Abstract
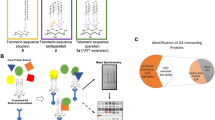
Molecular simulations suggest that the stability of a folded macromolecule increases in a confined space due to entropic effects. However, due to the interactions between the confined molecular structure and the walls of the container, clear-cut experimental evidence for this prediction is lacking. Here, using DNA origami nanocages, we show the pure effect of confined space on the property of individual human telomeric DNA G-quadruplexes. We induce targeted mechanical unfolding of the G-quadruplex while leaving the nanocage unperturbed. We find that the mechanical and thermodynamic stabilities of the G-quadruplex inside the nanocage increase with decreasing cage size. Compared to the case of diluted or molecularly crowded buffer solutions, the G-quadruplex inside the nanocage is significantly more stable, showing a 100 times faster folding rate. Our findings suggest the possibility of co-replicational or co-transcriptional folding of G-quadruplex inside the polymerase machinery in cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Tang, Y.-C. et al. Structural features of the GroEL-GroES nano-cage required for rapid folding of encapsulated protein. Cell 125, 903–914 (2006).
Brinker, A. et al. Dual function of protein confinement in chaperonin-assisted protein folding. Cell 107, 223–233 (2001).
Baumketner, A., Jewett, A. & Shea, J. E. Effects of confinement in chaperonin assisted protein folding: rate enhancement by decreasing the roughness of the folding energy landscape. J. Mol. Biol. 332, 701–713 (2003).
Jewett, A. I., Baumketner, A. & Shea, J.-E. Accelerated folding in the weak hydrophobic environment of a chaperonin cavity: creation of an alternate fast folding pathway. Proc. Natl Acad. Sci. USA 101, 13192–13197 (2004).
Takagi, F., Koga, N. & Takada, S. How protein thermodynamics and folding mechanisms are altered by the chaperonin cage: molecular simulations. Proc. Natl Acad. Sci. USA 100, 11367–11372 (2003).
Zhou, H.-X. Protein folding and binding in confined spaces and in crowded solutions. J. Mol. Recognit. 17, 368–375 (2004).
Zhou, H.-X. & Dill, K. A. Stabilization of proteins in confined spaces†. Biochemistry 40, 11289–11293 (2001).
Eggers, D. K. & Valentine, J. S. Molecular confinement influences protein structure and enhances thermal protein stability. Protein Sci. 10, 250–261 (2001).
Senske, M., Smith, A. E. & Pielak, G. J. Protein stability in reverse micelles. Angew. Chem. Int. Ed. 55, 3586–3589 (2016).
Zhou, H. X., Rivas, G. & Minton, A. P. Macromolecular crowding and confinement: biochemical, biophysical, and potential physiological consequences. Annu. Rev. Biophys. 37, 375–397 (2008).
Mitchell, M., Gillis, A., Futahashi, M., Fujiwara, H. & Skordalakes, E. Structural basis for telomerase catalytic subunit TERT binding to RNA template and telomeric DNA. Nat. Struct. Mol. Biol. 17, 513–518 (2010).
Jansson, L. I. et al. Structural basis of template-boundary definition in Tetrahymena telomerase. Nat. Struct. Mol. Biol. 22, 883–888 (2015).
Koirala, D. et al. Intramolecular folding in three tandem guanine repeats of human telomeric DNA. Chem. Commun. 48, 2006–2008 (2012).
Li, W., Hou, X.-M., Wang, P.-Y., Xi, X.-G. & Ming Li, M. Direct measurement of sequential folding pathway and energy landscape of human telomeric G-quadruplex structures. J. Am. Chem. Soc. 135, 6423–6426 (2013).
Han, H. & Hurley, L. H. G-quadruplex DNA: a potential target for anti-cancer drug design. Trends Pharmacol. Sci. 21, 136–142 (2000).
Balasubramanian, S. & Neidle, S. G-quadruplex nucleic acids as therapeutic targets. Curr. Opin. Chem. Biol. 13, 345–353 (2009).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Moffitt, J. R., Chemla, Y. R., Smith, S. B. & Bustamante, C. Recent advances in optical tweezers. Annu. Rev. Biochem. 77, 205–228 (2008).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Koirala, D. et al. A single-molecule platform for investigation of interactions between G-quadruplexes and small-molecule ligands. Nat. Chem. 3, 782–787 (2011).
Venczel, E. A. & Sen, D. Parallel and antiparallel G-DNA structures from a complex telomeric sequence. Biochemistry 32, 6220–6228 (1993).
Yu, Z. et al. Click chemistry assisted single-molecule fingerprinting reveals a 3D biomolecular folding funnel. J. Am. Chem. Soc. 134, 12338–12341 (2012).
An, N., Fleming, A. M., Middleton, E. G. & Burrows, C. J. Single-molecule investigation of G-quadruplex folds of the human telomere sequence in a protein nanocavity. Proc. Natl Acad. Sci. USA 111, 14325–14331 (2014).
Zhou, J. et al. Formation and stabilization of G-quadruplex in nanosized water pools. Chem. Commun. 46, 1700–1702 (2010).
Gore, J., Ritort, F. & Bustamante, C. Bias and error in estimates of equilibrium free-energy differences from nonequilibrium measurements. Proc. Natl Acad. Sci. USA 100, 12564–12569 (2003).
Bennett, C. H. Efficient estimation of free energy differences from Monte Carlo data. J. Comput. Phys. 22, 245–268 (1976).
Crooks, G. E. Path-ensemble averages in systems driven far from equilibrium. Phys. Rev. E 61, 2361–2366 (2000).
Collin, D. et al. Verification of the Crooks fluctuation theorem and recovery of RNA folding free energies. Nature 437, 231–234 (2005).
Dhakal, S. et al. Structural and mechanical properties of individual human telomeric G-quadruplexes in molecularly crowded solutions. Nucleic Acids Res. 41, 3915–3923 (2013).
Dudko, O. K., Hummer, G. & Szabo, A. Theory, analysis, and interpretation of single-molecule force spectroscopy experiments. Proc. Natl Acad. Sci. USA 105, 15755–15760 (2008).
Greenleaf, W. J., Frieda, K. L., Foster, D. A., Woodside, M. T. & Block, S. M. Direct observation of hierarchical folding in single riboswitch aptamers. Science 319, 630–633 (2008).
Xie, M., Podlevsky, J. D., Qi, X., Bley, C. J. & Chen, J. J.-L. A novel motif in telomerase reverse transcriptase regulates telomere repeat addition rate and processivity. Nucleic Acids Res. 38, 1982–1996 (2010).
Paeschke, K., Simonsson, T., Postberg, J., Rhodes, D. & Lipps, H. J. Telomere end-binding proteins control the formation of G-quadruplex DNA structures in vivo. Nat. Struct. Mol. Biol. 12, 847–854 (2005).
Miyoshi, D., Karimata, H. & Sugimoto, N. Hydration regulates thermodynamics of G-quadruplex formation under molecular crowding conditions. J. Am. Chem. Soc. 128, 7957–7963 (2006).
Dai, J. et al. Structure of the intramolecular human telomeric G-quadruplex in potassium solution: a novel adenine triple formation. Nucleic Acids Res. 35, 2440–2450 (2007).
Pelliccioli, A. P. & Wirz, J. Photoremovable protecting groups: reaction mechanisms and applications. Photochem. Photobiol. Sci. 1, 441–458 (2002).
Luchette, P., Abiy, N. & Mao, H. Microanalysis of clouding process at the single droplet level. Sens. Actuat. B 128, 154–160 (2007).
Mao, H. & Luchette, P. An integrated laser-tweezers instrument for microanalysis of individual protein aggregates. Sens. Actuat. B 129, 764–771 (2008).
Acknowledgements
This project was supported by the Japan Society for the Promotion of Science (JSPS) and the National Science Foundation (NSF) under the JSPS-NSF International Collaborations in Chemistry (ICC) (CHE-1415883, to H.S. and H.M.). H.M. acknowledges support from NSF (CHE-1609504). M.E. acknowledges supports from JSPS KAKENHI (grant nos. 24104002, 15H03837 and 16K14033).
Author information
Authors and Affiliations
Contributions
H.M. and H.S. conceived the project. H.M., H.S., M.E. and P.S. designed experiments. H.M. supervised mechanical unfolding experiments. H.S. and M.E. supervised origami synthesis and characterization. P.S. performed mechanical unfolding experiments and prepared origami constructs. P.S. and H.M. carried out data analyses. S.J. performed mechanical unfolding experiments. T.E. and K.H. prepared and characterized origami constructs. H.M., P.S., M.E. and H.S. co-wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 2231 kb)
Rights and permissions
About this article
Cite this article
Shrestha, P., Jonchhe, S., Emura, T. et al. Confined space facilitates G-quadruplex formation. Nature Nanotech 12, 582–588 (2017). https://doi.org/10.1038/nnano.2017.29
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2017.29
This article is cited by
-
Fabricating higher-order functional DNA origami structures to reveal biological processes at multiple scales
NPG Asia Materials (2023)
-
DNA origami
Nature Reviews Methods Primers (2021)
-
The regulation and functions of DNA and RNA G-quadruplexes
Nature Reviews Molecular Cell Biology (2020)
-
Overview of the “biophysics in nano-space” session at the 57th annual meeting of the biophysical society of Japan
Biophysical Reviews (2020)
-
Sites of high local frustration in DNA origami
Nature Communications (2019)