Key Points
-
Findings from next-generation sequencing (NGS) have led to a shift in phenotypic boundaries and reclassifications of some kidney diseases
-
NGS techniques are a valuable addition to the diagnostic toolbox in nephrology and findings from NGS can have important implications for therapeutic strategies and clinical outcomes
-
Interpretation of genetic variants and accurate prediction of the associated kidney phenotype can be challenging despite the increasing availability of bioinformatics tools and functional tests
-
Data sharing initiatives are imperative to establish clinically useful genotype–phenotype correlations and to maximize the benefit of genetic testing in routine nephrology practice
Abstract
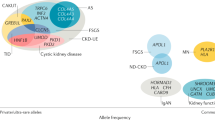
Next-generation sequencing (NGS) has led to the identification of previously unrecognized phenotypes associated with classic kidney disease genes. In addition to improving diagnostics for genetically heterogeneous diseases and enabling a faster rate of gene discovery, NGS has enabled an expansion and redefinition of nephrogenetic disease categories. Findings from these studies raise the question of whether disease diagnoses should be made on clinical grounds, on genetic evidence or a combination thereof. Here, we discuss the major kidney disease-associated genes and gene categories for which NGS has expanded the phenotypic spectrum. For example, COL4A3–5 genes, which are classically associated with Alport syndrome, are now understood to also be involved in the aetiology of focal segmental glomerulosclerosis. DGKE, which is associated with nephrotic syndrome, is also mutated in patients with atypical haemolytic uraemic syndrome. We examine how a shared genetic background between diverse clinical phenotypes can provide insight into the function of genes and novel links with essential pathophysiological mechanisms. In addition, we consider genetic and epigenetic factors that contribute to the observed phenotypic heterogeneity of kidney diseases and discuss the challenges in the interpretation of genetic data. Finally, we discuss the implications of the expanding phenotypic spectra associated with kidney disease genes for clinical practice, genetic counselling and personalized care, and present our recommendations for the use of NGS-based tests in routine nephrology practice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Eisenberger, T. et al. An efficient and comprehensive strategy for genetic diagnostics of polycystic kidney diseas. PLoS ONE 10, e0116680 (2015).
Tavira, B. et al. A labor and cost effective next generation sequencing of PKHD1 in autosomal recessive polycystic kidney disease patients. Gene 561, 165–169 (2015).
Morinière, V. et al. Improving mutation screening in familial hematuric nephropathies through next generation sequencing. J. Am. Soc. Nephrol. 25, 2740–2751 (2014).
Lohmann, K. & Klein, C. Next generation sequencing and the future of genetic diagnosis. Neurotherapeutics 11, 699–707 (2014).
Sampson, M. G. et al. Using population genetics to interrogate the monogenic nephrotic syndrome diagnosis in a case cohort. J. Am. Soc. Nephrol. 27, 1–14 (2015).
Otto, E. A. et al. Candidate exome capture identifies mutation of SDCCAG8 as the cause of a retinal-renal ciliopathy. Nat. Genet. 42, 840–850 (2010).
Gupta, I. R. et al. ARHGDIA: a novel gene implicated in nephrotic syndrome. J. Med. Genet. 50, 330–338 (2013).
Gee, H. Y. et al. Mutations in EMP2 cause childhood-onset nephrotic syndrome. Am. J. Hum. Genet. 94, 884–890 (2014).
Gbadegesin, R. A. et al. Mutations in the gene that encodes the F-actin binding protein anillin cause FSGS. J. Am. Soc. Nephrol. 25, 1991–2002 (2014).
Saisawat, P. et al. Whole-exome resequencing reveals recessive mutations in TRAP1 in individuals with CAKUT and VACTERL association. Kidney Int. 85, 1310–1317 (2014).
Humbert, C. et al. Integrin alpha 8 recessive mutations are responsible for bilateral renal agenesis in humans. Am. J. Hum. Genet. 94, 288–294 (2014).
Vivante, A. & Hildebrandt, F. Exploring the genetic basis of early-onset chronic kidney disease. Nat. Rev. Nephrol. 12, 133–146 (2016).
Nassirpour, R. et al. Identification of tubular injury microRNA biomarkers in urine: comparison of next-generation sequencing and qPCR-based profiling platforms. BMC Genomics 15, 485 (2014).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Marsaud, A. et al. Dismantling papillary renal cell carcinoma classification: the heterogeneity of genetic profiles suggests several independent diseases. Genes Chromosomes Cancer 54, 369–382 (2015).
Lan, J. & Zhang, Q. Clinical applications of next-generation sequencing in histocompatibility and transplantation. Curr. Opin. Organ Transplant. 20, 461–7 (2015).
Sun, Y. et al. Next-generation diagnostics: gene panel, exome, or whole genome? Hum. Mutat. 36, 648–655 (2015).
Bergmann, C. ARPKD and early manifestations of ADPKD: the original polycystic kidney disease and phenocopies. Pediatr. Nephrol. 30, 651–16 (2014).
Gee, H. Y. et al. Whole-exome resequencing distinguishes cystic kidney diseases from phenocopies in renal ciliopathies. Kidney Int. 85, 880–7 (2014).
Lemaire, M. et al. Recessive mutations in DGKE cause atypical hemolytic-uremic syndrome. Nat. Genet. 45, 531–536 (2013).
Mele, C. et al. Characterization of a new DGKE intronic mutation in genetically unsolved cases of familial atypical hemolytic uremic syndrome. Clin. J. Am. Soc. Nephrol. 10, 1011–1019 (2015).
Westland, R. et al. Phenotypic expansion of DGKE-associated diseases. J. Am. Soc. Nephrol. 25, 1408–14 (2014).
Kashtan, C. in GeneReviews 1–22 (Univ. of Washington, 2015).
Artuso, R. et al. Advances in Alport syndrome diagnosis using next-generation sequencing. Eur. J. Hum. Genet. 20, 50–7 (2012).
Braun, D. A. et al. Whole exome sequencing identifies causative mutations in the majority of consanguineous or familial cases with childhood-onset increased renal echogenicity. Kidney Int. 89, 468–475 (2016).
Nogueira, M. et al. Thin basement membrane disease with heavy proteinuria or nephrotic syndrome at presentation. Am. J. Kidney Dis. 35, E15 (2000).
Voskarides, K. et al. COL4A3/COL4A4 mutations producing focal segmental glomerulosclerosis and renal failure in thin basement membrane nephropathy. J. Am. Soc. Nephrol. 18, 3004–3016 (2007).
Pierides, A. et al. Clinico-pathological correlations in 127 patients in 11 large pedigrees, segregating one of three heterozygous mutations in the COL4A3/ COL4A4 genes associated with familial haematuria and significant late progression to proteinuria and chronic kidney dis. Nephrol. Dial. Transplant. 24, 2721–9 (2009).
Chatterjee, R. et al. Targeted exome sequencing integrated with clinicopathological information reveals novel and rare mutations in atypical, suspected and unknown cases of Alport syndrome or proteinuria. PLoS ONE 8, e76360 (2013).
McCarthy, H. J. et al. Simultaneous sequencing of 24 genes associated with steroid-resistant nephrotic syndrome. Clin. J. Am. Soc. Nephrol. 8, 637–648 (2013).
Gibson, J. et al. Exome analysis resolves differential diagnosis of familial kidney disease and uncovers a potential confounding variant. Genet. Res. 95, 165–73 (2013).
Bullich, G. et al. Targeted next-generation sequencing in steroid-resistant nephrotic syndrome: mutations in multiple glomerular genes may influence disease severity. Eur. J. Hum. Genet. 23, 1192–1199 (2015).
Xie, J. et al. COL4A3 mutations cause focal segmental glomerulosclerosis. J. Mol. Cell. Biol. 6, 498–505 (2014).
Malone, A. F. et al. Rare hereditary COL4A3/COL4A4 variants may be mistaken for familial focal segmental glomerulosclerosis. Kidney Int. 86, 1253–1259 (2014).
Gast, C. et al. Collagen (COL4A) mutations are the most frequent mutations underlying adult focal segmental glomerulosclerosis. Nephrol. Dial. Transplant. 31, 961–970 (2016).
Deltas, C., Savva, I., Voskarides, K., Papazachariou, L. & Pierides, A. Carriers of autosomal recessive alport syndrome with thin basement membrane nephropathy presenting as focal segmental glomerulosclerosis in later life. Nephron 130, 271–280 (2015).
Savige, J. et al. Thin basement membrane nephropathy. Kidney Int. 64, 1169–1178 (2003).
Miner, J. H. Pathology versus molecular genetics: (re)defining the spectrum of Alport syndrome. Kidney Int. 86, 1084–1086 (2014).
Ozaltin, F. et al. DGKE variants cause a glomerular microangiopathy that mimics membranoproliferative GN. J. Am. Soc. Nephrol. 24, 377–384 (2013).
Sánchez Chinchilla, D. et al. Complement mutations in diacylglycerol kinase-ε-associated atypical hemolytic uremic syndrome. Clin. J. Am. Soc. Nephrol. 9, 1611–1619 (2014).
Sadowski, C. E. et al. A single-gene cause in 29.5% of cases of steroid-resistant nephrotic syndrome. J. Am. Soc. Nephrol. 26, 1279–1289 (2015).
Noris, M. et al. Relative role of genetic complement abnormalities in sporadic and familial aHUS and their impact on clinical phenotype. Clin. J. Am. Soc. Nephrol. 5, 1844–1859 (2010).
Licht, C. & Fremeaux-Bacchi, V. Hereditary and acquired complement dysregulation in membranoproliferative glomerulonephritis. Thromb. Haemost. 101, 1271–278 (2009).
Noris, M., Mele, C. & Remuzzi, G. Podocyte dysfunction in atypical haemolytic uraemic syndrome. Nat. Rev. Nephrol. 11, 245–252 (2015).
Bruneau, S. et al. Loss of DGKε induces endothelial cell activation and death independently of complement activation. Blood 125, 1038–1046 (2015).
Offermanns, S. Activation of platelet function through G protein – coupled receptors. Circ. Res. 99, 1293–1304 (2006).
Takano, T. & Cybulsky, A. V. Complement C5b-9-mediated arachidonic acid metabolism in glomerular epithelial cells role of cyclooxygenase-1 and -2. Am. J. Pathol. 156, 2091–2101 (2000).
Winn, M. P., Daskalakis, N., Spurney, R. F. & Middleton, J. P. Unexpected role of TRPC6 channel in familial nephrotic syndrome: does it have clinical implications? J. Am. Soc. Nephrol. 378–387 (2006).
Renner, B. et al. Cyclosporine induces endothelial cell release of complement-activating microparticles. J. Am. Soc. Nephrol. 24, 1849–1862 (2013).
Tran, P. V. et al. THM1 negatively modulates mouse sonic hedgehog signal transduction and affects retrograde intraflagellar transport in cilia. Nat. Genet. 40, 403–410 (2008).
Davis, E. E. et al. TTC21B contributes both causal and modifying alleles across the ciliopathy spectrum. Nat. Genet. 43, 189–196 (2011).
Huynh Cong, E. et al. A homozygous missense mutation in the ciliary gene TTC21B causes familial FSGS. J. Am. Soc. Nephrol. 25, 2435–2443 (2014).
Bullich, G. et al. Contribution of the TTC21B gene to glomerular and cystic kidney diseases. Nephrol. Dial. Transplant. http://dx.doi.org/10.1093/ndt/gfv453 (2016).
Romani, M. et al. Mutations in B9D1 and MKS1 cause mild Joubert syndrome: expanding the genetic overlap with the lethal ciliopathy Meckel syndrome. Orphanet J. Rare Dis. 9, 72 (2014).
Slaats, G. G. et al. MKS1 regulates ciliary INPP5E levels in Joubert syndrome. J. Med. Genet. 53, 62–72 (2016).
Thomas, S. et al. A homozygous PDE6D mutation in Joubert syndrome impairs targeting of farnesylated INPP5E protein to the primary cilium. Hum. Mutat. 35, 137–146 (2014).
Bielas, S. L. et al. Mutations in INPP5E, encoding inositol polyphosphate-5-phosphatase E, link phosphatidyl inositol signaling to the ciliopathies. Nat. Genet. 41, 1032–1036 (2009).
Fehrenbach, H. et al. Mutations in WDR19 encoding the intraflagellar transport component IFT144 cause a broad spectrum of ciliopathies. Pediatr. Nephrol. 29, 1451–1456 (2014).
Habbig, S. & Liebau, M. C. Ciliopathies — from rare inherited cystic kidney diseases to basic cellular function. Mol. Cell. Pediatr. 2, 8–13 (2015).
Drivas, T. G., Wojno, A. P., Tucker, B. A., Stone, E. M. & Bennett, J. Basal exon skipping and genetic pleiotropy: a predictive model of disease pathogenesis. Sci. Transl. Med. 7, 291ra97 (2015).
Littink, K. W. et al. A novel nonsense mutation in CEP290 induces exon skipping and leads to a relatively mild retinal phenotype. Invest. Opthalmol. Vis. Sci. 51, 3646–3652 (2010).
Hoefele, J. et al. Evidence of oligogenic inheritance in nephronophthisis. J. Am. Soc. Nephrol. 18, 2789–2795 (2007).
Royer-Pokora, B. et al. Twenty-four new cases of WT1 germline mutations and review of the literature: genotype / phenotype correlations for wilms tumor development. Am. J. Med. Genet. A 257, 249–257 (2004).
Dome, J. & Huff, V. in GeneReviews (eds Pagon, R. A. et al.) 1–24 (Univ. of Washington, 2015).
Lipska, B. S. et al. Genotype–phenotype associations in WT1 glomerulopathy. Kidney Int. 85, 1169–1178 (2014).
Nielsen, S. M. et al. Von Hippel-Lindau disease: genetics and role of genetic counseling in a multiple neoplasia syndrome. J. Clin. Oncol. http://dx.doi.org/10.1200/JCO.2015.65.6140 (2016).
Stanescu, D. E., Hughes, N., Kaplan, B., Stanley, C. a. & De León, D. D. Novel presentations of congenital hyperinsulinism due to mutations in the MODY genes: HNF1A and HNF4A. J. Clin. Endocrinol. Metab. 97, 1–5 (2012).
Hamilton, A. J. et al. The HNF4A R76W mutation causes atypical dominant Fanconi syndrome in addition to a β cell phenotype. J. Med. Genet. 51, 165–169 (2014).
Edwards, N. et al. A novel LMX1B mutation in a family with end-stage renal disease of 'unknown cause'. Clin. Kidney J. 8, 113–119 (2014).
Isojima, T. et al. LMX1B mutation with residual transcriptional activity as a cause of isolated glomerulopathy. Nephrol. Dial. Transplant. 29, 81–88 (2014).
Boyer, O. et al. LMX1B mutations cause hereditary FSGS without extrarenal involvement. J. Am. Soc. Nephrol. 24, 1216–1222 (2013).
Hoopes, R. R. et al. Dent disease with mutations in OCRL1. Am. J. Hum. Genet. 76, 260–267 (2005).
Saisawat, P. et al. Identification of two novel CAKUT-causing genes by massively parallel exon resequencing of candidate genes in patients with unilateral renal agenesis. Kidney Int. 81, 196–200 (2012).
Kohl, S. et al. Mild recessive mutations in six Fraser Syndrome-related genes cause isolated congenital anomalies of the kidney and urinary tract. J. Am. Soc. Nephrol. 25, 1917–1922 (2014).
Adam, J. et al. Genetic testing can resolve diagnostic confusion in Alport syndrome. Clin. Kidney J. 7, 197–200 (2014).
Choi, M. et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc. Natl Acad. Sci. USA 106, 19096–19101 (2009).
Besbas, N., Ozaltin, F., Jeck, N., Seyberth, H. & Ludwig, M. CLCN5 mutation (R347X) associated with hypokalaemic metabolic alkalosis in a Turkish child: an unusual presentation of Dent's disease. Nephrol. Dial. Transplant. 20, 1476–1479 (2005).
Bogdanovic, R. et al. A novel CLCN5 mutation in a boy with Bartter-like syndrome and partial growth hormone deficiency. Pediatr. Nephrol. 25, 2363–2368 (2010).
Okamoto, T., Tajima, T., Hirayama, T. & Sasaki, S. A patient with Dent disease and features of Bartter syndrome caused by a novel mutation of CLCN5. Eur. J. Pediatr. 171, 401–404 (2012).
Sethi, S. K. et al. A boy with proteinuria and focal global glomerulosclerosis: answers. Pediatr. Nephrol. 30, 1945–1946 (2015).
Copelovitch, L., Nash, M. A. & Kaplan, B. S. Hypothesis: Dent disease is an underrecognized cause of focal glomerulosclerosis. J. Am. Soc. Nephrol. 2, 914–918 (2007).
Frishberg, Y. et al. Dent' s disease manifesting as focal glomerulosclerosis: is it the tip of the iceberg? Pediatr. Nephrol. 24, 2369–2373 (2009).
Valina, M. et al. A novel CLCN5 mutation in a boy with asymptomatic proteinuria and focal global glomerulosclerosis. Clin. Nephrol. 80, 377–384 (2013).
Cramer, M. T. et al. Expanding the phenotype of proteinuria in Dent disease. A case series. Pediatr. Nephrol. 29, 2051–2054 (2014).
Johnston, J. J. et al. Individualized iterative phenotyping for genome-wide analysis of loss-of-function mutations. Am. J. Hum. Genet. 96, 913–925 (2015).
Verhave, J. C., Bech, A. P., Wetzels, J. F. M. & Nijenhuis, T. Hepatocyte nuclear factor 1β-associated kidney disease: more than renal cysts and diabetes. J. Am. Soc. Nephrol. 27, 345–353 (2016).
Bergmann, C. et al. Mutations in multiple PKD genes may explain early and severe polycystic kidney disease. J. Am. Soc. Nephrol. 22, 2047–2056 (2011).
Khanna, H. et al. A common allele in RPGRIP1L is a modifier of retinal degeneration in ciliopathies. Nat. Genet. 41, 739–745 (2009).
Leitch, C. C. et al. Hypomorphic mutations in syndromic encephalocele genes are associated with Bardet–Biedl syndrome. Nat. Genet. 40, 443–448 (2008).
Renkema, K. Y., Stokman, M. F., Giles, R. H. & Knoers, N. V. A. M. Next-generation sequencing for research and diagnostics in kidney disease. Nat. Rev. Nephrol. 10, 433–444 (2014).
Saunier, S. et al. A novel gene that encodes a protein with a putative src homology 3 domain is a candidate gene for familial juvenile nephronophthisis. Hum. Mol. Genet. 6, 2317–2323 (1997).
Hildebrandt, F. et al. A novel gene encoding an SH3 domain protein is mutated in nephronophthisis type 1. Nat. Genet. 17, 149–153 (1997).
Arts, H. H. & Knoers, N. V. A. M. Current insights into renal ciliopathies: what can genetics teach us? Pediatr. Nephrol. 28, 863–874 (2013).
Schueler, M. et al. Large-scale targeted sequencing comparison highlights extreme genetic heterogeneity in nephronophthisis-related ciliopathies. J. Med. Genet. 53, 208–214 (2016).
Kubiak, M. & Lewandowska, M. A. Can chromatin conformation technologies bring light into human molecular pathology? Acta Biochim. Pol. 62, 483–489 (2015).
Mimura, I., Kanki, Y., Kodama, T. & Nangaku, M. Revolution of nephrology research by deep sequencing: ChIP-seq and RNA-seq. Kidney Int. 85, 31–38 (2014).
Elumalai, R., Periasamy, S., Ramanathan, G., Lakkakula, B. V. & Soundararajan, P. Journal of renal injury prevention role of endothelial nitric oxide synthase VNTR (intron 4 a/b) polymorphism on the progression of renal disease in autosomal dominant polycystic kidney disease. J. Renal Inj. Prev. 3, 69–73 (2014).
Merta, M., Reiterová, J., Tesar, V., Štekrová, J. & Viklický, O. Influence of the endothelial nitric oxide synthase polymorphism on the progression of autosomal dominant polycystic kidney disease and IgA nephropathy. Ren. Fail. 24, 585–593 (2002).
King, K., Flinter, F. A., Nihalani, V. & Green, P. M. Unusual deep intronic mutations in the COL4A5 gene cause X linked Alport syndrome. Hum. Genet. 111, 548–554 (2002).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
van de Hoek, G. et al. Functional models for congenital anomalies of the kidney and urinary tract. Nephron 129, 62–67 (2015).
Yao, X.-D. et al. Challenge in pathologic diagnosis of Alport syndrome: evidence from correction of previous misdiagnosis. Orphanet J. Rare Dis. 7, 100 (2012).
Jais, J. P. et al. X-linked Alport syndrome: natural history in 195 families and genotype–phenotype correlations in males. J. Am. Soc. Nephrol. 11, 649–657 (2000).
Stenson, P. D. et al. The Human Gene Mutation Database: building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Hum. Genet. 133, 1–9 (2014).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2011).
Perrichot, R. et al. DGGE screening of PKD1 gene reveals novel mutations in a large cohort of 146 unrelated patients. Hum. Genet. 105, 231–239 (1999).
Harris, P. C. & Torres, V. E. in GeneReviews (eds Pagon, R. A. et al.) 1–46 (Univ. of Washington, 2015).
Biesecker, L. G. et al. The ClinSeq Project: piloting large-scale genome sequencing for research in genomic medicine. Genome Res. 19, 1665–1674 (2009).
Nicolaou, N. et al. Prioritization and burden analysis of rare variants in 208 candidate genes suggest they do not play a major role in CAKUT. Kidney Int. 89, 476–486 (2016).
Roversi, G. et al. Constitutional de novo deletion of the FBXW7 gene in a patient with focal segmental glomerulosclerosis and multiple primitive tumors. Sci. Rep. 5, 15454 (2015).
Chaki, M. et al. Genotype–phenotype correlation in 440 patients with NPHP-related ciliopathies. Kidney Int. 80, 1239–1245 (2011).
Firth, H. V. et al. DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans Using Ensembl Resources. Am. J. Hum. Genet. 84, 524–533 (2009).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, 862–868 (2016).
Nicolaou, N., Renkema, K. Y., Bongers, E. M. H. F., Giles, R. H. & Knoers, N. V. A. M. Genetic, environmental, and epigenetic factors involved in CAKUT. Nat. Rev. Nephrol. 11, 720–731 (2015).
Prakash, S. & Gharavi, A. G. Diagnosing kidney disease in the genetic era. Curr. Opin. Nephrol. Hypertens. 24, 380–387 (2015).
Liebau, M. C. An emerging molecular understanding and novel targeted treatment approaches in pediatric kidney diseases. Front. Pediatr. 2, 68 (2014).
Weber, S. & Tonshoff, B. Recurrence of focal-segmental glomerulosclerosis in children after renal transplantation: clinical and genetic aspects. Transplantation 80, S128–S134 (2005).
Noris, M., Bresin, E., Mele, C. & Remuzzi, G. in GeneReviews 1–28 (2013).
Gross, O. et al. Early angiotensin-converting enzyme inhibition in Alport syndrome delays renal failure and improves life expectancy. Kidney Int. 81, 494–501 (2012).
Webb, N. J. et al. Losartan and enalapril are comparable in reducing proteinuria in children with Alport syndrome. Pediatr. Nephrol. 28, 737–743 (2013).
Slaats, G. G., Lilien, M. R. & Giles, R. H. Nephronophthisis: should we target cysts or fibrosis? Pediatr. Nephrol. 31, 545–554 (2016).
Lemmink, H. H. et al. Benign familial hematuria due to mutation of the type IV collagen A4 gene. J. Clin. Invest. 98, 1114–1118 (1996).
Badenas, C. et al. Mutations in the COL4A4 and COL4A3 genes cause familial benign hematuria. J. Am. Soc. Nephrol. 13, 1248–1254 (2002).
Mochizuki, T. et al. Identification of mutations in the α3(IV) and α4(IV) collagen genes in autosomal recessive Alport syndrome. Nat. Genet. 8, 77–82 (1994).
Barker, D. et al. Identification of mutations in the COL4A5 collagen gene in Alport syndrome. Science 248, 1224–1227 (1990).
Sanyanusin, P. et al. Mutation of the PAX2 gene in a family with optic nerve colobomas, renal anomalies and vesicoureteral reflux. Nat. Genet. 9, 358–364 (1995).
Nishimoto, K. et al. PAX2 gene mutation in a family with isolated renal hypoplasia. J. Am. Soc. Nephrol. 12, 1769–1772 (2001).
Barua, M. et al. Mutations in PAX2 associate with adult-onset FSGS. J. Am. Soc. Nephrol. 25, 1942–1953 (2014).
Horikawa, Y. et al. Mutation in hepatocyte nuclear factor-1β gene (TCF2) associated with MODY. Nat. Genet. 15, 57–61 (1997).
Weber, S. et al. Prevalence of mutations in renal developmental genes in children with renal hypodysplasia: results of the ESCAPE study. J. Am. Soc. Nephrol. 17, 2864–2870 (2006).
Hwang, D.-Y. et al. Mutations in 12 known dominant disease-causing genes clarify many congenital anomalies of the kidney and urinary tract. Kidney Int. 85, 1–5 (2014).
Thomas, R. et al. HNF1B and PAX2 mutations are a common cause of renal hypodysplasia in the CKiD cohort. Pediatr. Nephrol. 26, 897–903 (2011).
Hiesberger, H. H. et al. Mutation of hepatocyte nuclear factor-1B inhibits Pkhd1 gene expression and produces renal cysts in mice. J. Clin. Invest. 113, 814–825 (2004).
Gresh, L. et al. A transcriptional network in polycystic kidney disease. EMBO 23, 1657–1668 (2004).
Heidet, L. et al. Spectrum of HNF1B mutations in a large cohort of patients who harbor renal diseases. Clin. J. Am. Soc. Nephrol. 5, 1079–1090 (2010).
Kyttälä, M. et al. MKS1, encoding a component of the flagellar apparatus basal body proteome, is mutated in Meckel syndrome. Nat. Genet. 38, 155–157 (2006).
Hopp, K. et al. B9d1 is revealed as a novel Meckel syndrome (MKS) gene by targeted exon-enriched next-generation sequencing and deletion analysis. Hum. Mol. Genet. 20, 2524–2534 (2011).
Sayer, J. A. et al. The centrosomal protein nephrocystin-6 is mutated in Joubert syndrome and activates transcription factor ATF4. Nat. Genet. 38, 674–681 (2006).
Baala, L. et al. Pleiotropic effects of CEP290 (NPHP6) mutations extend to Meckel syndrome. Am. J. Hum. Genet. 81, 170–179 (2007).
Haber, D. A. et al. An internal deletion within an 11p13 zinc finger gene contributes to the development of Wilms' tumor. Cell 61, 1257–1269 (1990).
Hastie, N. Dominant negative mutations in the Wilms tumour (WT1) gene cause Denys-Drash syndrome — proof that a tumour-suppressor gene plays a crucial role in normal genitourinary development. Hum. Molec. Genet. 1, 293–295 (1992).
Jeanpierre, C. et al. Identification of constitutional WT1 mutations, in patients with isolated diffuse mesangial sclerosis, and analysis of genotype/phenotype correlations by use of a computerized mutation database. Am. J. Hum. Genet. 62, 824–833 (1998).
Yamagata, K. et al. Mutations in the hepatocyte nuclear factor-4α gene in maturity-onset diabetes of the young (MODY1). Nature 384, 458–460 (1996).
Dreyer, S. D. et al. Mutations in LMX1B cause abnormal skeletal patterning and renal dysplasia in nail patella syndrome. Nat. Genet. 19, 47–50 (1998).
Bongers, E. M. et al. Genotype-phenotype studies in nail-patella syndrome show that LMX1B mutation location is involved in the risk of developing nephropathy. Eur. J. Hum. Genet. 13, 935–946 (2005).
Bailey, L. J. et al. Characterization of a candidate gene for OCRL. Am. J. Hum. Genet. 51, 1 (1992).
Shrimpton, A. E. et al. OCRL1 mutations in dent 2 patients suggest a mechanism for phenotypic variability. Nephron. Physiol. 112, 27–36 (2009).
Hichri, H. et al. From lowe syndrome to Dent disease: Correlations between mutations of the OCRL1 gene and clinical and biochemical phenotypes. Hum. Mutat. 32, 379–388 (2011).
Mehta, Z. B., Pietka, G. & Lowe, M. The cellular and physiological functions of the lowe syndrome protein OCRL1. Traffic 15, 471–487 (2014).
McGregor, L. et al. Fraser syndrome and mouse blebbed phenotype caused by mutations in FRAS1/Fras1 encoding a putative extracellular matrix protein. Nat. Genet. 34, 203–208 (2003).
Jadeja, S. et al. Identification of a new gene mutated in Fraser syndrome and mouse myelencephalic blebs. Nat. Genet. 37, 520–525 (2005).
Vogel, M. J. et al. Mutations in GRIP1 cause Fraser syndrome. J. Med. Genet. 49, 303–306 (2012).
Jais, J. P. et al. X-linked Alport syndrome: natural history and genotype-phenotype correlations in girls and women belonging to 195 families: a 'European Community Alport Syndrome Concerted Action' Study. J. Am. Soc. Nephrol. 14, 2603–2610 (2003).
Acknowledgements
The researchers receive funding from the Dutch Kidney Foundation under grant agreements CP11.18 Kouncil (N.V.A.M.K. and R.H.G.) and KSTP12 010 (A.M.v.E.), the European Community's Seventh Framework Programme (FP7/2009) under grant agreement 305608 EURenOmics (N.V.A.M.K., F.S. and K.Y.R.) and Fonds NutsOhra grant 1303–070 (A.M.v.E.).
Author information
Authors and Affiliations
Contributions
M.F.S. researched data for the article and wrote the article. All authors made substantial contributions to discussions of the article's content and reviewed/edited the manuscript before submission
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Next-generation sequencing
-
A technique that enables the simultaneous investigation of mulitple genes and pathways in parallel. The term includes all forms of modern, high-throughput sequencing techniques, including gene panel sequencing, whole-exome sequencing and whole-genome sequencing.
- Nephrogenetic diseases
-
Kidney diseases with a genetic aetiology, including hereditary kidney disorders for which the responsible genes have not yet been identified.
- Pseudogenes
-
DNA sequences that are similar to genes but do not encode functional proteins.
- Gene panel sequencing
-
Targeted sequencing of a set of genes that are associated with a specific phenotype.
- Whole-exome sequencing
-
Targeted sequencing of all the protein-coding regions (1–2%) of the genome.
- Whole-genome sequencing
-
Untargeted sequencing of the complete genome.
- Causal
-
Variant(s) that are the cause of a specific phenotype.
- Missense
-
A variant that results in a single amino-acid substitution.
- Truncating
-
A variant that introduces a premature stop codon and results in a shortened protein.
- Biallelic
-
Variant present on both alleles of a specific gene. Biallelic variants can be homozygous or compound heterozygous.
- Probands
-
Patients who are the starting points of genetic studies in families.
- Pathogenic
-
Variant that has an effect on protein function that is associated with a specific disease phenotype.
- Hypomorphic
-
A mutation that results in reduced expression or reduced activity of a protein. The resulting disease phenotype is potentially milder than when mutations cause a complete loss of functional protein.
- Phenotypic heterogeneity
-
When mutations in the same gene can give rise to two or more distinct clinical phenotypes.
- Oligogenic inheritance
-
Inheritance model in which a phenotype is determined by the combination of a few genes.
- Chromosome conformation capture
-
A technique used to study the in vivo organization and interactions of genomic elements.
- Chromatin immunoprecipitation sequencing
-
A technique that combines chromatin immunoprecipitation with NGS to study the in vivo interaction between proteins (for example transcription factors) or epigenetic modifications (for example histone modifications) and the DNA.
- Incidental findings
-
Findings that are unrelated to the condition for which the DNA test is performed, including alleles that confer disease-risk to the patient as well as carriership for recessive or X-linked disease.
- Variants of unknown significance
-
Variants for which the association with disease risk is unknown.
- Copy number variation
-
A type of structural variation that alters the diploid status of DNA (deletions and duplications).
- Loss-of-function
-
Variant that results in a protein with reduced or no function.
- Segregation analysis
-
Study of the association between a genetic variant and a specific phenotype in a family. It is used to establish the mode of inheritance and to investigate if a specific genetic variant could potentially be causal for the disease in a family.
- Canonical
-
A canonical disease gene is a gene that is typically associated with a particular disease.
- Replication stress
-
Occurs when replication fork progression is slow or problematic and can result in DNA damage and genomic instability.
Rights and permissions
About this article
Cite this article
Stokman, M., Renkema, K., Giles, R. et al. The expanding phenotypic spectra of kidney diseases: insights from genetic studies. Nat Rev Nephrol 12, 472–483 (2016). https://doi.org/10.1038/nrneph.2016.87
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrneph.2016.87
This article is cited by
-
Expanding the genotype–phenotype correlations in Alport syndrome: novel mutations, digenic inheritance, and genetic modifiers
Egyptian Journal of Medical Human Genetics (2023)
-
Organs-on-chip technology: a tool to tackle genetic kidney diseases
Pediatric Nephrology (2022)
-
The utility of a genetic kidney disease clinic employing a broad range of genomic testing platforms: experience of the Irish Kidney Gene Project
Journal of Nephrology (2022)
-
Genomic medicine for kidney disease
Nature Reviews Nephrology (2018)
-
Could the interaction between LMX1B and PAX2 influence the severity of renal symptoms?
European Journal of Human Genetics (2018)