Abstract
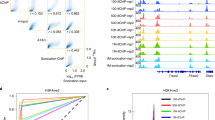
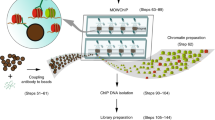
The sensitivity of chromatin immunoprecipitation (ChIP) assays poses a major obstacle for epigenomic studies of low-abundance cells. Here we present a microfluidics-based ChIP-seq protocol using as few as 100 cells via drastically improved collection of high-quality ChIP-enriched DNA. Using this technology, we uncovered many new enhancers and super enhancers in hematopoietic stem and progenitor cells from mouse fetal liver, suggesting that enhancer activity is highly dynamic during early hematopoiesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Park, P.J. Nat. Rev. Genet. 10, 669–680 (2009).
Adli, M., Zhu, J. & Bernstein, B.E. Nat. Methods 7, 615–618 (2010).
Shankaranarayanan, P. et al. Nat. Methods 8, 565–567 (2011).
Zwart, W. et al. BMC Genomics 14, 232 (2013).
Lara-Astiaso, D. et al. Science 345, 943–949 (2014).
Wu, A.R. et al. Lab Chip 9, 1365–1370 (2009).
Geng, T. et al. Lab Chip 11, 2842–2848 (2011).
Wu, A.R. et al. Lab Chip 12, 2190–2198 (2012).
Hong, J.W., Studer, V., Hang, G., Anderson, W.F. & Quake, S.R. Nat. Biotechnol. 22, 435–439 (2004).
Kundaje, A. et al. Nature 518, 317–330 (2015).
Landt, S.G. et al. Genome Res. 22, 1813–1831 (2012).
Göttgens, B. et al. EMBO J. 21, 3039–3050 (2002).
Thoms, J.A. et al. Blood 117, 7079–7089 (2011).
Nottingham, W.T. et al. Blood 110, 4188–4197 (2007).
Hnisz, D. et al. Cell 155, 934–947 (2013).
Buza-Vidas, N. et al. Blood 113, 3453–3460 (2009).
Jakobsen, J.S. et al. BMC Genomics 16, 46 (2015).
McKinney-Freeman, S. et al. Cell Stem Cell 11, 701–714 (2012).
McKinney-Freeman, S.L. et al. Blood 114, 268–278 (2009).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Genome Biol. 9, R137 (2008).
Kharchenko, P.V., Tolstorukov, M.Y. & Park, P.J. Nat. Biotechnol. 26, 1351–1359 (2008).
Marson, A. et al. Cell 134, 521–533 (2008).
Goren, A. et al. Nat. Methods 7, 47–49 (2010).
Wei, G. et al. Immunity 30, 155–167 (2009).
Heinz, S. et al. Mol. Cell 38, 576–589 (2010).
Firpi, H.A., Ucar, D. & Tan, K. Bioinformatics 26, 1579–1586 (2010).
Portales-Casamar, E. et al. Nucleic Acids Res. 38, D105–D110 (2010).
Newburger, D.E. & Bulyk, M.L. Nucleic Acids Res. 37, D77–D82 (2009).
Matys, V. et al. Nucleic Acids Res. 34, D108–D110 (2006).
Wilson, N.K. et al. Cell Stem Cell 7, 532–544 (2010).
Whitington, T., Frith, M.C., Johnson, J. & Bailey, T.L. Nucleic Acids Res. 39, e98 (2011).
Huang, D.W., Sherman, B.T. & Lempicki, R.A. Nat. Protoc. 4, 44–57 (2009).
Benjamini, Y. & Hochberg, Y. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Acknowledgements
We thank the Genomics Research Laboratory of Virginia Bioinformatics Institute and Genomics Division of Iowa Institute of Human Genetics for providing sequencing service. We thank L. Van Tol and University of Iowa Institute for Clinical and Translational Science for providing computing support. This work was supported by US National Institutes of Health grants EB017855 (K.T. and C.L.), CA174577 (C.L.), EB017235 (C.L.), HG006130 (K.T.) and GM104369 (K.T.) and a seed grant from Virginia Tech Institute for Critical Technology and Applied Science (C.L.).
Author information
Authors and Affiliations
Contributions
C.L. designed the microfluidic device. C.L., K.T. and Z.C. developed the MOWChIP-seq technology. Z.C. generated the MOWChIP-qPCR and MOWChIP-seq data. K.T., C.C. and B.H. designed the biological experiments, isolated the primary cells from mice and conducted data analysis. All authors wrote the manuscript together.
Corresponding authors
Ethics declarations
Competing interests
Virginia Tech (on behalf of C.L. and Z.C.) filed a US utility patent (Application no. 14/511,422; Publication no. US 2015/0105287 A1) on the MOWChIP-seq technology on 10 October 2014.
Integrated supplementary information
Supplementary Figure 1 Setup of the microfluidic device with ancillary control and liquid-delivery equipment.
Each microfluidic device has one inlet, one outlet, and one port for applying pressure to the control channel to actuate the on-chip pneumatic microvalve via its connection with a solenoid valve. (a) The inlet is connected to a syringe pump when a reagent is delivered (while the outlet is open to the atmosphere). (b) During oscillatory washing, the inlet and outlet are attached to two separate solenoid valves that are alternatingly actuated. (c) The microfluidic chip on a microscope stage. There are 6 parallel microfluidic devices on the same chip.
Supplementary Figure 2 Microscopic images that recorded the MOWChIP process.
The scale bar represents 1 mm. (a) The microfluidic chamber is filled with buffer. (b) The pressurization of the control channel partially closes the pneumatic microvalve (i.e. liquid passes the microvalve while beads get stopped). (c) The IP beads are stopped by the partially closed microvalve and form a packed bed. (d) Oscillatory washing by applying alternating pressure pulses on the inlet and the outlet after the flow of the chromatin fragments through the packed bed. (e) The retention of IP beads using a magnet (not shown) while flushing out undesired molecules and debris.
Supplementary Figure 3 The yield of ChIP DNA as a function of antibody concentration used during coating of the IP beads.
ChIP was done using anti-H3K4me3 antibody and GM12878 cells. The chromatin samples were processed by IPure kit (C03010012, Diagenode) to extract DNA before the measurement. The concentrations of DNA in these samples were measured by a Qubit 2.0 fluorometer using dsDNA HS Assay kit (Q32851, Life Technologies). Data points are mean of two replicate experiments. Error bars represent s.d.. Under the optimized MOWChIP-Seq conditions (i.e. antibody concentration of 5 µg/ml for 1000-cell samples and 6.6 µg/ml for 10000-cell samples), 1000 and 10000 cells yielded 180 pg and 1.3 ng ChIP DNA, respectively. In comparison, the same numbers of cells yielded 2.9 ng and 24.6 ng DNA without the ChIP step.
Supplementary Figure 4 Genome-wide correlations among MOWChIP-Seq data generated using various GM12878 cell sample sizes.
Normalized ChIP-Seq signals from promoter regions across the genome were used for computing correlations (n = 34342). Promoter regions were defined as +/- 2kb around transcription start sites (TSS). TSS annotation was based on RefSeq. Promoter regions with zero values in both samples were removed before computing correlation coefficients. Colors represent Pearson correlation coefficients. (a) H3K4me3. (b) H3K27Ac.
Supplementary Figure 5 Purification of hematopoietic stem and progenitor cells (HSPCs) from mouse fetal liver.
HSPCs were purified as Lineage- Sca-1+ Kit+ (LSK) cells using fluorescence activated cell sorting. Cells were double sorted with a yield sort first followed by a purity sort. Percentages shown in each panel indicate the purity of fetal liver LSK cells.
Supplementary Figure 6 Genome-wide correlations between MOWChIP-seq data on FL HSPC data and published data.
Normalized ChIP-Seq signals from promoter regions across the genome were used for computing correlations (n = 34342). Promoter regions were defined as +/- 2kb around transcription start sites (TSS). TSS annotation was based on RefSeq. Promoter regions with zero values in both samples were removed before computing correlation coefficients. Colors represent Pearson correlation coefficients. Bone marrow (BM) LSK data was obtained from Adli et al. BM LT-HSC, B cell and macrophage data were obtained from Lara-Astiaso et al. (a) H3K4me3. (b) H3K27Ac.
Supplementary Figure 7 Scatter plots of histone modification signals and transcript expression levels.
FL LSK microarray data was downloaded from the GEO database (accession no.: GSE21404). All transcripts annotated in the RefSeq database were used for computing Spearman correlation coefficient (n = 24658). X-axis, normalized expression values of transcripts. Y-axis, mean value of the normalized MOWChIP-Seq signals in the +/- 2Kb region of a transcription start site. r denotes Spearman correlation coefficient. a,c,e,g: H3K4me3; b,d,f,h: H3K27Ac. a,b: data generated using 100 cells. c,d: data generated using 600 cells. e,f: data generated using 1000 cells. g,h: data generated using 10000 cells.
Supplementary Figure 9 Normalized H3K27ac MOWChIP-seq signals at known enhancers in FL HSPCs.
Data generated using different numbers of cells are shown. (a) Erg +85 enhancer. (b) Runx1 +24 enhancer. BM_HSC data was obtained from Lara-Astiaso et al.
Supplementary Figure 11 DNA fragment size distribution after sonication and library preparation with starting cell numbers of 600 and 100.
DNA size distribution was examined by an Agilent 2200 TapeStation using high sensitivity DNA analysis kit (5067-4626, Agilent). (a) DNA size distribution using 600 cells as the starting material and before library preparation. DNA was purified after pre-ChIP processing and MOWChIP. (b) DNA size distribution using 600 cells as the starting material and after library preparation. DNA was purified after pre-ChIP processing, MOWChIP, and library preparation. (c) DNA size distribution using 100 cells as the starting material and after library preparation. DNA was purified after pre-ChIP processing, MOWChIP, and library preparation.
Supplementary Figure 12 Genome-wide correlations between MOWChIP-seq data generated using two protocols for chromatin preparation.
Chromatin was prepared from GM12878 cells. “From stock chromatin”, cross-linked and sonicated chromatin was prepared from 10000 cells. Chromatin amounts equivalent to 100 and 600 cells were then aliquoted from the stock chromatin preparation. “From cells directly”, chromatin was prepared directly from 100 and 600 starting cells. Values shown are Pearson correlation coefficients. Normalized ChIP-Seq signals from promoter regions across the genome were used for computing correlations (n = 50961). Promoter regions were defined as +/- 2kb around transcription start sites (TSS). TSS annotation was based on RefSeq. Promoter regions with zero values in both samples were removed before computing correlation coefficients. Duplicate experiments were done using each protocol. (a) H3K4me3 data using 600 cells. (b) H3K4me3 data using 100 cells; (c) H3K27Ac data using 600 cells. (d) H3K27Ac data using 100 cells.
Supplementary Figure 13 MOWChIP-seq data using 100 and 600 starting cells.
Chromatin was prepared from GM12878 cells. (a) and (b), Receiver Operating Characteristic (ROC) curves. ROC curves were constructed by comparing MOWChIP-Seq data to published gold-standard data generated using conventional protocols with millions of cells. Values shown are average Area Under the ROC curve (AUC) of two replicate experiments. (a) H3K4me3 data. (b) H3K27Ac data. (c) and (d), Normalized MOWChIP-Seq signals at the SPI1 gene locus (c) and at the immunoglobin heavy chain locus (d). (c) H3K4me3 data. (d) H3K27Ac data.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Tables 1–3 (PDF 2908 kb)
Supplementary Table 4
Enriched transcription factor motifs in FL HSPC enhancers (XLSX 37 kb)
Supplementary Table 5
FL HSPC specific enhancers identified in this study (XLSX 96 kb)
Supplementary Table 6
FL HSPC super enhancers identified in this study (XLSX 13 kb)
Supplementary Table 7
Enriched Gene Ontology terms among target genes of super enhancers (XLSX 42 kb)
Microfluidic oscillatory washing to remove nonspecific adsorption and trapping after ChIP
Pressure pulses (each at 3 psi, with a pulse width of 0.5 s and an interval of 0.5 s between two pulses) were applied alternatingly at either end of the microfluidic chamber for washing of the IP beads. (AVI 14031 kb)
Rights and permissions
About this article
Cite this article
Cao, Z., Chen, C., He, B. et al. A microfluidic device for epigenomic profiling using 100 cells. Nat Methods 12, 959–962 (2015). https://doi.org/10.1038/nmeth.3488
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3488
This article is cited by
-
Epigenomic and transcriptomic analyses reveal differences between low-grade inflammation and severe exhaustion in LPS-challenged murine monocytes
Communications Biology (2022)
-
Transcription factor chromatin profiling genome-wide using uliCUT&RUN in single cells and individual blastocysts
Nature Protocols (2021)
-
Chromatin integration labeling for mapping DNA-binding proteins and modifications with low input
Nature Protocols (2020)
-
Microfluidic techniques for enhancing biofuel and biorefinery industry based on microalgae
Biotechnology for Biofuels (2019)
-
Profiling chromatin states using single-cell itChIP-seq
Nature Cell Biology (2019)