Abstract
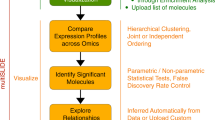
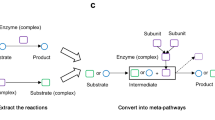
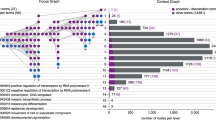
The Systems Biology Graphical Notation (SBGN) is an emerging standard for the uniform representation of biological processes and networks. By using examples from gene regulation and metabolism, this protocol shows the construction of SBGN maps by either manual drawing or automatic translation using the tool SBGN-ED. In addition, it discusses the enrichment of SBGN maps with different kinds of -omics data to bring numerical data into the context of these networks in order to facilitate the interpretation of experimental data. Finally, the export of such maps to public websites, including clickable images, supports the communication of results within the scientific community. With regard to the described functionalities, other tools partially overlap with SBGN-ED. However, currently, SBGN-ED is the only tool that combines all of these functions, including the representation in SBGN, data mapping and website export. This protocol aims to assist scientists with the step-by-step procedure, which altogether takes ∼90 min.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Le Novere, N. et al. The systems biology graphical notation. Nat. Biotechnol. 27, 735–741 (2009).
Junker, A., Hartmann, A., Schreiber, F. & Baumlein, H. An engineer's view on regulation of seed development. Trends Plant Sci. 15, 303–307 (2010).
Caron, E. et al. A comprehensive map of the mTOR signaling network. Mol. Syst. Biol. 6, 453 (2010).
Kierzek, A.M., Zhou, L. & Wanner, B.L. Stochastic kinetic model of two component system signalling reveals all-or-none, graded and mixed mode stochastic switching responses. Mol. Biosyst. 6, 531–542 (2010).
Jansson, A. & Jirstrand, M. Biochemical modeling with systems biology graphical notation. Drug Discov. Today 15, 365–370 (2010).
Funahashi, A. et al. CellDesigner 3.5: a versatile modeling tool for biochemical networks. Proceedings of the IEEE 96, 1254–1265 (2008).
Zinovyev, A., Viara, E., Calzone, L. & Barillot, E. BiNoM: a Cytoscape plugin for manipulating and analyzing biological networks. Bioinformatics 24, 876–877 (2008).
van Iersel, M.P. et al. Presenting and exploring biological pathways with PathVisio. BMC Bioinformatics 9, 399 (2008).
Czauderna, T., Klukas, C. & Schreiber, F. Editing, validating and translating of SBGN maps. Bioinformatics 26, 2340–2341 (2010).
Gehlenborg, N. et al. Visualization of omics data for systems biology. Nat. Methods 7, S56 S (2010).
Suderman, M. & Hallett, M. Tools for visually exploring biological networks. Bioinformatics 23, 2651–2659 (2007).
Sorokin, A. et al. The pathway editor: a tool for managing complex biological networks. IBM J. Res. Dev. 50, 561–573 (2006).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Schreiber, F. et al. MetaCrop 2.0: managing and exploring information about crop plant metabolism. Nucleic Acid Res. published online, doi:10.1093/nar/GKR1004 (15 November, 2011).
Moodie, S., Le Novère, N., Emek, D., Huaiyu, M. & Villeger, A. Systems biology graphical notation: process description language level 1V3. Nature Precedings published online, doi:10.1038/npre.2011.3721.4 (17 February, 2011).
Grafahrend-Belau, E. et al. MetaCrop: a detailed database of crop plant metabolism. Nucleic Acids Res. 36, D954–D958 (2008).
Zimmermann, P., Hirsch-Hoffmann, M., Hennig, L. & Gruissem, W. GENEVESTIGATOR. Arabidopsis microarray database and analysis toolbox. Plant Physiol. 136, 2621–2632 (2004).
Junker, B.H. et al. Temporally regulated expression of a yeast invertase in potato tubers allows dissection of the complex metabolic phenotype obtained following its constitutive expression. Plant Mol. Biol. 56, 91–110 (2004).
Stone, S.L. et al. Arabidopsis LEAFY COTYLEDON2 induces maturation traits and auxin activity: implications for somatic embryogenesis. Proc. Natl. Acad. Sci. USA 105, 3151–3156 (2008).
To, A. et al. A network of local and redundant gene regulation governs Arabidopsis seed maturation. Plant Cell 18, 1642–1651 (2006).
Villeger, A.C., Pettifer, S.R. & Kell, D.B. Arcadia: a visualization tool for metabolic pathways. Bioinformatics 26, 1470–1471 (2010).
Chandran, D., Bergmann, F.T. & Sauro, H.M. Athena: modular CAM/CAD software for synthetic biology. arXiv.org published online, arXiv:0902.2598v1 (2009).
Chabrier-Rivier, N., Fages, F. & Soliman, S. in Computational Methods in Systems Biology. Lecture Notes in Computer Science Vol. 3082 (eds. Danos, V. & Schachter, V.) 172–191 (Springer-Verlag, 2005).
Olivier, B.G. & Snoep, J.L. Web-based kinetic modelling using JWS Online. Bioinformatics 20, 2143–2144 (2004).
Battke, F., Symons, S. & Nieselt, K. Mayday—integrative analytics for expression data. BMC Bioinformatics 11 (2010).
Wegner, K. et al. The NetBuilder' project: development of a tool for constructing, simulating, evolving, and analysing complex regulatory networks. BMC Syst. Biol. 1, P72 (2007).
Deckard, A., Bergmann, F.T. & Sauro, H.M. Supporting the SBML layout extension. Bioinformatics 22, 2966–2967 (2006).
Lammers, C.R. et al. Connecting parts with processes: SubtiWiki and SubtiPathways integrate gene and pathway annotation for Bacillus subtilis. Microbiology 156, 849–859 (2010).
Reyes Palomares, A. et al. Systems biology metabolic modeling assistant: an ontology-based tool for the integration of metabolic data in kinetic modeling. Bioinformatics 25, 834–835 (2009).
Dilek, A., Belviranli, M.E. & Dogrusoz, U. VISIBIOweb: visualization and layout services for BioPAX pathway models. Nucleic Acids Res. 38, W150–W154 (2010).
Elowitz, M. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Acknowledgements
We are grateful to J. Hüge and B. Junker for helpful discussions and advice.
Author information
Authors and Affiliations
Contributions
All authors contributed extensively to the work presented in this paper. A.J. and H.R. wrote the paper. T.C. and C.K. wrote the code. A.H. developed the tutorial. A.H., T.C. and F.S. edited the manuscript. F.S. supervised the project and gave conceptual advice.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Tutorial
Creating interactive, web-based and data-enriched maps using the Systems Biology Graphical Notation. (PDF 10646 kb)
Supplementary Data 1
LEC1/AFL-B3 regulatory network. This file represents the result of the manual drawing process (Steps 9Ai-xxiv). It can directly be loaded into SBGN-ED for further layouting or data mapping. (ZIP 1 kb)
Supplementary Data 2
TCA cycle metabolic network. This file represents the result of the import of a SBGN style metabolic network from the MetaCrop database (Steps 9Bi-iii). It can directly be loaded into SBGN-ED for further layouting or data mapping. (ZIP 2 kb)
Supplementary Data 3
Excel template with expression data. This template file contains expression data obtained from the Genevestigator database for four transcription factors of the LEC1/AFL-B3 regulatory network. It can directly be used for upload into SBGN-ED and for mapping onto the LEC1/AFL-B3 regulatory network as described in Steps 10Ai-x. (XLS 29 kb)
Supplementary Data 4
Mapping table for mapping of expression data. This mapping table provides alternative IDs (AGI gene IDs for Arabidopsis) for the mapping of expression data on the LEC1/AFL-B3 regulatory network which contains only the gene names. (XLS 26 kb)
Supplementary Data 5
Regulatory network mapping. This file shows the LEC1/AFL-B3 regulatory network enriched by expression data as the result of Steps 10Ai-x. It can directly be loaded into SBGN-ED for further layouting or data mapping. (ZIP 3 kb)
Supplementary Data 6
Excel template with metabolic data. This template file contains metabolite measurement data obtained from literature. It can directly be used for upload into SBGN-ED and for mapping onto the TCA cycle metabolic network as described in Steps 10Bi-iii. (XLS 76 kb)
Supplementary Data 7
Metabolic pathway mapping. This file shows the TCA cycle metabolic network enriched by metabolite data as the result of steps 10Bi-iii. It can directly be loaded into SBGN-ED for further layouting or data mapping. (ZIP 8 kb)
Rights and permissions
About this article
Cite this article
Junker, A., Rohn, H., Czauderna, T. et al. Creating interactive, web-based and data-enriched maps with the Systems Biology Graphical Notation. Nat Protoc 7, 579–593 (2012). https://doi.org/10.1038/nprot.2012.002
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.002
This article is cited by
-
Aadh2p: an Arxula adeninivorans alcohol dehydrogenase involved in the first step of the 1-butanol degradation pathway
Microbial Cell Factories (2016)
-
Translation of SBGN maps: Process Description to Activity Flow
BMC Systems Biology (2013)
-
OPTIMAS-DW: A comprehensive transcriptomics, metabolomics, ionomics, proteomics and phenomics data resource for maize
BMC Plant Biology (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.