Abstract
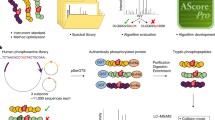
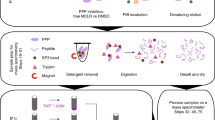
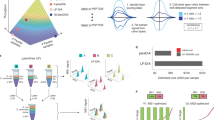
The success in profiling the phosphoproteome by mass spectrometry-based proteomics has been intimately related to the availability of methods that selectively enrich for phosphopeptides. To this end, we describe a protocol that combines two sequential enrichment steps. First, strong cation exchange (SCX) chromatography separates peptides by solution charge. Phosphate groups contribute to solution charge by adding a negative charge at pH 2.7. Therefore, at that pH, phosphopeptides are expected to elute earlier than their nonphosphorylated homologs. Second, immobilized metal affinity chromatography (IMAC) takes advantage of phosphate's affinity for metal ions such as Fe3+ to uniformly enrich for phosphopeptides from the previously collected SCX fractions. We have successfully employed the SCX/IMAC enrichment strategy in the exploration of phosphoproteomes from several systems including mouse liver and Drosophila embryos characterizing over 5,500 and 13,000 phosphorylation events, respectively. The SCX/IMAC enrichment protocol requires 2 days, and the entire procedure from cells to a phosphorylation data set can be completed in less than 10 days.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hastie, C.J., McLauchlan, H.J. & Cohen, P. Assay of protein kinases using radiolabeled ATP: a protocol. Nat. Protoc. 1, 968–971 (2006).
Collins, M.O., Yu, L. & Choudhary, J.S. Analysis of protein phosphorylation on a proteome-scale. Proteomics 7, 2751–2768 (2007).
Hunter, T. & Cooper, J.A. Protein-tyrosine kinases. Annu. Rev. Biochem. 54, 897–930 (1985).
Rush, J. et al. Immunoaffinity profiling of tyrosine phosphorylation in cancer cells. Nat. Biotechnol. 23, 94–101 (2005).
Andersson, L. & Porath, J. Isolation of phosphoproteins by immobilized metal (Fe3+) affinity chromatography. Anal. Biochem. 154, 250–254 (1986).
Posewitz, M.C. & Tempst, P. Immobilized gallium(III) affinity chromatography of phosphopeptides. Anal. Chem. 71, 2883–2892 (1999).
Pinkse, M.W., Uitto, P.M., Hilhorst, M.J., Ooms, B. & Heck, A.J. Selective isolation at the femtomole level of phosphopeptides from proteolytic digests using 2D-NanoLC-ESI-MS/MS and titanium oxide precolumns. Anal. Chem. 76, 3935–3943 (2004).
Sano, A. & Nakamura, H. Titania as a chemo-affinity support for the column-switching HPLC analysis of phosphopeptides: application to the characterization of phosphorylation sites in proteins by combination with protease digestion and electrospray ionization mass spectrometry. Anal. Sci. 20, 861–864 (2004).
Kweon, H.K. & Hakansson, K. Selective zirconium dioxide-based enrichment of phosphorylated peptides for mass spectrometric analysis. Anal. Chem. 78, 1743–1749 (2006).
Holmes, C.F. A new method for the selective isolation of phosphoserine-containing peptides. FEBS Lett. 215, 21–24 (1987).
Oda, Y., Nagasu, T. & Chait, B.T. Enrichment analysis of phosphorylated proteins as a tool for probing the phosphoproteome. Nat. Biotechnol. 19, 379–382 (2001).
Zhou, H., Watts, J.D. & Aebersold, R. A systematic approach to the analysis of protein phosphorylation. Nat. Biotechnol. 19, 375–378 (2001).
Beausoleil, S.A. et al. Large-scale characterization of HeLa cell nuclear phosphoproteins. Proc. Natl. Acad. Sci. USA 101, 12130–12135 (2004).
Li, X. et al. Large-scale phosphorylation analysis of alpha-factor-arrested Saccharomyces cerevisiae. J. Proteome Res. 6, 1190–1197 (2007).
Wilson-Grady, J.T., Villen, J. & Gygi, S.P. Phosphoproteome analysis of fission yeast. J. Proteome Res. 7, 1088–1097 (2008).
Gruhler, A. et al. Quantitative phosphoproteomics applied to the yeast pheromone signaling pathway. Mol. Cell. Proteomics 4, 310–327 (2005).
Trinidad, J.C., Specht, C.G., Thalhammer, A., Schoepfer, R. & Burlingame, A.L. Comprehensive identification of phosphorylation sites in postsynaptic density preparations. Mol. Cell. Proteomics 5, 914–922 (2006).
Villen, J., Beausoleil, S.A., Gerber, S.A. & Gygi, S.P. Large-scale phosphorylation analysis of mouse liver. Proc. Natl. Acad. Sci. USA 104, 1488–1493 (2007).
Zhai, B., Villen, J., Beausoleil, S.A., Mintseris, J. & Gygi, S.P. Phosphoproteome analysis of Drosophila melanogaster embryos. J. Proteome Res. 7, 1675–1682 (2008).
Olsen, J.V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648 (2006).
Pinkse, M.W. et al. Highly robust, automated, and sensitive online TiO2-based phosphoproteomics applied to study endogenous phosphorylation in Drosophila melanogaster. J. Proteome Res. 7, 687–697 (2008).
Wu, J., Shakey, Q., Liu, W., Schuller, A. & Follettie, M.T. Global profiling of phosphopeptides by titania affinity enrichment. J. Proteome Res. 6, 4684–4689 (2007).
Mazanek, M. et al. Titanium dioxide as a chemo-affinity solid phase in offline phosphopeptide chromatography prior to HPLC-MS/MS analysis. Nat. Protoc. 2, 1059–1069 (2007).
Thingholm, T.E., Jorgensen, T.J., Jensen, O.N. & Larsen, M.R. Highly selective enrichment of phosphorylated peptides using titanium dioxide. Nat. Protoc. 1, 1929–1935 (2006).
Bakalarski, C.E., Haas, W., Dephoure, N.E. & Gygi, S.P. The effects of mass accuracy, data acquisition speed, and search algorithm choice on peptide identification rates in phosphoproteomics. Anal. Bioanal. Chem. 389, 1409–1419 (2007).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Eng, J.K., McCormack, A.L. & Yates, J.R. III. An approach to correlate tandem mass-spectral data of peptides with amino-acid-sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Beausoleil, S.A., Villen, J., Gerber, S.A., Rush, J. & Gygi, S.P. A probability-based approach for high-throughput protein phosphorylation analysis and site localization. Nat. Biotechnol. 24, 1285–1292 (2006).
Haas, W. et al. Optimization and use of peptide mass measurement accuracy in shotgun proteomics. Mol. Cell Proteomics 5, 1326–1337 (2006).
Albuquerque, C.P. et al. A multidimensional chromatography technology for in-depth phosphoproteome analysis. Mol. Cell. Proteomics 7, 1389–1396 (2008).
Bodenmiller, B. et al. PhosphoPep—a phosphoproteome resource for systems biology research in Drosophila Kc167 cells. Mol. Syst. Biol. 3, 139 (2007).
Zarling, A.L. et al. Phosphorylated peptides are naturally processed and presented by major histocompatibility complex class I molecules in vivo. J. Exp. Med. 192, 1755–1762 (2000).
Stensballe, A., Jensen, O.N., Olsen, J.V., Haselmann, K.F. & Zubarev, R.A. Electron capture dissociation of singly and multiply phosphorylated peptides. Rapid Commun. Mass Spectrom. 14, 1793–1800 (2000).
Chi, A. et al. Analysis of phosphorylation sites on proteins from Saccharomyces cerevisiae by electron transfer dissociation (ETD) mass spectrometry. Proc. Natl. Acad. Sci. USA 104, 2193–2198 (2007).
Schroeder, M.J., Shabanowitz, J., Schwartz, J.C., Hunt, D.F. & Coon, J.J. A neutral loss activation method for improved phosphopeptide sequence analysis by quadrupole ion trap mass spectrometry. Anal. Chem. 76, 3590–3598 (2004).
Acknowledgements
We thank Andrew Alpert from PolyLC for kindly providing columns for SCX chromatography and Manuel Rodriguez-Falcon for the initial tests of the combined IMAC-desalting procedure. We are also grateful to Joshua T. Wilson-Grady for constructive comments on the manuscript. This work was supported by NIH grant HG3456 to S.P.G.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Villén, J., Gygi, S. The SCX/IMAC enrichment approach for global phosphorylation analysis by mass spectrometry. Nat Protoc 3, 1630–1638 (2008). https://doi.org/10.1038/nprot.2008.150
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.150
This article is cited by
-
Pilot investigation of magnetic nanoparticle–based immobilized metal affinity chromatography for efficient enrichment of phosphoproteoforms for mass spectrometry–based top-down proteomics
Analytical and Bioanalytical Chemistry (2023)
-
Proteomic investigation of neural stem cell to oligodendrocyte precursor cell differentiation reveals phosphorylation-dependent Dclk1 processing
Cellular and Molecular Life Sciences (2023)
-
Global phosphoproteomic analysis identified key kinases regulating male meiosis in mouse
Cellular and Molecular Life Sciences (2022)
-
Listeria monocytogenes exposed to antimicrobial peptides displays differential regulation of lipids and proteins associated to stress response
Cellular and Molecular Life Sciences (2022)
-
A Cdk4/6-dependent phosphorylation gradient regulates the early to late G1 phase transition
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.