Abstract
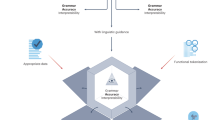
The correspondence between biology and linguistics at the level of sequence and lexical inventories, and of structure and syntax, has fuelled attempts to describe genome structure by the rules of formal linguistics. But how can we define protein linguistic rules? And how could compositional semantics improve our understanding of protein organization and functional plasticity?
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Boguski, M. S. Biosequence exegesis. Science 286, 453–455 (1999).
Baker, M. C. The atoms of language (Basic books, New York, 2001).
Pesole, G., Attimonelli, M. & Saccone, C. Linguistic approaches to the analysis of sequence information. Trends Biotechnol. 12, 401–408 (1994).
Mantegna, R. N et al. Linguistic features of noncoding DNA sequences. Phys. Rev. Lett. 73, 3169–3172 (1994).
Popov, O., Segal, D. M. & Trifonov, E. N. Linguistic complexity of protein sequences as compared to texts of human languages. Biosystems 38, 65–74 (1996).
Doerfler, W. In search of more complex genetic codes — can linguistics be a guide? Med. Hypotheses 9, 563–579 (1982).
Ji, S. Isomorphism between cell and human languages: molecular biological, bioinformatic and linguistic implications. Biosynthesis 44, 17–39 (1997).
Ji, S. & Ciobanu, G. Conformon-driven biopolymer shape changes in cell modelling. Biosystems 70, 165–181 (2002).
Botstein, D. & Cherry, J. M. Molecular linguistics: extracting information from gene and protein sequences. Proc. Natl Acad. Sci. USA 94, 5506–5507 (1997).
Editorial. Folding as grammar. Nature Struct. Biol. 9, 713 (2002).
Brendel, V. & Busse, H. G. Genome structure described by formal languages. Nucleic Acids Res. 12, 2561–2568 (1984).
Brendel, V., Beckman, J. S. & Trifonov, E. N. Linguistics of nucleotide sequences: morphology and comparison of vocabularies. J. Biomol. Struct. Dyn. 4, 11–21 (1986).
Werner, E. Genome semantics, in silico multicellular systems and the central dogma. FEBS Lett. 579, 1779–1782 (2005).
Searls, D. B. Linguistic approaches to biological sequences. Comput. Appl. Biosci. 13, 333–344 (1997).
Searls, D. B. in Artificial Intelligence and Molecular Biology (ed. Hunter, L.) 47–121 (The MIT Press Classics Series and AAAI press, Cambridge, USA, 1993).
Searls, D. B. Using bioinformatics in gene and drug discovery. Drug Discov. Today 5, 135–143 (2000).
Searls, D. B. Reading the book of life. Bioinformatics, 17, 579–580 (2001).
Searls, D. B. The language of genes. Nature, 420, 211–217 (2002).
Searls, D. B. Trees of life and of language, Nature 426, 391–392 (2003).
Dong, S. & Searls, D. B. Gene structure prediction by linguistic methods. Genomics 23, 540–551 (1994).
Koonin, E. V., Wolf, Y. I. & Karev, G. P. The structure of the protein universe and genome evolution. Nature 420, 218–223 (2002).
Modular Protein Domains. (eds Cesareni,G., Gimona, M., Sudol, M. & Yaffe, M.) (WILEY-VCH, Weinheim, 2004).
Papin, J. A., Hunter, T., Palsson, B. O. & Subramaniam, S. Reconstruction of cellular signalling networks and analysis of their properties. Nature Rev. Mol. Cell Biol. 6, 99–111 (2005).
Barabasi, A. -L. & Oltvai, Z. N. Network biology: understanding the cell's functional organization. Nature Rev. Genet. 5, 101–113 (2004).
Han, J. -D. et al. Evidence for dynamically organized modularity in the yeast protein–protein interaction network. Nature 430, 88–93 (2004).
Wuchty, S. Scale-free behaviour in protein domain networks. Mol. Biol. Evol. 18, 1694–1702 (2001).
Hartwell, L. H., Hopfield, J. J., Leibler, S. & Murray, A. W. From molecular to modular cell biology. Nature 402, C47–C52 (1999).
Wuchty, S., Oltvai, Z. N. & Barabasi, A. -L. Evolutionary conservation of motif constituents in the yeast interaction network. Nature Genet. 35, 176–179 (2003).
Pietrokovski, S, Hishon, J. & Trifonov, E. N. Linguistic measure of taxonomic and functional relatedness of nucleotide sequences. J. Biomol. Struct. 7, 1251–1268 (1990).
Pietrokovski, S & Trifonov, E. N. Imported sequences in the mitochondrial yeast genome identified by nucleotide linguistics. Gene 122, 129–137 (1992).
Pawson, T. Protein modules and signalling networks. Nature 373, 573–580 (1995).
Przytycka, T., Aurora, R. & Rose, G. D. A protein taxonomy based on secondary structure. Nature Struct. Biol. 6, 672–682 (1999).
Przytycka, T., Srinivasan, R. & Rose, G. D. Recursive domains in proteins. Prot. Sci. 11, 409–417 (2002).
Sim, J., Kim, S. Y. & Lee, J. PPRODO: prediction of protein domain boundaries using neural networks. Proteins 59, 627–632 (2005).
Sonnhammer, E. L. L. & Kahn, D. Modular arrangement of proteins as inferred from analysis of homology. Prot. Sci. 3, 482–492 (1994).
Galzitskaya, O. V. & Melnik, B. S. Prediction of protein domain boundaries from sequence alone. Prot. Sci. 12, 696–701 (2003).
Aasland, R. et al. Normalization of nomenclature for peptide motifs as ligands of modular protein domains. FEBS Lett. 513, 141–144 (2002).
Arlinghaus, R. B. Bcr: a negative regulator of the Bcr–Abl oncoprotein in leukemia. Oncogene 21, 8560–8567 (2002).
Park, S. -H., Zarrinpar, A. & Lim, W. A. Rewiring MAP kinase pathways using alternative scaffold assembly mechanisms. Science 299, 1061–1064 (2003).
Dyson, H. J. & Wright, P. E. Intrinsically unstructured proteins and their functions. Nature Rev. Mol. Cell Biol. 6, 197–208 (2005).
George, R. A. & Heringa, J. An analysis of protein domain linkers: their classification and role in protein folding. Prot. Eng. 15, 871–879 (2002).
Pawson, T. Specificity in signal transduction: from phosphotyrosine–SH2 domain interactions to complex cellular systems. Cell 116, 191–203 (2004).
Farooq, A., Sudol, M. & Zhou, M. -M. Two is better than one: structure function and mechanism of tandem domains. Nova Publications (in the press).
Benner, S. A. & Gaucher, E. A. Evolution, language and analogy in functional genomics. Trends Genet. 17, 414–418 (2001).
Vidal, M. Interactome modelling FEBS Lett. 579, 1834–1838 (2005).
Zanzoni, A. et al. MINT: a Molecular INTeraction database. FEBS Lett. 513, 135–140 (2002).
Sudol, M. From src homology modules to other signalling domains: proposal of the „Protein Recognition Code”. Oncogene 17, 1469–1474 (1998).
Wuchty, S. & Almaas, E. Evolutionary cores of domain co-occurrence networks. BMC Evol. Biol. 5, 24 (2005).
Acknowledgements
I wish to thank M. C. Baker and M. Sudol for critically commenting on this manuscript, and the members of the Protein Modules Consortium for inspiring discussions. The author is supported by a Marie Curie Excellence Grant of the Framework Program 6 of the European Union.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Related links
DATABASES
Artificial Intelligence and Molecular Biology (electronic text (PDF) of the out-of-print book)
The DIMA domain interaction map
FURTHER INFORMATION
Glossary
- Affix
-
A meaningful element that cannot stand on its own but it is added to another element.
- Automaton
-
A device that reads input, conventionally from left to right, and either recognizes or generates language.
- Clause
-
A basic unit of grammatical structure that expresses a single thought.
- Grammar
-
The part of a language that is responsible for assembling basic words into larger words, phrases and clauses in systematic ways. For simplicity, grammar may be viewed as a combination of syntax and morphology.
- Lexica
-
The stocks of basic words.
- Linguistics
-
The study of the nature, structure and variation of language (includes the sub-disciplines of morphology, syntax, semantics and pragmatics).
- Module
-
Different languages have different concepts of a module but there are several shared ideas. Modules are similar to objects in an object-orientated language, although a module might contain many procedures and/or functions, which would correspond to many objects. In computer science, modules are described as a portion of a program that carries out a specific function and might be used alone or combined with other modules of the same program.
- Phrase
-
A group of words that appear next to each other or stay together in the arrangement of a sentence and that form a syntactic unit.
- Prefix
-
A meaningful element that cannot stand on its own but it is added to the beginning of another element.
- Root
-
The core of a word, before prefixes and suffixes are attached.
- Semantics
-
The branch of linguistics concerned with the meaning of linguistic expression.
- Sentence
-
A basic unit of a language that expresses a complete thought.
- Stem
-
Prefixes and suffixes attach to a stem in order to form a longer word.
- Suffix
-
A meaningful element that cannot stand on its own but it is added to the end of another element.
- Syntax
-
The branch of linguistics that studies how words are combined to make phrases and sentences.
- Word
-
A freestanding portion of language with a coherent meaning.
Rights and permissions
About this article
Cite this article
Gimona, M. Protein linguistics — a grammar for modular protein assembly?. Nat Rev Mol Cell Biol 7, 68–73 (2006). https://doi.org/10.1038/nrm1785
Issue Date:
DOI: https://doi.org/10.1038/nrm1785
This article is cited by
-
Probing ion channel functional architecture and domain recombination compatibility by massively parallel domain insertion profiling
Nature Communications (2021)
-
On the Verge of Life: Distribution of Nucleotide Sequences in Viral RNAs
Biosemiotics (2021)
-
Unity and disunity in evolutionary sciences: process-based analogies open common research avenues for biology and linguistics
Biology Direct (2016)
-
Evolutionary dynamics of selfish DNA explains the abundance distribution of genomic subsequences
Scientific Reports (2016)
-
Probabilistic grammatical model for helix‐helix contact site classification
Algorithms for Molecular Biology (2013)