Abstract
The functional role of protein phosphorylation is impacted by its fractional stoichiometry. Thus, a comprehensive strategy to study phosphorylation dynamics should include an assessment of site stoichiometry. Here we report an integrated method that relies on phosphatase treatment and stable-isotope labeling to determine absolute stoichiometries of protein phosphorylation on a large scale. This approach requires the measurement of only a single ratio relating phosphatase-treated and mock-treated samples. Using this strategy we determined stoichiometries for 5,033 phosphorylation sites in triplicate analyses from Saccharomyces cerevisiae growing through mid-log phase. We validated stoichiometries at ten sites that represented the full range of values obtained using synthetic phosphopeptides and found excellent agreement. Using bioinformatics, we characterized the biological properties associated with phosphorylation sites with vastly differing absolute stoichiometries.
Similar content being viewed by others
Main
Reversible protein phosphorylation has an important role in biological systems and is involved in virtually every cellular function. Mass spectrometry–based proteomics can be used to globally characterize this widespread post-translational modification and has been extensively reviewed1,2,3. In combination with stable-isotope labeling, mass spectrometry–based proteomics has produced large-scale datasets with quantification of phosphorylation changes between cell states1,4,5. Current large-scale comparisons provide an informative view of the protein-phosphorylation landscape1,6,7,8,9. However, their biological interpretation is complex. If the difference in protein abundance is known, a change in the extent of a site's phosphorylation results in a known relative stoichiometry change10, but the absolute fractional occupancy remains unknown. For example, a twofold downregulation of site stoichiometry could result from either fractional occupancy changes of 0.2% to 0.1% or 100% to 50%, which likely represent fundamentally different cellular strategies. Therefore, it appears crucial to assess absolute phosphorylation site occupancy on a proteome scale to correctly and comprehensively understand its functional importance.
Conventionally, biochemical methods, such as western blotting, have been used to measure phosphorylation stoichiometry. Phosphoproteins and non-phosphoproteins are separated physically via sodium dodecyl sulfate polyacrylamide gel electrophoresis, and their quantities are estimated using antibodies11,12. This method is time-consuming and requires a phosphorylation-induced migration difference in the gel. Protein phosphorylation stoichiometry can also be measured by mass spectrometry9,13,14. For example, using a label-free approach, stoichiometry can be measured from the ratios of ion signals of phosphopeptides and the corresponding non-phosphopeptides14,15,16,17. An assumption of this method is that differences in the ionization and detection efficiencies of a peptide's phosphorylated and nonphosphorylated forms are negligible. To overcome this shortcoming, a recently reported method18 has been used to determine the response ratios of phosphopeptides and non-phosphopeptides using synthetic peptide standards, and to measure the stoichiometries of two tyrosine residues in the Lyn protein.
Another mass spectrometry–based method, termed absolute quantification (AQUA)13, is based on stable-isotope dilution. Heavy versions of phosphorylated and non-phosphorylated peptides are synthesized and spiked into the sample in known quantities as internal standards. The phosphorylation stoichiometry can be obtained by measuring and comparing the absolute abundance of each peptide's phosphorylated and nonphosphorylated forms. Notably, using this method, we have demonstrated that Ser1526 from human separase is maintained at full occupancy until the metaphase-to-anaphase transition when it is partially dephosphorylated and activated, allowing the release of tethered sister chromatids. Knowing the stoichiometry of Ser1526 across the cell cycle proved vital to understanding its specialized mitotic function19. In addition to separase, this method has been adapted to measure occupancy in Akt20.
Another elegant strategy has been reported to measure phosphorylation stoichiometry by mass spectrometry21. Phosphatase treatment has been used in combination with specific and differential labeling of the N termini of all peptides in a sample with either a D5 or D0–propionyl group, followed by measuring the ratio of the abundance of the D5-D0 peptide pairs simultaneously using matrix-assisted laser desorption/ionization–mass spectrometry (MALDI-MS). Stoichiometry is obtained based on the signal increase of the peptide from the dephosphorylation of the corresponding phosphopeptide. A similar strategy22 has been used to measure the phosphorylation stoichiometries of different sites of protein Npr1 by protease-dependent incorporation of 18O- or 16O-labeled peptides, followed by phosphopeptide enrichment, phosphatase treatment and MALDI-MS analysis. This strategy has also been used in other reports23,24,25,26,27,28.
All of the methods discussed above have been used for small-scale studies, ranging from one to several proteins or a protein complex. With the development of mass spectrometry–based proteomics methods, thousands of unique phosphorylation sites can be analyzed in one experiment, yet the underlying fractional occupancy of these sites remains unknown. Recently, a method9 has been reported to measure site stoichiometry on a large scale by obtaining a minimum of three different ratios representing protein, phosphopeptide and unmodified peptide changes based on stable-isotope labeling with amino acids in cell culture (SILAC). But this approach is biased in that it can only be applied to sites after detecting a change in phosphopeptide levels.
In this study, we integrated phosphatase treatment and stable-isotope labeling to determine site stoichiometries of protein phosphorylations on a large scale. The method can provide thousands of stoichiometry measurements for a single cellular condition. We measured basal stoichiometry levels for more than 5,000 events in exponentially growing yeast in triplicate. We assessed the accuracy of this method for ten sites of different stoichiometries using the AQUA strategy13. Bioinformatic analyses indicated that acidic sites were, on average, of higher stoichiometry than other motifs and that high-stoichiometry sites were not more conserved across yeast species than low-stoichiometry sites. Several biological and functional categories were statistically enriched for high or low stoichiometry.
Results
A method to assess absolute stoichiometry on a large scale
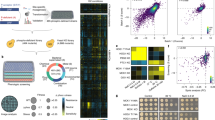
An overview of the method is shown in Figure 1. In this method, a cell lysate is first digested with endoproteinase lys-C. Two identical 0.5-mg peptide aliquots are subjected to either phosphatase treatment or a mock reaction. After the reactions, peptides in the phosphatase-treated sample are chemically labeled by reductive dimethylation using deuteroformaldehyde to dimethylate free amines29. The mock reaction sample is chemically labeled using formaldehyde. The two aliquots are then mixed, which results in a 1:1 ratio for all peptides unaffected by phosphatase treatment. The mixed sample is then analyzed by deep mass spectrometry sequencing to identify and quantify as many peptide species as possible. This involved separation by hydrophilic interaction liquid chromatography and then analysis of each fraction by liquid chromatography–tandem mass spectrometry (LC-MS/MS) techniques. Among the thousands of detected peptides will be those that were previously phosphorylated. Such peptides will display and increase in the heavy partner intensity over the co-eluting light version, directly representing the fraction of the peptide that was phosphorylated. Each of these peptides is termed an 'occupancy-determining peptide' (ODP), and they directly encode the absolute stoichiometry based on the ratio of heavy/light species ((1 – 1/ratio) × 100%). A list of ODPs can be collected from previously published datasets of site-specific phosphorylation. The overlap of these published datasets with the phosphatase-treated one defines the number of sites for which stoichiometries can be determined.
Two identical aliquots of a proteolyzed protein lysate are either mock- or phosphatase-treated followed by differential chemical labeling with stable isotopes. After mixing, fractional occupancy is encoded in a single ratio comparing the nonphosphorylated form of a peptide with and without phosphatase treatment. P, phosphorylation site. Peptide sequences obtained are then examined against a database of known sites from the literature. We term these overlapping, nonphosphorylated forms ODPs. Based on the ratio of the ODPs, stoichiometries are calculated.
Applying the method to define absolute stoichiometries
We evaluated the potential for this method by using it to define the absolute stoichiometries of yeast growing through log phase with glucose as a carbon source. We grew three separate yeast cultures and isolated protein. We performed the experiments as described in Figure 1 with deep mass spectrometry sequencing of 20 fractions for each experiment. The total numbers of peptides detected in cultures 1–3 were 76,893; 77,439 and 81,012, respectively (Supplementary Tables 1,2,3 and 4). We calculated false discovery rates based on the target-decoy database approach30. In a separate experiment, the phosphatase reaction performed at the peptide level was >99% effective (Online Methods).
To use this method, we needed a list of phosphorylation sites. We used localized sites determined in five published yeast phosphorylation studies7,10,31,32,33. We assessed the overlap between our datasets for phosphatase-treated samples and the sequences surrounding these published sites. Owing to the shotgun nature of LC-MS/MS, not every ODP is identified. In this proof-of-principle experiment, for 5,033 sites reported in the literature, we detected the nonphosphorylated peptide forms. We plotted their heavy/light species ratios (Fig. 2a), and show an example of the occupancy calculation from the measured ratio and stoichiometry distribution (Fig. 2b,c). Very high occupancy (>90%) was not common, occurring at only about one in ten sites. The majority of sites had low occupancy, with half displaying occupancies of <30%. Some examples with different motifs are shown in Supplementary Table 5.
(a) Three biological yeast replicates were grown to mid-log phase. Lysates were proteolyzed with endoproteinase lys-C. An identical aliquot of each proteolyzed lysate was either mock- or phosphatase-treated as illustrated in Figure 1. ODPs (5,033 peptides) were identified based on the overlap with sites in five published studies7,10,31,32,33. Shown is a scatter plot of all ODP ratios. Ratios directly encode fractional occupancies (orange lines). S/N, signal-to-noise ratio. CIP, calf intestinal phosphatase. (b) Example of an ODP (EAENDEDS*EVNAK, where the asterisk indicates the phosphorylated serine) from protein Bbc1, with signals from CIP-treated (yellow) and untreated (blue) samples highlighted. (c) Site stoichiometry distribution for 5,033 events from wild-type yeast undergoing exponential growth.
One limitation of this method stems from the fact that bottom-up proteomics methods measure ratios for individual peptides whose sequences can contain one or more phosphorylation sites. Although the ratios observed for singly phosphorylated peptides represent stoichiometries for individual sites, peptides containing multiple phosphorylation sites reflect the extent of modification for all phosphorylation sites in any combination. Our mass spectrometry–based method cannot be used to distinguish the contribution of each site in these multiply modified peptides. For simplicity, we assigned the stoichiometries calculated from the peptide ratio to each site in the peptide. Thus, these stoichiometries should be considered as maximum stoichiometries when more than one site is known on the peptide. We identified 4,016 sites (80%) from peptides displaying only a single site (Supplementary Table 6).
To assess the reproducibility of the method, we performed biological triplicate experiments and obtained stoichiometries for 3,843, 4,077 and 3,756 events. The Pearson correlations between replicates was strong, as shown for experiments 1 and 2, and 1 and 3 (Supplementary Fig. 1). For 2,664 site stoichiometries measured in all experiments, the average s.d. was 6.3% (Supplementary Fig. 1c). Using one s.d., stoichiometries were accurate to ± 6%. Values could be made even more accurate by increasing the number of biological replicates. However, these measurements were accurate enough for site classification, allowing biological characterization.
Validation of absolute site stoichiometries using AQUA
To validate our stoichiometry measurements, we used another independent method, the AQUA strategy13, to accurately determine the stoichiometries of ten sites chosen to represent the full range of occupancy states (Fig. 3 and Table 1). We synthesized ten pairs of heavy phosphopeptides and corresponding non-phosphopeptides. Into a proteolyzed whole-cell lysate, we spiked known amounts of heavy peptides and analyzed the samples by LC-MS/MS (Online Methods). We analyzed the extracted chromatographic peaks of non-phosphopeptide (RIIEHSDVENENVK) and phosphopeptide (RIIEHS*DVENENVK) ions for site Ser618 of the protein UBP1, a ubiquitin-specific protease that removes ubiquitin from conjugated proteins, and identified the corresponding phosphopeptide by MS/MS (Fig. 3a,b). According to the heavy/light species ratio and the amount of heavy peptides spiked into the sample, we calculated the amount of the peptide and phosphopeptide in the lysate to be 20.5 fmol and 46.0 fmol, respectively. Thus, the stoichiometry of this site was 69.2%, which agrees with the values of 59%, 59% and 66% in the biological triplicate experiments. The stoichiometries of sites Ser266 of IPP1 and Thr710 of DCP2 were very low based on the results of our large-scale experiments; we confirmed this by AQUA, which yielded stoichiometries of 1.8% and 2%, respectively. Additionally, acidic sites, Ser562 of YML093W and Ser379 of BFR2, had full or nearly full occupancies (Table 1). The AQUA results were again consistent. The ten site stoichiometries quantified by AQUA strongly suggest that our large-scale method for determination of phosphorylation site occupancy is reliable.
(a,b) Synthetic peptides were generated representing heavy phosphorylated and nonphosphorylated versions of the peptide. The synthetic peptides were spiked into proteolyzed lysates and separated by LC-MS/MS techniques. Using the heavy peptides as internal standards, the light versions were quantified and stoichiometries were calculated. An example of the measured amount of peptide (RIIEHSDVENENVK) and phosphopeptide (RIIEHS*DVENENVK, where the asterisk indicates the phosphorylation site) by AQUA in the protein UBP1 (a). The calculated stoichiometry was 69.2% (= 46.0/(20.5 + 46.0) × 100). An example of phosphopeptide identification (RIIEHS*DVENENVK) by MS/MS (b). Xcorr is the primary score from the Sequest algorithm.
Relationship between kinase motifs and site stoichiometry
We next examined the relationship between specific sequence motifs favored by kinases and the resulting site stoichiometries. Phosphorylation sites containing an acidic (casein kinase II–like) motif had higher occupancy than other motifs (Fig. 4a). Proline-directed sites, which are often modified by cyclin-dependent kinase (CDK) and mitogen-activated protein kinase (MAPK), had lower than average occupancies.
(a) Analysis of phosphorylation events in indicated motifs. (b) Analysis of phosphorylation events in ordered and disordered regions. (c,d) Clustering of the phosphoproteins according to their highest stoichiometry site based on their enrichment in specific cell compartments (c) and biological processes (d). Categories without a P value were assigned a conservative value of 1. The P values were log-transformed and then z-transformed. Phosphoproteins were then grouped based on their z scores via hierarchical clustering.
For each site with occupancy, we also applied secondary-structure prediction algorithms34 to classify events according to likely structure. Most sites (91%) were predicted to be in disordered regions (Fig. 4b). However, sites predicted to be in ordered regions had a strong preference for low stoichiometry. As we have shown previously7, phosphorylation is overwhelmingly detected in protein regions predicted or known to be disordered. Yet when phosphorylation does occur in ordered regions, we found that it is almost always of low occupancy in exponentially growing yeast. This suggests that (i) the frequency of high-occupancy sites serving structural roles is low and (ii) phosphorylation events occurring in ordered regions are nearly always activating in nature. Indeed, the best example of this phenomenon is that phosphorylation in the kinase domains is often required for kinase activation35.
Correlating protein function to stoichiometry levels
To obtain functional insights into site stoichiometry, we separated phosphoproteins into four groups based on their highest stoichiometries (low (<10%), medium (10–30%), high (30–75%) and very high (75–100%)) to examine differences in biological processes and cellular compartment enrichment using the database for annotation, visualization and integrated discovery (DAVID)36. We z-transformed P values and then clustered them (Fig. 4c,d). Gene Ontology biological processes enriched in high-stoichiometry sites included 'chromatin silencing' and 'cytokinesis during cell cycle'. Full versions of these figures are shown in Supplementary Figure 2.
Next we examined site conservation across 25 fungal proteomes for thousands of sites (Fig. 5 and Supplementary Fig. 3). We identified homologs from 25 fungal genomes from the same family, Saccharomycetaceae, using a Tblastn search (search translated nucleotide databases using a protein query) and then aligned the sequences. If a homolog was found, we determined conservation based on identifying the same phospho-acceptor residue at the same position in both species (Online Methods). To facilitate this analysis, we selected only singly phosphorylated sites with high-confidence localizations (3,443 sites), and we binned stoichiometries into three categories of low, medium and high occupancies. Site residue conservation was generally localized to only a few very closely related species. However, ∼10% of sites were highly conserved. Previously, we had found a similarly poor conservation of 541 Cdk1-phosphorylated residues across fungal species7. Notably, residues in high-occupancy sites were actually less conserved, on average, than residues in low-occupancy ones. Most sites were poorly conserved even in budding yeast species, and we did not find high-occupancy sites associated with more conserved residues. Indeed, the most conserved site residues had very-low-stoichiometry sites. These most conserved sites were also the ones predicted to be in ordered regions (data not shown). We created a website for visualizing these conservation data and the alignments (http://gygi.med.harvard.edu/pubs/occupancy_evolution/).
Sites were subgrouped by fractional occupancy into high, medium and low sets, and then clustered based on overall conservation levels. Each column represents a single site residue. If a homolog was identified and the site residue was conserved, the corresponding cell is yellow otherwise it is blue. If no homolog was identified, the cell is black.
Discussion
Our method of determining absolute site stoichiometry provides unbiased insight into this functionally important protein property for thousands of basally phosphorylated sites. The approach requires only the measurement of a single ratio in a single experiment for ODPs. The stoichiometry relationship is straightforward to determine for singly phosphorylated peptides. Determining stoichiometries for multiply phosphorylated peptides is more complex and may not accurately reflect values at individual sites, but the results still represent occupancy levels encompassing the sequence region shown. They should be considered to represent the maximum potential occupancy at each site. Of the 5,033 stoichiometry measurements we collected, 1,017 were derived from peptides in which multiple sites were present.
We averaged our stoichiometry measurements from three separate yeast cultures. The s.d. of these measurements was ∼6%. This implies that the method is not suitable for distinguishing small differences in stoichiometry but can be used to classify sites into at least 5–10 categories of increasing stoichiometry. Also, in our validation of the absolute occupancy state for ten sites by AQUA13, the agreement between the AQUA stoichiometries and the phosphatase-based method was strong.
A special stoichiometry case is full occupancy, which predicts constitutive kinase activity and/or little phosphatase activity at the site. Many cyclin-dependent kinase sites have been shown to achieve full occupancy under mitotic conditions9,19. However, we collected our measurements from asynchronous yeast in which the vast majority of cells would not be in mitosis. Thus, most targets of CDK (mitotic sites) would be expected to be detected at low stoichiometry. We found that sites determined in that study7 to be substrates for Cdk1 and overlapping with ours (95 sites) contained a similar distribution of stoichiometries as the entire phosphoproteome with many high and low stoichiometry events (mean = 40.6%; Supplementary Fig. 4 and Supplementary Table 7). This indicates that many of these 'mitotic' sites are actually phosphorylated during other phases of the cell cycle. They likely have additional, nonmitotic roles, which are then overwhelmed or repurposed during mitosis.
During exponential growth in asynchronous cultures, we found that ∼10% of sites had full or almost full occupancy (>90%) (Fig. 2c). This finding has important implications. It suggests that kinase pathways are generally inactive or less active in their default state and that most phosphorylation events function by influencing only a fraction of the available protein molecules. Maintaining sites at full occupancy thus requires considerable energy investment. Casein kinase II (CKII) is a constitutively active and essential yeast kinase with hundreds of substrates37. Strong casein kinase II motifs contain acidic residues at the +1 and +3 positions (for example SerAspXxxGlu, in which Xxx denotes any amino acid). This acidic motif encompassed higher stoichiometries, on average, than basophilic or proline-directed motifs, demonstrating that strong or even constitutive pathway activation can result in high basal stoichiometries.
Only one other method has been reported to our knowledge that can be used to assess absolute site stoichiometry on a large or global scale9. The authors determined occupancy by obtaining a minimum of three different SILAC ratios representing protein, phosphopeptide and unmodified peptide changes. This is accomplished in separate proteome- and phosphorylation-based analyses. Using these three ratios from two experiments, a fractional occupancy level is calculated. There is a requirement that a substantial change in phosphopeptide amount occurs. In addition, because of the complex math involved, even slight errors in measurement can cause wildly changing stoichiometry calculations, and there are many undefined possibilities among the three ratios. For example, a change of any amount at the protein level with no corresponding change at the phosphopeptide level results in a change in occupancy but is undefined by their method.
A major question is how stoichiometry as a biological property might influence protein function or regulation. We analyzed differences in the enrichment of Gene Ontology categories for low, medium, high and very high stoichiometries. This provided hints of where high- and low-occupancy may be important for biological function. Phosphoproteins with reported localization to the cytoplasm, ribosome and mitochondria generally had lower stoichiometries whereas proteins in the nucleus and cellular buds were enriched in high phosphorylation occupancies. Nuclear and budding proteins are enriched in regulatory proteins, suggesting fine regulatory control. Proteins in many biological processes such as 'cytokinesis during cell cycle' and 'chromatin silencing' were enriched with high stoichiometries, but phosphorylation site stoichiometries of proteins related to, for example, 'mitochondrial organization' and 'RNA transport' were enriched in low-stoichiometry sites. As high stoichiometry sites are rare, their association with biological processes reflect their important role in regulating these events. Examples of very-high-stoichiometry sites are shown in Supplementary Table 8.
Our analysis of 25 fungal species with sequenced genomes from the Saccharomycetaceae family, which reproduce based on 'budding', suggests that stoichiometry does not positively correlate with a site's biological essentialness. The strong lack of conservation in general and even in very closely related species implies that phosphorylation networks are set up such that (perhaps through multiple phosphorylation events per protein8) extensive evolutionary freedom even at previous sites of regulation is permissible.
Methods
Cell culture and lysis, and protein extraction and digestion.
The yeast strain used was BY4742 MATα, derived from S288c. Three biological replicate experiments were carried out in this work. Wild-type cells were grown in yeast peptone dextrose (YPD) medium. After ∼18 h, cell populations had undergone over ten doublings until they reached log phase (OD600 = 1.0). Yeast cells were collected and then resuspended at 4 °C in a buffer containing 50 mM Tris (pH 8.2), 8 M urea, 75 mM NaCl, 50 mM NaF, 50 mM β-glycerophosphate, 1 mM sodium orthovanadate, 10 mM sodium pyrophosphate and one protease inhibitor cocktail tablet (complete mini, EDTA-free, Roche) per 10 ml. Cells were lysed using the MiniBeadbeater (Biospec) for microcentrifuge tubes at maximum speed, four cycles of 60 s each, with 2 min pauses between cycles to avoid overheating of the lysates. After centrifugation, lysates were transferred to new tubes. The protein concentration in the lysate was determined using the BCA protein assay (Pierce) and proteins were subjected to disulfide reduction with 5 mM DTT (56 °C, 25 min) and alkylation with 15 mM iodoacetamide (room temperature (25 °C), 30 min in the dark). Excess iodoacetamide was quenched with 5 mM DTT (room temperature, 15 min in the dark). The denatured protein extract was then digested with 5 ng μl−1 lys-C (Wako) in 25 mM Tris (pH 8.8) with 1.5 M urea, at 37 °C for 15 h.
Peptide separation.
Peptide mixtures were acidified by addition of 10% trifluoroacetic acid (TFA) to a final concentration of 0.4%, centrifuged and desalted using a C18 SepPak cartridge (Waters). Purified peptides were separated using hydrophilic interaction chromatography (HILIC) into 20 fractions using a 4.6 mm × 250 mm TSKgel Amide-80 5-μm particle column (Tosoh Biosciences) with a 50-min gradient of 5%–28% H2O in acetonitrile (0.1% TFA).
LC-MS/MS analyses.
Dried peptides were dissolved in 5% acetonitrile and 4% formic acid, and 2 μl of the solution was loaded onto a microcapillary column packed with C18 beads (Magic C18AQ, 5 μm, 200 Å, 125 μm × 16 cm) using a Famos autosampler (LC Packings). Peptides were separated by reversed-phase chromatography using an Agilent 1100 binary pump with a 70-min gradient of 5–30% acetonitrile (in 0.125% formic acid) and detected in a hybrid dual-cell quadruple linear ion trap–orbitrap mass spectrometer (LTQ Orbitrap Velos, ThermoFisher) using a data-dependent Top20 method38. For each cycle, one full mass spectrometry scan in the Orbitrap at 3 × 106 automatic gain control (AGC) target was followed by up to 20 MS/MS scans in the LTQ for the most intense ions. Selected ions were excluded from further analysis for 30 s. Ions with charge +1 or unassigned were also rejected. Maximum ion accumulation times were 1,000 ms for each full mass spectrometry scan and 150 ms for MS/MS scans.
Database searches.
After acquisition of mass spectrometry data, Xcalibur .RAW format files were converted into mzXML format. Individual precursors selected for MS/MS fragmentation were checked using algorithms that detect and correct errors in monoisotopic peak assignment and refine precursor ion mass measurements. All MS/MS spectra were then exported as DTA files and searched using the Sequest algorithm39. Spectra were matched against a database containing sequences of all proteins in the yeast open reading frame database (6,607 entries) downloaded from the Saccharomyces Genome Database in both forward and reverse orientations. The following parameters were selected to identify peptides for protein expression experiments: 50 p.p.m. precursor mass tolerance; 1.0 Da product-ion mass tolerance; up to two missed cleavages; variable modifications: oxidation of methionine (+15.9949 Da); fixed modifications: carbamidomethylation of cysteine (+57.0214 Da). False discovery rates were controlled using the target-decoy strategy to distinguish correct and incorrect identifications30.
Peptide quantification.
Peptide quantification was performed using the Vista program40. We required a signal-to-noise ratio (S/N) value >3 for both heavy and light species for quantification. For peptides found exclusively as singlets (only heavy or only light peak present), we reported the peak S/N ratio or its inverse, as a proxy for relative abundance measurement. For such peptides, we required an S/N value >5 for the observed species. In addition, if the S/N value of one member of a pair was <3, the partner value was required to be >5. Finally, to avoid quantifying false positives, any identification from a singlet peak was required to pass a tenfold more stringent identification threshold (Q value < 0.001; precision > 99.9%). Raw abundance ratios from each experiment were normalized based on the median distribution ratio.
Phosphatase reaction.
Desalted peptides from each lys-C digest were split into two tubes equally and then dissolved in phosphatase reaction buffer (100 mM NaCl, 50 mM Tris-HCl, 10 mM MgCl2 and 1 mM DTT; pH 7.9). Calf intestinal phosphatase (1,000 units; CIP; New England BioLabs) was incubated with one tube to dephosphorylate the phosphopeptides. To the other tube, no CIP was added. The reaction proceeded for both tubes at 37 °C for 2 h. Reactions were quenched with TFA to final concentration of 0.5%.
To assess phosphatase efficiency, 0.5-mg aliquots from parallel mock- and phosphatase-treated samples were enriched for phosphopeptides using immobilized metal affinity chromatography as described previously41. LC-MS/MS analysis resulted in the identification of 720 phosphopeptides (no reverse hits) in the mock-treated sample. No phosphopeptides were detected in the treated sample at a 1% false discovery rate. It is important to note that reactions occurred on peptides and not proteins. We found no bias toward any sites at the peptide level with CIP.
Phosphorylation site stoichiometry.
Yeast were seeded, grown and collected in three separate experiments. For each replicate, lysate was proteolyzed with endoproteinase lys-C. Two identical 0.5-mg peptide aliquots were subjected to either phosphatase treatment or a mock reaction. After the reactions, peptides in the phosphatase-treated sample were chemically labeled by reductive dimethylation using deuteroformaldehyde to dimethylate free amines29. The mock reaction sample was chemically labeled by reductive dimethylation using formaldehyde. The two aliquots were then mixed, resulting in a 1:1 ratio for nearly all peptides analyzed by LC–mass spectrometry techniques. However, sequences bearing phosphorylation would result in an increase in their levels directly proportional to the amount tied up by phosphorylation.
Our method requires a dataset of phosphorylation sites to identify the ODPs. We used five published datasets7,10,31,32,33. Each of these presented large-scale analyses of yeast phosphorylation and, notably, localization information was also presented. We only used localized sites in these experiments. The localization was measured differently in each report. We used the following cutoff values for localization: sites with Phospho (STY) probabilities32 > 0.9; Ascore7,10,33 > 13 or PLScore31 > 10. Phosphorylation site stoichiometries were directly calculated according to the ratio of heavy/light peptides (H/L) ((1 – 1/ratio) × 100%). Values less than 1% were assigned to 1%.
Validation of site stoichiometries using AQUA.
Ten pairs of heavy phosphopeptides and corresponding non-phosphopeptides were synthesized replacing one residue with an isotopically enriched version (13C and 15N) (Cell Signaling Technology). A known amount of heavy phosphopeptides and non-phosphopeptides were spiked into the proteolyzed sample. For the non-phosphopeptide experiment, proteolyzed lysates were separated by HILIC into 20 fractions. In the phosphopeptide experiment, during purification using the C18 SepPak column, the peptides were eluted into five fractions using different concentrations of acetonitrile and followed by immobilized metal affinity chromatography enrichment of phosphopeptides. Finally, all samples were analyzed by the same LC-MS/MS techniques.
Secondary structure prediction.
The likely secondary structural environment and level of disorder was predicted for each phosphorylation site using VSL2 (ref. 34), which estimates the probability that each phosphorylation site was located in a region of high disorder. Those sites with disorder probabilities exceeding 0.5 were classified as 'disordered', whereas those with probabilities below 0.5 were classified as 'ordered'.
Clustering of phosphoproteins.
Phosphoproteins were grouped into four classes according to their highest stoichiometry site: that is, 0–10%, 10–30%, 30–75% and 75–100%. The enrichment analysis for Gene Ontology (GO) biological process (BP) and cellular compartment (CC) were performed separately for every class using DAVID36, and phosphoproteins identified by mass spectrometric proteomics method were used as background7,10,31,32,33. Categories without a P value were assigned a conservative value of 1. The P values were transformed by the function x = −log10(P), and then they were z-transformed. Phosphoproteins were then grouped based on their z-scores via hierarchical clustering (Euclidean distance, Centroid linkage) using Gene Cluster 3.0.
Calculation procedure of evolutionary conservation.
Whole genome DNA sequences for the organisms listed in Supplementary Figure 3 were retrieved from the US National Center for Biotechnology Information and then used in a Tblastn search with the appropriate Saccharomyces cerevisiae protein as the query. This was necessary as many of these organisms do not yet have fully annotated proteomes. These search results were then analyzed using in-house–developed software to examine the residue at the corresponding position in the subject's protein sequence to that of the phosphorylated residue from the S. cerevisiae protein. For comparison, only the top matching sequence from each organism was selected and when no sequence was found the conservation was designated as 'no homologous sequence found'. Because we confined our analysis to just fungal genomes, the alignment was easier. A residue was considered conserved if the corresponding residue position in the subject's sequence was a perfect match to that of S. cerevisiae. After this procedure was applied to each alignment, conservation information was partitioned into three occupancy categories (high, ≥80%; moderate, <80%, >20%; low, ≤20%) and then conservation data for each of these categories was clustered using Pearson correlation and visualized using the MultiExperiment viewer (http://www.tm4.org/mev/).
References
Grimsrud, P.A., Swaney, D.L., Wenger, C.D., Beauchene, N.A. & Coon, J.J. Phosphoproteomics for the masses. ACS Chem. Biol. 5, 105–119 (2010).
Macek, B., Mann, M. & Olsen, J.V. Global and site-specific quantitative phosphoproteomics: principles and applications. Annu. Rev. Pharmacol. Toxicol. 49, 199–221 (2009).
Thingholm, T.E., Jensen, O.N. & Larsen, M.R. Analytical strategies for phosphoproteomics. Proteomics 9, 1451–1468 (2009).
Boersema, P.J., Mohammed, S. & Heck, A.J.R. Phosphopeptide fragmentation and analysis by mass spectrometry. J. Mass Spectrom. 44, 861–878 (2009).
Witze, E.S., Old, W.M., Resing, K.A. & Ahn, N.G. Mapping protein post-translational modifications with mass spectrometry. Nat. Methods 4, 798–806 (2007).
Choudhary, C. et al. Mislocalized activation of oncogenic RTKs switches downstream signaling outcomes. Mol. Cell 36, 326–339 (2009).
Holt, L.J. et al. Global analysis of Cdk1 substrate phosphorylation sites provides insights into evolution. Science 325, 1682–1686 (2009).
Huttlin, E.L. et al. A tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–1189 (2010).
Olsen, J.V. et al. Quantitative phosphoproteomics reveals widespread full phosphorylation site occupancy during mitosis. Sci. Signal. 3, ra3 (2010).
Wu, R. et al. Correct interpretation of comprehensive phosphorylation dynamics requires normalization by protein expression changes. Mol. Cell. Proteomics published online, doi:10.1074/mcp.M111.009654 (7 May 2011).
Cooper, J.A. & Hunter, T. Identification and characterization of cellular targets for tyrosine protein kinases. J. Biol. Chem. 258, 1108–1115 (1983).
Stukenberg, P.T. et al. Systematic identification of mitotic phosphoproteins. Curr. Biol. 7, 338–348 (1997).
Gerber, S.A., Rush, J., Stemman, O., Kirschner, M.W. & Gygi, S.P. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl. Acad. Sci. USA 100, 6940–6945 (2003).
Steen, H., Jebanathirajah, J.A., Springer, M. & Kirschner, M.W. Stable isotope-free relative and absolute quantitation of protein phosphorylation stoichiometry by MS. Proc. Natl. Acad. Sci. USA 102, 3948–3953 (2005).
Carr, S.A., Huddleston, M.J. & Annan, R.S. Selective detection and sequencing of phosphopeptides at the femtomole level by mass spectrometry. Anal. Biochem. 239, 180–192 (1996).
Guo, L. et al. Studies of ligand-induced site-specific phosphorylation of epidermal growth factor receptor. J. Am. Soc. Mass Spectrom. 14, 1022–1031 (2003).
Steen, J.A.J. et al. Different phosphorylation states of the anaphase promoting complex in response to antimitotic drugs: a quantitative proteomic analysis. Proc. Natl. Acad. Sci. USA 105, 6069–6074 (2008).
Jin, L.L. et al. Measurement of protein phosphorylation stoichiometry by selected reaction monitoring mass spectrometry. J. Proteome Res. 9, 2752–2761 (2010).
Stemmann, O., Zou, H., Gerber, S.A., Gygi, S.P. & Kirschner, M.W. Dual inhibition of sister chromatid separation at metaphase. Cell 107, 715–726 (2001).
Atrih, A. et al. Stoichiometric quantification of Akt phosphorylation using LC-MS/MS. J. Proteome Res. 9, 743–751 (2010).
Zhang, X., Jin, Q.K., Carr, S.A. & Annan, R.S. N-terminal peptide labeling strategy for incorporation of isotopic tags: a method for the determination of site-specific absolute phosphorylation stoichiometry. Rapid Commun. Mass Spectrom. 16, 2325–2332 (2002).
Bonenfant, D. et al. Quantitation of changes in protein phosphorylation: a simple method based on stable isotope labeling and mass spectrometry. Proc. Natl. Acad. Sci. USA 100, 880–885 (2003).
Domanski, D., Murphy, L.C. & Borchers, C.H. Assay development for the determination of phosphorylation stoichiometry using multiple reaction monitoring methods with and without phosphatase treatment: application to breast cancer signaling pathways. Anal. Chem. 82, 5610–5620 (2010).
Hegeman, A.D., Harms, A.C., Sussman, M.R., Bunner, A.E. & Harper, J.F. An isotope labeling strategy for quantifying the degree of phosphorylation at multiple sites in proteins. J. Am. Soc. Mass Spectrom. 15, 647–653 (2004).
Johnson, H., Eyers, C.E., Eyers, P.A., Beynon, R.J. & Gaskell, S.J. Rigorous determination of the stoichiometry of protein phosphorylation using mass spectrometry. J. Am. Soc. Mass Spectrom. 20, 2211–2220 (2009).
Kanshin, E. et al. The stoichiometry of protein phosphorylation in adipocyte lipid droplets: Analysis by N-terminal isotope tagging and enzymatic dephosphorylation. Proteomics 9, 5067–5077 (2009).
Pflieger, D. et al. Quantitative proteomic analysis of protein complexes. Mol. Cell. Proteomics 7, 326–346 (2008).
Previs, M.J. et al. Quantification of protein phosphorylation by liquid chromatography-mass spectrometry. Anal. Chem. 80, 5864–5872 (2008).
Broberg, A. High-performance liquid chromatography/electrospray ionization ion-trap mass spectrometry for analysis of oligosaccharides derivatized by reductive amination and N,N-dimethylation. Carbohydr. Res. 342, 1462–1469 (2007).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Albuquerque, C.P. et al. A multidimensional chromatography technology for in-depth phosphoproteome analysis. Mol. Cell. Proteomics 7, 1389–1396 (2008).
Gnad, F. et al. High-accuracy identification and bioinformatic analysis of in vivo protein phosphorylation sites in yeast. Proteomics 9, 4642–4652 (2009).
Li, X. et al. Large-scale phosphorylation analysis of alpha-factor-arrested Saccharomyces cerevisiae. J. Proteome Res. 6, 1190–1197 (2007).
Peng, K., Radivojac, P., Vucetic, S., Dunker, A.K. & Obradovic, Z. Length-dependent prediction of protein intrinsic disorder. BMC Bioinformatics 7, 208 (2006).
Taylor, S.S., Knighton, D.R., Zheng, J.H., Teneyck, L.F. & Sowadski, J.M. Structural framework for the protein-kinase family. Annu. Rev. Cell Biol. 8, 429–462 (1992).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Guerra, B. & Issinger, O.G. Protein kinase CK2 in human diseases. Curr. Med. Chem. 15, 1870–1886 (2008).
Haas, W. et al. Optimization and use of peptide mass measurement accuracy in shotgun proteomics. Mol. Cell. Proteomics 5, 1326–1337 (2006).
Eng, J.K., McCormack, A.L. & Yates, J.R. An approach to correlate tandem mass-spectral data of peptides with amino-acid-sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Bakalarski, C.E. et al. The impact of peptide abundance and dynamic range on stable-isotope-based quantitative proteomic analyses. J. Proteome Res. 7, 4756–4765 (2008).
Villen, J. & Gygi, S.P. The SCX/IMAC enrichment approach for global phosphorylation analysis by mass spectrometry. Nat. Protoc. 3, 1630–1638 (2008).
Acknowledgements
This work was supported in part by US National Institutes of Health grants (HG3456) to S.P.G. We thank all members of the Gygi lab for help, especially R.A. Everley for his help with instrumentation and L. Ting for critically reading the manuscript.
Author information
Authors and Affiliations
Contributions
S.P.G. and R.W. designed the research. R.W., W.H., N.D., E.L.H., B.Z., M.E.S. and S.P.G. participated in the data generation, analysis and interpretation. R.W. and S.P.G. wrote the manuscript and all authors edited it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1 and 5 (PDF 381 kb)
Supplementary Table 2
Peptides identified in experiment 1. (XLSX 14310 kb)
Supplementary Table 3
Peptides identified in experiment 2. (XLSX 14521 kb)
Supplementary Table 4
Peptides identified in experiment 3. (XLSX 15000 kb)
Supplementary Table 6
Site stoichiometries obtained in biological triplicate experiments. (XLSX 584 kb)
Supplementary Table 7
Site stoichiometries for events described as Cdk1substrates. (XLSX 21 kb)
Supplementary Table 8
Examples of phosphorylation sites with high stoichiometries. (XLSX 11 kb)
Rights and permissions
About this article
Cite this article
Wu, R., Haas, W., Dephoure, N. et al. A large-scale method to measure absolute protein phosphorylation stoichiometries. Nat Methods 8, 677–683 (2011). https://doi.org/10.1038/nmeth.1636
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1636
This article is cited by
-
A streamlined tandem tip-based workflow for sensitive nanoscale phosphoproteomics
Communications Biology (2023)
-
A time-resolved multi-omics atlas of Acanthamoeba castellanii encystment
Nature Communications (2022)
-
Global profiling of distinct cysteine redox forms reveals wide-ranging redox regulation in C. elegans
Nature Communications (2021)
-
Wittig reagents for chemoselective sulfenic acid ligation enables global site stoichiometry analysis and redox-controlled mitochondrial targeting
Nature Chemistry (2021)
-
Proteomic analysis of degradation ubiquitin signaling by ubiquitin occupancy changes responding to 26S proteasome inhibition
Clinical Proteomics (2020)