Abstract
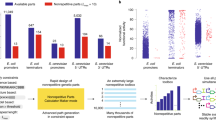
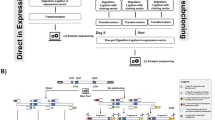
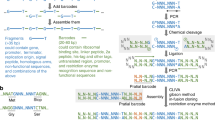
Randomized gene libraries may be constructed and screened to find novel candidates with particular functions, and the applications can range widely, from protein engineering to selecting new microRNAs. Here we describe a technique to construct gene libraries using semi-randomized weighted oligonucleotide synthesis and end-to-end ligation. This method makes it possible to search the combinatorial space around a particular nucleotide sequence for a greater number of positions than is possible with fully randomized oligonucleotides. As an alternative to full cassette construction, library mutations can also be introduced through 'round-the-world PCR' approaches. Construction of a randomized gene cassette and cloning can typically be achieved in 2 weeks. Therefore, these are rapid and convenient methods to generate successive generations of libraries for iterative selection and optimization.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Smith, G.P. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science 228, 1315–1317 (1985).
Winter, G., Griffiths, A.D., Hawkins, R.E. & Hoogenboom, H.R. Making antibodies by phage display technology. Annu. Rev. Immunol. 12, 433–455 (1994).
Hanes, J. & Pluckthun, A. In vitro selection and evolution of functional proteins by using ribosome display. Proc. Natl. Acad. Sci. USA 94, 4937–4942 (1997).
Lipovsek, D. & Pluckthun, A. In-vitro protein evolution by ribosome display and mRNA display. J. Immunol. Methods 290, 51–67 (2004).
Tawfik, D.S. & Griffiths, A.D. Man-made cell-like compartments for molecular evolution. Nat. Biotechnol. 16, 652–656 (1998).
Campbell, T.B. & Cech, T.R. Identification of ribozymes within a ribozyme library that efficiently cleave a long substrate RNA. RNA 1, 598–609 (1995).
Zhao, H.F. et al. High-throughput screening of effective siRNAs from RNAi libraries delivered via bacterial invasion. Nat. Methods 2, 967–973 (2005).
Isalan, M., Santori, M.I., Gonzalez, C. & Serrano, L. Localized transfection on arrays of magnetic beads coated with PCR products. Nat. Methods 2, 113–118 (2005).
Isalan, M., Klug, A. & Choo, Y. A rapid, generally applicable method to engineer zinc fingers illustrated by targeting the HIV-1 promoter. Nat. Biotechnol. 19, 656–660 (2001).
Griffiths, A.D. & Tawfik, D.S. Directed evolution of an extremely fast phosphotriesterase by in vitro compartmentalization. EMBO J. 22, 24–35 (2003).
Scott, J.K. & Smith, G.P. Searching for peptide ligands with an epitope library. Science 249, 386–390 (1990).
Tong, A.H. et al. A combined experimental and computational strategy to define protein interaction networks for peptide recognition modules. Science 295, 321–324 (2002).
Reynolds, L. et al. Repression of the HIV-1 5′ LTR promoter and inhibition of HIV-1 replication by using engineered zinc-finger transcription factors. Proc. Natl. Acad. Sci. USA 100, 1615–1620 (2003).
Santori, M.I., Serrano, L. & Isalan, M. Transfection with magnetic beads coated with PCR products and other nucleic acids. Nat. Protocols 1, (2006) (doi:10.1038/nprot2006.74).
del Rio, G., Osuna, J. & Soberon, X. Combinatorial libraries of proteins: analysis of efficiency of mutagenesis techniques. Biotechniques 17, 1132–1139 (1994).
Hermes, J.D., Parekh, S.M., Blacklow, S.C., Koster, H. & Knowles, J.R. A reliable method for random mutagenesis: the generation of mutant libraries using spiked oligodeoxyribonucleotide primers. Gene 84, 143–151 (1989).
Isalan, M., Klug, A. & Choo, Y. Comprehensive DNA recognition through concerted interactions from adjacent zinc fingers. Biochemistry 37, 12026–12033 (1998).
Schymkowitz, J. et al. The FoldX web server: an online force field. Nucleic Acids Res. 33, W382–W388 (2005).
Innis, M.A. PCR Protocols: A Guide to Methods and Applications (Academic Press, San Diego, 1990).
Miyazaki, K. & Takenouchi, M. Creating random mutagenesis libraries using megaprimer PCR of whole plasmid. Biotechniques 33, 1033–1034, 1036–1038 (2002).
Kim, Y.G. & Maas, S. Multiple site mutagenesis with high targeting efficiency in one cloning step. Biotechniques 28, 196–198 (2000).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual 2nd ed. (Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, USA, 1989).
Isalan, M. & Choo, Y. Rapid, high-throughput engineering of sequence-specific zinc finger DNA-binding proteins. Methods Enzymol. 340, 593–609 (2001).
Papworth, C., Bauer, J.C. & Braman, J. Site-directed mutagenesis in one day with >80% efficiency. Strategies 9, 3–4 (1996).
Acknowledgements
The author would like to thank P. Beltrao for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Isalan, M. Construction of semi-randomized gene libraries with weighted oligonucleotide synthesis and PCR. Nat Protoc 1, 468–475 (2006). https://doi.org/10.1038/nprot.2006.68
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.68
This article is cited by
-
Intracellular directed evolution of proteins from combinatorial libraries based on conditional phage replication
Nature Protocols (2017)
-
Engineering orthogonal dual transcription factors for multi-input synthetic promoters
Nature Communications (2016)
-
Iterative saturation mutagenesis (ISM) for rapid directed evolution of functional enzymes
Nature Protocols (2007)
-
Localized transfection with magnetic beads coated with PCR products and other nucleic acids
Nature Protocols (2006)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.