Abstract
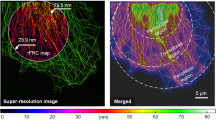
Super-resolution fluorescence microscopy has become a widely used tool in many areas of research. However, designing and validating super-resolution experiments to address a research question in a technically feasible and scientifically rigorous manner remains a fundamental challenge. We developed SuReSim, a software tool that simulates localization data of arbitrary three-dimensional structures represented by ground truth models, allowing users to systematically explore how changing experimental parameters can affect potential imaging outcomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Dani, A., Huang, B., Bergan, J., Dulac, C. & Zhuang, X. Neuron 68, 843–856 (2010).
Xu, K., Zhong, G. & Zhuang, X. Science 339, 452–456 (2013).
Xu, K., Babcock, H.P. & Zhuang, X. Nat. Methods 9, 185–188 (2012).
Endesfelder, U. & Heilemann, M. Nat. Methods 11, 235–238 (2014).
Nieuwenhuizen, R.P. et al. Nat. Methods 10, 557–562 (2013).
Banterle, N., Bui, K.H., Lemke, E.A. & Beck, M. J. Struct. Biol. 183, 363–367 (2013).
Deschout, H. et al. Nat. Methods 11, 253–266 (2014).
Endesfelder, U., Malkusch, S., Fricke, F. & Heilemann, M. Histochem. Cell Biol. 141, 629–638 (2014).
Takamori, S. et al. Cell 127, 831–846 (2006).
Horstmann, H., Korber, C., Satzler, K., Aydin, D. & Kuner, T. PLoS ONE 7, e35172 (2012).
Weber, K., Rathke, P.C. & Osborn, M. Proc. Natl. Acad. Sci. USA 75, 1820–1824 (1978).
Kremer, J.R., Mastronarde, D.N. & McIntosh, J.R. J. Struct. Biol. 116, 71–76 (1996).
Dempsey, G.T., Vaughan, J.C., Chen, K.H., Bates, M. & Zhuang, X. Nat. Methods 8, 1027–1036 (2011).
Buchwalow, I., Samoilova, V., Boecker, W. & Tiemann, M. Sci. Rep. 1, 28 (2011).
Deschout, H. et al. Nat. Methods 11, 253–266 (2014).
Nieuwenhuizen, R.P. et al. Nat. Methods 10, 557–562 (2013).
Sage, D. et al. Nat. Methods 12, 717–724 (2015).
Huang, B., Jones, S.A., Brandenburg, B. & Zhuang, X. Nat. Methods 5, 1047–1052 (2008).
Finan, K., Raulf, A. & Heilemann, M. Angew. Chem. Int. Ed. Engl. 54, 12049–12052 (2015).
Früh, S.M., Schoen, I., Ries, J. & Vogel, V. Nat. Commun. 6, 7275 (2015).
Heilemann, M. et al. Angew. Chem. Int. Edn. Engl. 47, 6172–6176 (2008).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer Control of Microscopes Using μManager (John Wiley & Sons, 2010).
Wolter, S. et al. Nat. Methods 9, 1040–1041 (2012).
Ovesný, M., Krizek, P., Borkovec, J., Svindrych, Z. & Hagen, G.M. Bioinformatics 30, 2389–2390 (2014).
Acknowledgements
We thank I. Schön and S. Früh (Laboratory of Applied Mechanobiology, Department of Health Sciences and Technology, ETH Zurich, Zurich, Switzerland) for providing the experimental SMLM data on fibronectin fibers, M. Cyrklaff (Centre for Infectious Diseases, Parasitology, Heidelberg, Germany) for the F-actin model data on erythrocytes, S. Srismith and M. Lanzer (Centre for Infectious Diseases, Parasitology, Heidelberg, Germany) for providing materials for erythrocyte stainings, and D. Mastronarde (Laboratory for Three-dimensional Fine Structure, Department of Molecular, Cellular, and Developmental Biology, University of Colorado, Boulder, Colorado, USA) and J. McIntosh for providing various electron-tomographic models of organelles. We thank B. Rieger and R. Nieuwenhuizen for discussions, M. Scheurer for help with rewriting the software in Java, and C. Kocksch and M. Kaiser for excellent technical assistance. This work was supported by the German Science Foundation through the CellNetworks Cluster of Excellence (EXC 81 to T.K.) and the Cluster of Excellence Macromolecular Complexes (EXC 115 to M.H.).
Author information
Authors and Affiliations
Contributions
V.V., F.H. and T.K. conceived the project and designed the software. F.H. and V.V. programmed the software and designed, performed and analyzed experiments. M.H. and T.K. supervised the project and designed experiments. F.H., V.V., M.H. and T.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–18, Supplementary Tables 1–8 and Supplementary Notes 1–3 (PDF 9039 kb)
Supplementary Software
SuReSim Software (ZIP 128199 kb)
Rights and permissions
About this article
Cite this article
Venkataramani, V., Herrmannsdörfer, F., Heilemann, M. et al. SuReSim: simulating localization microscopy experiments from ground truth models. Nat Methods 13, 319–321 (2016). https://doi.org/10.1038/nmeth.3775
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3775
This article is cited by
-
Machine learning framework to segment sarcomeric structures in SMLM data
Scientific Reports (2023)
-
Single molecule imaging simulations with advanced fluorophore photophysics
Communications Biology (2023)
-
Maximum-likelihood model fitting for quantitative analysis of SMLM data
Nature Methods (2023)
-
FluoSim: simulator of single molecule dynamics for fluorescence live-cell and super-resolution imaging of membrane proteins
Scientific Reports (2020)
-
Super-resolution microscopy as a powerful tool to study complex synthetic materials
Nature Reviews Chemistry (2019)