Abstract
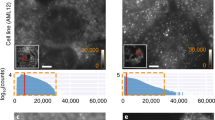
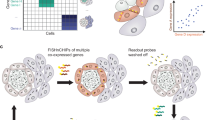
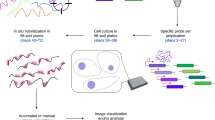
Fluorescence in situ hybridization (FISH) is widely used to obtain information about transcript copy number and subcellular localization in single cells. However, current approaches do not readily scale to the analysis of whole transcriptomes. Here we show that branched DNA technology combined with automated liquid handling, high-content imaging and quantitative image analysis allows highly reproducible quantification of transcript abundance in thousands of single cells at single-molecule resolution. In addition, it allows extraction of a multivariate feature set quantifying subcellular patterning and spatial properties of transcripts and their cell-to-cell variability. This has multiple implications for the functional interpretation of cell-to-cell variability in gene expression and enables the unbiased identification of functionally relevant in situ signatures of the transcriptome without the need for perturbations. Because this method can be incorporated in a wide variety of high-throughput image-based approaches, we expect it to be broadly applicable.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Brown, P.O. & Botstein, D. Exploring the new world of the genome with DNA microarrays. Nat. Genet. 21, 33–37 (1999).
Nagalakshmi, U. et al. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 320, 1344–1349 (2008).
Wilhelm, B.T. et al. Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453, 1239–1243 (2008).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Tang, F. et al. mRNA-seq whole-transcriptome analysis of a single cell. Nat. Methods 6, 377–382 (2009).
Hashimshony, T., Wagner, F., Sher, N. & Yanai, I. CEL-Seq: single-cell RNA-Seq by multiplexed linear amplification. Cell Rep. 2, 666–673 (2012).
Ramsköld, D. et al. Full-length mRNA-seq from single-cell levels of RNA and individual circulating tumor cells. Nat. Biotechnol. 30, 777–782 (2012).
Shalek, A.K. et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498, 236–240 (2013).
Goetz, J.J. & Trimarchi, J.M. Transcriptome sequencing of single cells with Smart-Seq. Nat. Biotechnol. 30, 763–765 (2012).
Lau, J.Y. et al. Significance of serum hepatitis C virus RNA levels in chronic hepatitis C. Lancet 341, 1501–1504 (1993).
Kern, D. et al. An enhanced-sensitivity branched-DNA assay for quantification of human immunodeficiency virus type 1 RNA in plasma. J. Clin. Microbiol. 34, 3196–3202 (1996).
Player, A.N., Shen, L.P., Kenny, D., Antao, V.P. & Kolberg, J.A. Single-copy gene detection using branched DNA (bDNA) in situ hybridization. J. Histochem. Cytochem. 49, 603–612 (2001).
Kenny, D., Shen, L. & Kolberg, J.A. Detection of viral infection and gene expression in clinical tissue specimens using branched DNA (bDNA) in situ hybridization. J. Histochem. Cytochem. 50, 1219–1227 (2002).
Ma, X.-J., Wu, X. & Luo, Y. Biomarkers for differentiating melanoma from benign nevus in the skin. US patent application 20120071343 (2012).
Femino, A.M., Fay, F.S., Fogarty, K. & Singer, R.H. Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998).
Raj, A., van den Bogaard, P., Rifkin, S.A., van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 5, 877–879 (2008).
Chartrand, P., Bertrand, E., Singer, R.H. & Long, R.M. Sensitive and high-resolution detection of RNA in situ. Methods Enzymol. 318, 493–506 (2000).
Bhatt, D.M. et al. Transcript dynamics of proinflammatory genes revealed by sequence analysis of subcellular RNA fractions. Cell 150, 279–290 (2012).
Carpenter, A.E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Raj, A. & Tyagi, S. Detection of individual endogenous RNA transcripts in situ using multiple singly labeled probes. Methods Enzymol. 472, 365–386 (2010).
So, L.H. et al. General properties of transcriptional time series in Escherichia coli. Nat. Genet. 43, 554–560 (2011).
Rämö, P., Sacher, R., Snijder, B., Begemann, B. & Pelkmans, L. CellClassifier: supervised learning of cellular phenotypes. Bioinformatics 25, 3028–3030 (2009).
Snijder, B. et al. Single-cell analysis of population context advances RNAi screening at multiple levels. Mol. Syst. Biol. 8, 579 (2012).
Hebenstreit, D. et al. RNA sequencing reveals two major classes of gene expression levels in metazoan cells. Mol. Syst. Biol. 7, 497 (2011).
Raj, A., Peskin, C.S., Tranchina, D., Vargas, D.Y. & Tyagi, S. Stochastic mRNA synthesis in mammalian cells. PLoS Biol. 4, e309 (2006).
Trcek, T., Larson, D., Moldón, A., Query, C. & Singer, R. Single-molecule mRNA decay measurements reveal promoter- regulated mRNA stability in yeast. Cell 147, 1484–1497 (2011).
Buganim, Y. et al. Single-cell expression analyses during cellular reprogramming reveal an early stochastic and a late hierarchic phase. Cell 150, 1209–1222 (2012).
Szklarczyk, D. et al. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res. 39, D561–D568 (2011).
Lubeck, E. & Cai, L. Single-cell systems biology by super-resolution imaging and combinatorial labeling. Nat. Methods 9, 743–748 (2012).
Nguyen, Q.N., Lipshutz, R.J. & Ma, Y. Methods of labeling cells, labeled cells, and uses thereof. US patent application 20120178081 (2012).
Levesque, M.J. & Raj, A. Single-chromosome transcriptional profiling reveals chromosomal gene expression regulation. Nat. Methods 10, 246–248 (2013).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Martinez, W.L. & Martinez, A.R. Computational Statistics Handbook with MATLAB 2nd edn. (CRC Press, 2008).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
We would like to acknowledge B. Snijder and Y. Yakimovich for help with computational analysis and infrastructure, J. Patterson for assistance, Q. Nguyen and S. Lai from Affymetrix for helpful comments on experimental procedures, J. Ellenberg (European Molecular Biology Laboratory) for reagents, and all members of the lab for useful comments on the manuscript. L.P. acknowledges financial support for this project from SystemsX.ch, the European Union, University of Zurich and University of Zurich Research Priority Program in Systems Biology and Functional Genomics.
Author information
Authors and Affiliations
Contributions
L.P. initiated the study. N.B., T.S. and L.P. designed and analyzed the experiments and wrote the manuscript. N.B. and T.S. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Tables 1 and 2, Supplementary Protocol and Supplementary Notes 1–6 (PDF 26058 kb)
Supplementary Table 3
High-throughput bDNA sm-FISH comparison to RNA-seq (XLSX 198 kb)
Supplementary Table 4
Features used for gene clustering (XLSX 7511 kb)
Supplementary Table 5
Equipment and settings (XLSX 11 kb)
Supplementary Table 6
Full RNA-seq dataset (XLSX 881 kb)
Supplementary Software
Spots detection and pattern recognition (ZIP 7358 kb)
Rights and permissions
About this article
Cite this article
Battich, N., Stoeger, T. & Pelkmans, L. Image-based transcriptomics in thousands of single human cells at single-molecule resolution. Nat Methods 10, 1127–1133 (2013). https://doi.org/10.1038/nmeth.2657
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2657
This article is cited by
-
Spatiotemporal control of RNA metabolism and CRISPR–Cas functions using engineered photoswitchable RNA-binding proteins
Nature Protocols (2024)
-
Cellular state landscape and herpes simplex virus type 1 infection progression are connected
Nature Communications (2023)
-
Cohesin couples transcriptional bursting probabilities of inducible enhancers and promoters
Nature Communications (2022)
-
Optogenetic control of RNA function and metabolism using engineered light-switchable RNA-binding proteins
Nature Biotechnology (2022)
-
Visualization and modeling of inhibition of IL-1β and TNF-α mRNA transcription at the single-cell level
Scientific Reports (2021)