Abstract
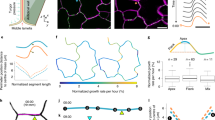
The emerging field of computational morphodynamics aims to understand the changes that occur in space and time during development by combining three technical strategies: live imaging to observe development as it happens; image processing and analysis to extract quantitative information; and computational modelling to express and test time-dependent hypotheses. The strength of the field comes from the iterative and combined use of these techniques, which has provided important insights into plant development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Reddy, G. V. & Roy-Chowdhury, A. Live-imaging and image processing of shoot apical meristems of Arabidopsis thaliana. Methods Mol. Biol. 553, 305–316 (2009).
Ball, E. Cell division patterns in living shoot apices. Phytomorphology 10, 377–396 (1960).
Dumais, J. & Kwiatkowska, D. Analysis of surface growth in shoot apices. Plant J. 31, 229–241 (2002).
Reddy, G. V., Gordon, S. P. & Meyerowitz, E. M. Unravelling developmental dynamics: transient intervention and live imaging in plants. Nature Rev. Mol. Cell Biol. 8, 491–501 (2007).
Chickarmane, V. et al. Computational morphodynamics: a modeling framework to understand plant growth. Annu. Rev. Plant Biol. 61, 65–87 (2010).
Oates, A. C., Gorfinkiel, N., González-Gaitán, M. & Heisenberg, C. P. Quantitative approaches in developmental biology. Nature Rev. Genet. 10, 517–530 (2009).
Grieneisen, V. A. & Scheres, B. Back to the future: evolution of computational models in plant morphogenesis. Curr. Opin. Plant Biol. 12, 606–614 (2009).
Sawai, S., Thomason, P. A. & Cox, E. C. An autoregulatory circuit for long-range self-organization in Dictyostelium cell populations. Nature 433, 323–326 (2005).
Aigouy, B. et al. Cell flow reorients the axis of planar polarity in the wing epithelium of Drosophila. Cell 142, 773–786.
Amonlirdviman, K. et al. Mathematical modeling of planar cell polarity to understand domineering nonautonomy. Science 307, 423–426 (2005).
Vavylonis, D., Wu, J. Q., Hao, S., O'Shaughnessy, B. & Pollard, T. D. Assembly mechanism of the contractile ring for cytokinesis by fission yeast. Science 319, 97–100 (2008).
Fayant, P. et al. Finite element model of polar growth in pollen tubes. Plant Cell 22, 2579–2593 (2010).
Scarpella, E., Marcos, D., Friml, J. & Berleth, T. Control of leaf vascular patterning by polar auxin transport. Genes Dev. 20, 1015–1027 (2006).
Bayer, E. M. et al. Integration of transport-based models for phyllotaxis and midvein formation. Genes Dev. 23, 373–384 (2009).
Galway, M. E. et al. The TTG gene is required to specify epidermal cell fate and cell patterning in the Arabidopsis root. Dev. Biol. 166, 740–754 (1994).
Kwak, S. H., Shen, R. & Schiefelbein, J. Positional signaling mediated by a receptor-like kinase in Arabidopsis. Science 307, 1111–1113 (2005).
Benítez, M., Espinosa-Soto, C., Padilla-Longoria, P., Díaz, J. & Alvarez-Buylla, E. R. Equivalent genetic regulatory networks in different contexts recover contrasting spatial cell patterns that resemble those in Arabidopsis root and leaf epidermis: a dynamic model. Int. J. Dev. Biol. 51, 139–155 (2007).
Savage, N. S. et al. A mutual support mechanism through intercellular movement of CAPRICE and GLABRA3 can pattern the Arabidopsis root epidermis. PLoS Biol. 6, e235 (2008).
Lee, M. M. & Schiefelbein, J. Cell pattern in the Arabidopsis root epidermis determined by lateral inhibition with feedback. Plant Cell 14, 611–618 (2002).
Steeves, T. A. & Sussex, I. M. Patterns in Plant Development (Cambridge Univ. Press, Cambridge, UK, 1989).
Hohm, T., Zitzler, E. & Simon, R. A dynamic model for stem cell homeostasis and patterning in Arabidopsis meristems. PLoS ONE 5, e9189 (2010).
Jönsson, H. et al. Modeling the organization of the WUSCHEL expression domain in the shoot apical meristem. Bioinformatics 21, I232–I240 (2005).
Nikolaev, S. V., Penenko, A. V., Lavreha, V. V., Mjolsness, E. D. & Kolchanov, N. A. A model study of the role of proteins CLV1, CLV2, CLV3, and WUS in regulation of the structure of the shoot apical meristem. Russ. J. Dev. Biol. 38, 383–388 (2007).
Reddy, G. V. & Meyerowitz, E. M. Stem-cell homeostasis and growth dynamics can be uncoupled in the Arabidopsis shoot apex. Science 310, 663–667 (2005).
Geier, F. et al. A quantitative and dynamic model for plant stem cell regulation. PLoS ONE 3, e3553 (2008).
Domagalska, M. A. & Leyser, O. Signal integration in the control of shoot branching. Nature Rev. Mol. Cell Biol. 12, 211–221 (2011).
Mundermann, L., Erasmus, Y., Lane, B., Coen, E. & Prusinkiewicz, P. Quantitative modeling of Arabidopsis development. Plant Physiol. 139, 960–968 (2005).
Hall, S. M. & Hillman, J. R. Correlative inhibition of lateral bud growth in Phaseolus vulgaris L. timing of bud growth following decapitation. Planta 123, 137–143 (1975).
Thimann, K. V., Skoog, F. & Kerckhoff, W. G. On the inhibition of bud development and other functions of growth substance in Vicia faba. Proc. R. Soc. Lond. B 114, 317–339 (1934).
Stirnberg, P., Ward, S. & Leyser, O. Auxin and strigolactones in shoot branching: intimately connected? Biochem. Soc. Trans. 38, 717–722 (2010).
Prusinkiewicz, P. et al. Control of bud activation by an auxin transport switch. Proc. Natl Acad. Sci. USA 106, 17431–17436 (2009).
Bossinger, G. & Smyth, D. R. Initiation patterns of flower and floral organ development in Arabidopsis thaliana. Development 122, 1093–1102 (1996).
Irish, V. F. & Sussex, I. M. A fate map of the Arabidopsis embryonic shoot apical meristem. Development 115, 745–753 (1992).
Poethig, R. S. & Sussex, I. M. The cellular parameters of leaf development in tobacco: a clonal analysis. Planta 165, 170–184 (1985).
Green, A. A., Kennaway, J. R., Hanna, A. I., Bangham, J. A. & Coen, E. Genetic control of organ shape and tissue polarity. PLoS Biol. 8, e1000537 (2010).
Cui, M. L., Copsey, L., Green, A. A., Bangham, J. A. & Coen, E. Quantitative control of organ shape by combinatorial gene activity. PLoS Biol. 8, e1000538 (2010).
Campilho, A., Garcia, B., Toorn, H. V., Wijk, H. V. & Scheres, B. Time-lapse analysis of stem-cell divisions in the Arabidopsis thaliana root meristem. Plant J. 48, 619–627 (2006).
Harrison, C. J., Roeder, A. H., Meyerowitz, E. M. & Langdale, J. A. Local cues and asymmetric cell divisions underpin body plan transitions in the moss Physcomitrella patens. Curr. Biol. 19, 461–471 (2009).
Reddy, G. V., Heisler, M. G., Ehrhardt, D. W. & Meyerowitz, E. M. Real-time lineage analysis reveals oriented cell divisions associated with morphogenesis at the shoot apex of Arabidopsis thaliana. Development 131, 4225–4237 (2004).
Roeder, A. H. K. et al. Variability in the control of cell division underlies sepal epidermal patterning in Arabidopsis thaliana. PLoS Biol. 8, e1000367 (2010).
Fernandez, R. et al. Imaging plant growth in 4D: robust tissue reconstruction and lineaging at cell resolution. Nature Methods 7, 547–553 (2010).
Traas, J., Hulskamp, M., Gendreau, E. & Hofte, H. Endoreduplication and development: rule without dividing? Curr. Opin. Plant Biol. 1, 498–503 (1998).
Grieneisen, V. A., Xu, J., Maree, A. F., Hogeweg, P. & Scheres, B. Auxin transport is sufficient to generate a maximum and gradient guiding root growth. Nature 449, 1008–1013 (2007).
Jönsson, H., Heisler, M. G., Shapiro, B. E., Meyerowitz, E. M. & Mjolsness, E. An auxin-driven polarized transport model for phyllotaxis. Proc. Natl Acad. Sci. USA 103, 1633–1638 (2006).
Smith, R. S. et al. A plausible model of phyllotaxis. Proc. Natl Acad. Sci. USA 103, 1301–1306 (2006).
Fortin, M. C., Pierce, F. J. & Poff, K. L. The pattern of secondary root formation in curving roots of Arabidopsis thaliana (L.) Heynh. Plant Cell Environ. 12, 337–339 (1989).
De Smet, I. et al. Auxin-dependent regulation of lateral root positioning in the basal meristem of Arabidopsis. Development 134, 681–690 (2007).
Boerjan, W. et al. superroot, a recessive mutation in Arabidopsis, confers auxin overproduction. Plant Cell 7, 1405–1419 (1995).
Dubrovsky, J. G. et al. Auxin acts as a local morphogenetic trigger to specify lateral root founder cells. Proc. Natl Acad. Sci. USA 105, 8790–8794 (2008).
Fukaki, H., Tameda, S., Masuda, H. & Tasaka, M. Lateral root formation is blocked by a gain-of-function mutation in the SOLITARY-ROOT/IAA14 gene of Arabidopsis. Plant J. 29, 153–168 (2002).
Okada, K., Ueda, J., Komaki, M. K., Bell, C. J. & Shimura, Y. Requirement of the auxin polar transport system in early stages of Arabidopsis floral bud formation. Plant Cell 3, 677–684 (1991).
Reinhardt, D., Mandel, T. & Kuhlemeier, C. Auxin regulates the initiation and radial position of plant lateral organs. Plant Cell 12, 507–518 (2000).
Benkova, E. et al. Local, efflux-dependent auxin gradients as a common module for plant organ formation. Cell 115, 591–602 (2003).
Heisler, M. G. et al. Patterns of auxin transport and gene expression during primordium development revealed by live imaging of the Arabidopsis inflorescence meristem. Curr. Biol. 15, 1899–1911 (2005).
Reinhardt, D. et al. Regulation of phyllotaxis by polar auxin transport. Nature 426, 255–260 (2003).
Vernoux, T., Kronenberger, J., Grandjean, O., Laufs, P. & Traas, J. PIN-FORMED 1 regulates cell fate at the periphery of the shoot apical meristem. Development 127, 5157–5165 (2000).
Hamant, O. et al. Developmental patterning by mechanical signals in Arabidopsis. Science 322, 1650–1655 (2008).
Heisler, M. G. et al. Alignment between PIN1 polarity and microtubule orientation in the shoot apical meristem reveals a tight coupling between morphogenesis and auxin transport PLoS Biol. 8, e1000516 (2010).
Laskowski, M. et al. Root system architecture from coupling cell shape to auxin transport. PLoS Biol. 6, e307 (2008).
Gor, V., Elowitz, M., Bacarian, T. & Mjolsness, E. Tracking cell signals in fluorescent images in Proc. 2005 IEEE Comp. Soc. Conf. on Computer Vision and Pattern Recognition 142 (2005).
Liu, M., Roy-Chowdhury, A. K. & Reddy, G. V. Robust estimation of stem cell lineages using local graph matching in IEEE Comp. Soc. Workshop on Mathematical Methods in Biomedical Image Analysis 194 (2009).
Cunha, A., Roeder, A. & Meyerowitz, E. M. Segmenting the sepal and shoot apical meristem of Arabidopsis thaliana in Eng. Med. Biol. Soc. (EMBC), 2010 Ann. Int. Conf. IEEE 5338–5342 (2010).
Clark, S. E. Cell signalling at the shoot meristem. Nature Rev. Mol. Cell Biol. 2, 276–284 (2001).
Nishimura, A., Tamaoki, M., Sato, Y. & Matsuoka, M. The expression of tobacco knotted1-type class 1 homeobox genes correspond to regions predicted by the cytohistological zonation model. Plant J. 18, 337–347 (1999).
Xie, M., Tataw, M. & Venugopala Reddy, G. Towards a functional understanding of cell growth dynamics in shoot meristem stem-cell niche. Semin. Cell Dev. Biol. 20, 1126–1133 (2009).
Sandersius, S. A. & Newman, T. J. Modeling cell rheology with the Subcellular Element Model. Phys. Biol. 5, 015002 (2008).
Van Liedekerke, P. et al. A particle-based model to simulate the micromechanics of single-plant parenchyma cells and aggregates. Phys. Biol. 7, 026006 (2010).
Acknowledgements
We apologize to the numerous members of these fields whose work we could not include owing to space limitations. We thank E. Mjolsness, W. Li, Y. Zhou and L. Ben-Ghaly for insightful comments. We acknowledge support from the Gordon and Betty Moore Cell Center at California Institute of Technology, the US National Institutes of Health (grants F32GM090543 to P.T.T. and R01 GM086639 to E.M.M.), the US Department of Energy (grant DE-FG02-88ER13873 to E.M.M.) and the US National Science Foundation (grant IOS-0846192 to E.M.M.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Related links
Rights and permissions
About this article
Cite this article
Roeder, A., Tarr, P., Tobin, C. et al. Computational morphodynamics of plants: integrating development over space and time. Nat Rev Mol Cell Biol 12, 265–273 (2011). https://doi.org/10.1038/nrm3079
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3079
This article is cited by
-
An optimized pipeline for live imaging whole Arabidopsis leaves at cellular resolution
Plant Methods (2023)
-
What determines cell size?
BMC Biology (2012)
-
The Measurement of Local Variation in Shape
Evolutionary Biology (2012)
-
Flowering and Plant Development at the 38th Spanish Society of Genetics Congress, Murcia, 2011
Journal of Plant Growth Regulation (2012)
-
Mutually reinforcing patterning mechanisms
Nature Reviews Molecular Cell Biology (2011)