Abstract
Powerful methods now allow the imaging of specific mRNAs in living cells. These methods enlist fluorescent proteins to illuminate mRNAs, use labeled oligonucleotide probes and exploit aptamers that render organic dyes fluorescent. The intracellular dynamics of mRNA synthesis, transport and localization can be analyzed at higher temporal resolution with these methods than has been possible with traditional fixed-cell or biochemical approaches. These methods have also been adopted to visualize and track single mRNA molecules in real time. This review explores the promises and limitations of these methods.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dirks, R.W. & Tanke, H.J. Advances in fluorescent tracking of nucleic acids in living cells. Biotechniques 40, 489–496 (2006).
Rodriguez, A.J., Condeelis, J., Singer, R.H. & Dictenberg, J.B. Imaging mRNA movement from transcription sites to translation sites. Semin. Cell Dev. Biol. 18, 202–208 (2007).
Querido, E. & Chartrand, P. Using fluorescent proteins to study mRNA trafficking in living cells. Methods Cell Biol. 85, 273–292 (2008).
LeCuyer, K.A., Behlen, L.S. & Uhlenbeck, O.C. Mutagenesis of a stacking contact in the MS2 coat protein–RNA complex. EMBO J. 15, 6847–6853 (1996).
Bertrand, E. et al. Localization of ASH1 mRNA particles in living yeast. Mol. Cell 2, 437–445 (1998). First paper to describe the MS2 coat protein–GFP approach for imaging mRNA dynamics in live cells.
Beach, D.L., Salmon, E.D. & Bloom, K. Localization and anchoring of mRNA in budding yeast. Curr. Biol. 9, 569–578 (1999).
Forrest, K.M. & Gavis, E.R. Live imaging of endogenous RNA reveals a diffusion and entrapment mechanism for nanos mRNA localization in Drosophila. Curr. Biol. 13, 1159–1168 (2003).
Zimyanin, V.L. et al. In vivo imaging of oskar mRNA transport reveals the mechanism of posterior localization. Cell 134, 843–853 (2008).
Weil, T.T., Forrest, K.M. & Gavis, E.R. Localization of bicoid mRNA in late oocytes is maintained by continual active transport. Dev. Cell 11, 251–262 (2006).
Jaramillo, A.M., Weil, T.T., Goodhouse, J., Gavis, E.R. & Schupbach, T. The dynamics of fluorescently labeled endogenous gurken mRNA in Drosophila. J. Cell Sci. 121, 887–894 (2008).
Rook, M.S., Lu, M. & Kosik, K.S. CaMKIIalpha 3′ untranslated region-directed mRNA translocation in living neurons: visualization by GFP linkage. J. Neurosci. 20, 6385–6393 (2000).
Dynes, J.L. & Steward, O. Dynamics of bidirectional transport of Arc mRNA in neuronal dendrites. J. Comp. Neurol. 500, 433–447 (2007).
Chubb, J.R., Trcek, T., Shenoy, S.M. & Singer, R.H. Transcriptional pulsing of a developmental gene. Curr. Biol. 16, 1018–1025 (2006).
Mili, S., Moissoglu, K. & Macara, I.G. Genome-wide screen reveals APC-associated RNAs enriched in cell protrusions. Nature 453, 115–119 (2008).
Golding, I. & Cox, E.C. RNA dynamics in live Escherichia coli cells. Proc. Natl. Acad. Sci. USA 101, 11310–11315 (2004).
Shav-Tal, Y. et al. Dynamics of single mRNPs in nuclei of living cells. Science 304, 1797–1800 (2004).
Haim, L., Zipor, G., Aronov, S. & Gerst, J.E. A genomic integration method to visualize localization of endogenous mRNAs in living yeast. Nat. Methods 4, 409–412 (2007).
Daigle, N. & Ellenberg, J. λN-GFP: an RNA reporter system for live-cell imaging. Nat. Methods 4, 633–636 (2007). A keenly awaited alternative to MS2 coat protein–GFP tagging; it creates the possibility of tracking two mRNAs in the same cells: one using MS2 hairpin and MS2 coat protein–GFP fusion and the other using boxB RNA motif and λ N -RFP fusion.
Lange, S. et al. Simultaneous transport of different localized mRNA species revealed by live-cell imaging. Traffic 9, 1256–1267 (2008).
Calapez, A. et al. The intranuclear mobility of messenger RNA binding proteins is ATP dependent and temperature sensitive. J. Cell Biol. 159, 795–805 (2002).
Fusco, D. et al. Single mRNA molecules demonstrate probabilistic movement in living Mammalian cells. Curr. Biol. 13, 161–167 (2003).
Kerppola, T.K. Visualization of molecular interactions by fluorescence complementation. Nat. Rev. Mol. Cell Biol. 7, 449–456 (2006).
Rackham, O. & Brown, C.M. Visualization of RNA-protein interactions in living cells: FMRP and IMP1 interact on mRNAs. EMBO J. 23, 3346–3355 (2004). long with references 24 and 25, this paper describes the split-GFP approach: only GFP that is bound to the mRNA yields fluorescence, whereas the excess and free GFP does not produce any signal.
Valencia-Burton, M., McCullough, R.M., Cantor, C.R. & Broude, N.E. RNA visualization in live bacterial cells using fluorescent protein complementation. Nat. Methods 4, 421–427 (2007).
Ozawa, T., Natori, Y., Sato, M. & Umezawa, Y. Imaging dynamics of endogenous mitochondrial RNA in single living cells. Nat. Methods 4, 413–419 (2007).
Magliery, T.J. et al. Detecting protein-protein interactions with a green fluorescent protein fragment reassembly trap: scope and mechanism. J. Am. Chem. Soc. 127, 146–157 (2005).
Anderie, I. & Schmid, A. In vivo visualization of actin dynamics and actin interactions by BiFC. Cell Biol. Int. 31, 1131–1135 (2007).
Morrison, L.E., Halder, T.C. & Stols, L.M. Solution-phase detection of polynucleotides using interacting fluorescent labels and competitive hybridization. Anal. Biochem. 183, 231–244 (1989).
Li, Q., Luan, G., Guo, Q. & Liang, J. A new class of homogeneous nucleic acid probes based on specific displacement hybridization. Nucleic Acids Res. 30, E5 (2002).
Sixou, S. et al. Intracellular oligonucleotide hybridization detected by fluorescence resonance energy transfer (FRET). Nucleic Acids Res. 22, 662–668 (1994).
Cardullo, R.A., Agrawal, S., Flores, C., Zamecnik, P.C. & Wolf, D.E. Detection of nucleic acid hybridization by nonradiative fluorescence resonance energy transfer. Proc. Natl. Acad. Sci. USA 85, 8790–8794 (1988).
Tsuji, A. et al. Direct observation of specific messenger RNA in a single living cell under a fluorescence microscope. Biophys. J. 78, 3260–3274 (2000).
Sando, S. & Kool, E.T. Imaging of RNA in bacteria with self-ligating quenched probes. J. Am. Chem. Soc. 124, 9686–9687 (2002).
Abe, H. & Kool, E.T. Flow cytometric detection of specific RNAs in native human cells with quenched autoligating FRET probes. Proc. Natl. Acad. Sci. USA 103, 263–268 (2006).
Tyagi, S. & Kramer, F.R. Molecular beacons: probes that fluoresce upon hybridization. Nat. Biotechnol. 14, 303–308 (1996).
Tyagi, S., Bratu, D.P. & Kramer, F.R. Multicolor molecular beacons for allele discrimination. Nat. Biotechnol. 16, 49–53 (1998).
Bratu, D.P., Cha, B.J., Mhlanga, M.M., Kramer, F.R. & Tyagi, S. Visualizing the distribution and transport of mRNAs in living cells. Proc. Natl. Acad. Sci. USA 100, 13308–13313 (2003). Using rigorous cell biological and probe-based controls, this report shows that endogenous mRNAs can be imaged and tracked in live cells using nuclease-resistant molecular beacons.
Santangelo, P.J., Nix, B., Tsourkas, A. & Bao, G. Dual FRET molecular beacons for mRNA detection in living cells. Nucleic Acids Res. 32, e57 (2004).
Wang, W. et al. Imaging and characterizing influenza A virus mRNA transport in living cells. Nucleic Acids Res. 36, 4913–4928 (2008).
Tyagi, S. & Alsmadi, O. Imaging native beta-actin mRNA in motile fibroblasts. Biophys. J. 87, 4153–4162 (2004).
Santangelo, P., Nitin, N., LaConte, L., Woolums, A. & Bao, G. Live-cell characterization and analysis of a clinical isolate of bovine respiratory syncytial virus, using molecular beacons. J. Virol. 80, 682–688 (2006).
Santangelo, P.J. & Bao, G. Dynamics of filamentous viral RNPs prior to egress. Nucleic Acids Res. 35, 3602–3611 (2007).
Kloc, M. et al. Potential structural role of non-coding and coding RNAs in the organization of the cytoskeleton at the vegetal cortex of Xenopus oocytes. Development 132, 3445–3457 (2005).
Cui, Z.Q. et al. Visualizing the dynamic behavior of poliovirus plus-strand RNA in living host cells. Nucleic Acids Res. 33, 3245–3252 (2005).
Vargas, D.Y., Raj, A., Marras, S.A., Kramer, F.R. & Tyagi, S. Mechanism of mRNA transport in the nucleus. Proc. Natl. Acad. Sci. USA 102, 17008–17013 (2005).
Yeh, H.Y., Yates, M.V., Mulchandani, A. & Chen, W. Visualizing the dynamics of viral replication in living cells via Tat peptide delivery of nuclease-resistant molecular beacons. Proc. Natl. Acad. Sci. USA 105, 17522–17525 (2008).
Politz, J.C., Browne, E.S., Wolf, D.E. & Pederson, T. Intranuclear diffusion and hybridization state of oligonucleotides measured by fluorescence correlation spectroscopy in living cells. Proc. Natl. Acad. Sci. USA 95, 6043–6048 (1998).
Molenaar, C., Abdulle, A., Gena, A., Tanke, H.J. & Dirks, R.W. Poly(A)+ RNAs roam the cell nucleus and pass through speckle domains in transcriptionally active and inactive cells. J. Cell Biol. 165, 191–202 (2004).
Siebrasse, J.P. et al. Discontinuous movement of mRNP particles in nucleoplasmic regions devoid of chromatin. Proc. Natl. Acad. Sci. USA 105, 20291–20296 (2008).
Svanvik, N., Westman, G., Wang, D. & Kubista, M. Light-up probes: thiazole orange-conjugated peptide nucleic acid for detection of target nucleic acid in homogeneous solution. Anal. Biochem. 281, 26–35 (2000).
Mhlanga, M.M., Vargas, D.V., Fung, C., Kramer, F.R. & Tyagi, S. tRNA-linked molecular beacons for imaging mRNAs in the cytoplasm of living cells. Nucleic Acids Res. 33, 1902–1912 (2005).
Walev, I. et al. Delivery of proteins into living cells by reversible membrane permeabilization with streptolysin-O. Proc. Natl. Acad. Sci. USA 98, 3185–3190 (2001).
Chen, A.K., Behlke, M.A. & Tsourkas, A. Efficient cytosolic delivery of molecular beacon conjugates and flow cytometric analysis of target RNA. Nucleic Acids Res. 36, e69 (2008).
Melikov, K. & Chernomordik, L.V. Arginine-rich cell penetrating peptides: from endosomal uptake to nuclear delivery. Cell. Mol. Life Sci. 62, 2739–2749 (2005).
Nitin, N., Santangelo, P.J., Kim, G., Nie, S. & Bao, G. Peptide-linked molecular beacons for efficient delivery and rapid mRNA detection in living cells. Nucleic Acids Res. 32, e58 (2004).
Leonetti, J.P., Mechti, N., Degols, G., Gagnor, C. & Lebleu, B. Intracellular distribution of microinjected antisense oligonucleotides. Proc. Natl. Acad. Sci. USA 88, 2702–2706 (1991).
Fisher, T.L., Terhorst, T., Cao, X. & Wagner, R.W. Intracellular disposition and metabolism of fluorescently-labeled unmodified and modified oligonucleotides microinjected into mammalian cells. Nucleic Acids Res. 21, 3857–3865 (1993). This early paper shows that deoxyoligonucleotides with natural phosphodiester backbone are sequestered in the nucleus and are degraded rapidly, and points to backbone modifications that improve the intracellular stability of oligonucleotides.
Walder, R.Y. & Walder, J.A. Role of RNase H in hybrid-arrested translation by antisense oligonucleotides. Proc. Natl. Acad. Sci. USA 85, 5011–5015 (1988).
Hoke, G.D. et al. Effects of phosphorothioate capping on antisense oligonucleotide stability, hybridization and antiviral efficacy versus herpes simplex virus infection. Nucleic Acids Res. 19, 5743–5748 (1991).
Wu, Y., Yang, C.J., Moroz, L.L. & Tan, W. Nucleic acid beacons for long-term real-time intracellular monitoring. Anal. Chem. 80, 3025–3028 (2008).
Luebke, K.J., Balog, R.P. & Garner, H.R. Prioritized selection of oligodeoxyribonucleotide probes for efficient hybridization to RNA transcripts. Nucleic Acids Res. 31, 750–758 (2003).
Bratu, D.P. Molecular beacons: fluorescent probes for detection of endogenous mRNAs in living cells. Methods Mol. Biol. 319, 1–14 (2006).
Tsien, R.Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Lakowicz, J.R. Principles of Fluorescence Spectroscopy. (Kluwer Academic/Plenum Publishers, New York, 1999).
Babendure, J.R., Adams, S.R. & Tsien, R.Y. Aptamers switch on fluorescence of triphenylmethane dyes. J. Am. Chem. Soc. 125, 14716–14717 (2003).
Szent-Gyorgyi, C. et al. Fluorogen-activating single-chain antibodies for imaging cell surface proteins. Nat. Biotechnol. 26, 235–240 (2008).
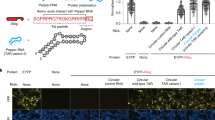
Sparano, B.A. & Koide, K. Fluorescent sensors for specific RNA: a general paradigm using chemistry and combinatorial biology. J. Am. Chem. Soc. 129, 4785–4794 (2007). Description of a new approach for RNA imaging using an aptamer that binds to a cell-permeant dye and renders the aptamer dye fluorescent.
Sando, S., Narita, A. & Aoyama, Y. Light-up Hoechst-DNA aptamer pair: generation of an aptamer-selective fluorophore from a conventional DNA-staining dye. ChemBioChem 8, 1795–1803 (2007).
Narita, A. Development of new sensing technologies toward non-invasive nucleic acid analysis. Thesis, Koyoto Univ. (2008).
Grate, D. & Wilson, C. Laser-mediated, site-specific inactivation of RNA transcripts. Proc. Natl. Acad. Sci. USA 96, 6131–6136 (1999).
Shaner, N.C. et al. Improving the photostability of bright monomeric orange and red fluorescent proteins. Nat. Methods 5, 545–551 (2008).
Richards, C.I. et al. Oligonucleotide-stabilized Ag nanocluster fluorophores. J. Am. Chem. Soc. 130, 5038–5039 (2008).
Kim, J.H., Morikis, D. & Ozkan, M. Adaptation of inorganic quantum dots for stable molecular beacons. Sensors and Actuators B 102, 315–319 (2004).
Cady, N.C., Strickland, A.D. & Batt, C.A. Optimized linkage and quenching strategies for quantum dot molecular beacons. Mol. Cell. Probes 21, 116–124 (2007).
Weissleder, R., Tung, C.H., Mahmood, U. & Bogdanov, A. Jr., In vivo imaging of tumors with protease-activated near-infrared fluorescent probes. Nat. Biotechnol. 17, 375–378 (1999).
So, M.K., Gowrishankar, G., Hasegawa, S., Chung, J.K. & Rao, J. Imaging target mRNA and siRNA-mediated gene silencing in vivo with ribozyme-based reporters. ChemBioChem 9, 2682–2691 (2008).
Andou, T., Endoh, T., Mie, M. & Kobatake, E. RNA detection using peptide-inserted Renilla luciferase. Anal. Bioanal. Chem. 393, 661–668 (2009).
Marras, S.A., Kramer, F.R. & Tyagi, S. Efficiencies of fluorescence resonance energy transfer and contact-mediated quenching in oligonucleotide probes. Nucleic Acids Res. 30, e122 (2002).
Bernacchi, S. & Mely, Y. Exciton interaction in molecular beacons: a sensitive sensor for short range modifications of the nucleic acid structure. Nucleic Acids Res. 29, e62 (2001).
Johansson, M.K., Fidder, H., Dick, D. & Cook, R.M. Intramolecular dimers: a new strategy to fluorescence quenching in dual-labeled oligonucleotide probes. J. Am. Chem. Soc. 124, 6950–6956 (2002).
Acknowledgements
I thank S. Marras of Public Health Research Institute for assistance with figures. The author's research is supported by National Institute of Mental Health grant MH079197.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
S.T. and his institution receive royalties from sale and use of molecular beacons.
Rights and permissions
About this article
Cite this article
Tyagi, S. Imaging intracellular RNA distribution and dynamics in living cells. Nat Methods 6, 331–338 (2009). https://doi.org/10.1038/nmeth.1321
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1321
This article is cited by
-
Binding and stabilizating effect of RNA triplex poly(U)⋅poly(A)*poly(U) by enantiomers of ruthenium(II) polypyridyl complex [Ru(bpy)2(dppx)]2+
JBIC Journal of Biological Inorganic Chemistry (2023)
-
Functional nucleic acid-based cell imaging and manipulation
Science China Chemistry (2021)
-
Expanding detection windows for discriminating single nucleotide variants using rationally designed DNA equalizer probes
Nature Communications (2020)
-
Visualizing RNA dynamics in live cells with bright and stable fluorescent RNAs
Nature Biotechnology (2019)
-
Real-time analysis of quantum dot labeled single porcine epidemic diarrhea virus moving along the microtubules using single particle tracking
Scientific Reports (2019)