microbial susceptibility and epigenetic activation of the interferon-gamma locus. Cell 152, 743–754 (2013).
Poliseno, L. et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465, 1033–1038 (2010).
Salmena, L., Poliseno, L., Tay, Y., Kats, L. & Pandolfi, P.P. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146, 353–358 (2011).
Marangoni, F. et al. The transcription factor NFAT exhibits signal memory during serial T cell interactions with antigen-presenting cells. Immunity 38, 237–249 (2013).
Acknowledgements
We thank K.M. Ansel for encouragement. Supported by the Swiss National Science Foundation (31003A_138343 for the S.M. laboratory), Novartis Stiftung für Medizinisch Biologische Forschung (S.M. laboratory) and the European Research Council (G.N. laboratory).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Monticelli, S., Natoli, G. Short-term memory of danger signals and environmental stimuli in immune cells. Nat Immunol 14, 777–784 (2013). https://doi.org/10.1038/ni.2636
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2636
This article is cited by
-
Interleukin-18-primed human umbilical cord-mesenchymal stem cells achieve superior therapeutic efficacy for severe viral pneumonia via enhancing T-cell immunosuppression
Cell Death & Disease (2023)
-
Diabetic microenvironment preconditioning of adipose tissue-derived mesenchymal stem cells enhances their anti-diabetic, anti-long-term complications, and anti-inflammatory effects in type 2 diabetic rats
Stem Cell Research & Therapy (2022)
-
Adaptation and memory in immune responses
Nature Immunology (2019)
-
Manufacturing of primed mesenchymal stromal cells for therapy
Nature Biomedical Engineering (2019)
-
Interferon target-gene expression and epigenomic signatures in health and disease
Nature Immunology (2019)