Abstract
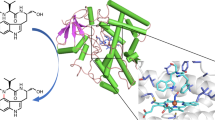
Cytochrome P450eryF catalyzes the 6S-hydroxylation of 6-deoxyerythronolide B, the initial reaction in a multistep pathway to convert 6-deoxyerythronolide B into the antibiotic, erythromycin. The overall structure of P450eryF is similar to that of P450cam but differs in the exact positioning of several α-helices. The largest difference occurs in the B′ helix and results in the enlargement of the substrate-binding pocket of P450eryF. The substrate is positioned with the macrolide ring perpendicular to the haem plane and contacts seven hydrophobic residues and three solvent molecules. The substrate participates in a network of hydrogen bonds that may provide a proton shuttle pathway in the oxygen cleavage reaction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Andersen, J.F. & Hutchinson, C.R. Characterization of Saccharopolyspora erythaea cytochrome P450 genes and enzymes, including 6-deoxyerythronolide B hydroxylase. J. Bacteriol. 174, 725–735 (1992).
Shafiee, A. & Hutchinson, C.R. Purification and reconstitution of the electron transport components of 6-deoxyerthronolide B hydroxylase, a cytochrome P450 enzyme macrolide antibiotic (erythromycin) biosynthesis. J. Bacteriol. 170, 1548–1553 (1988).
Zotchev, S.B. & Hutchinson, C.R. Cloning and heterologous expression of the genes encoding nonspecific electron transport components for a cytochrome P450 system of Saccharopolyspora erythraea involved in erythromycin production. Biochemistry (1994)
Raag, R. & Poulos, T.L. X-ray crystallographic studies of P450cam; factors controlling substrate metabolism in Frontiers in Biotransformations, (eds Ruckpaul, K. & Rein, H.) 1–43, (Akademia Verlag; 1992).
Poulos, T.L., Finzel, B.C. & Howard, A.J. Crystal structure of substrate-free Pseudomonas putida cytochrome P–450. Biochemistry 25, 5314–5322 (1986).
Poulos, T.L., Finzel, B.C. & Howard, A.J. High-resolution crystal structure of cytochrome P450cam. J. molec. Biol. 195, 687–700 (1987).
Nelson, D.R. & Stobel, H.W. On the membrane topology of vertebrate cytochrome P450 proteins. J. Biol. Chem. 263, 6038–6050 (1988).
Vijayakumar, S. & Salerno, J.C. Molecular modeling of the 3-D structure of cytochrome P450scc. Biochim. biophys. Acta 1160, 281–286 (1992).
Cupp-Vickery, J.R., Li, H. & Poulos, T.L. Preliminary crystallographic analysis of an enzyme involved in erythromycin biosynthesis: cytochrome P450eryF. Proteins (1994).
Nelson, D.R. & Strobel, H.W. Secondary structure prediction of 52 membrane-bound cytochromes P-450 shows a stron structural similarity to P450cam. Biochemistry 28, 656–660 (1989).
Ravichandran, K.G., Boodupalli, S.S., Hasemann, C.A., Peterson, J.A., Deisenhofer, J. Crystal structure of hemoprotein domain of P450BM-3′ a prototype for microsomal P-450s. Science 261, 731–736 (1993).
Li, H. & Poulos, T.L. Modeling protein-substrate interactions in the haem domain of cytochrome P450BM-3. Acta crystallogr. D in the press (1994).
Haseman, C.A., Ravichandran, K.G., Peterson, J.A. & Deisenhofer, J. Crystal structure and refinement of cytochrome P450terp at 2.3 resolution. J. molec. Biol. 236, 1169–1185 (1994).
Evans, S.V. SETOR: Hardware-lighted three-dimensional solid model representations of macromolecules. J. molec. Graphics. 11, 134–138 (1993).
Raag, R. & Poulos, T.L. Crystal structures of cytochrorme P450cam complexed with camphane, thiocamphor, and adamantane: factors controlling P450 substrate hydroxylation. Biochemistry 30, 2674–2684 (1991).
Li, H. & Poulos, T.L. Structural variation in haem enzymes: a comparative analysis of peroxidase and P450 crystal structures. Structure 2, 461–464 (1994).
Poulos, T.L., Edwards, S.L., Wariishi, H. & Gold, M.H. Crystallographic refinement of lignin peroxidase at 2 Å. Biol. Chem. 268, 4429–4440 (1993).
Quiocho, F.A., Wilson, D.K., Wilson, D.K. & Vyas, N.K. Substrate specificity and affinity of a protein modulated by bound water molecules. Nature 340, 404–407 (1989).
Andersen, J.F., Tatsuta, K., Gunji, H., Ishiyama, T. & Hutchinson, C.R. Substrate specificity of 6-deoxyerythronlide B hydroxylase, a bacterial cytochrome P450 of erythromycin A biosynthesis. Biochemistry 32, 1905–1913 (1993).
Matinis, S.A., Atkins, W.M., Stayton, P.S. & Sligar, S.G. A conserved residue of P450 involved in haem-oxygen stability and activation. J. Am. chem. Soc. 111, 9252–9253 (1989).
Imai, M. et al. Uncoupling the cytochrome P450cam monooxygenase reaction by a single mutation, threonine-252 to alanine or valine: a possible role of hydroxy amino acid in oxygen activation. Proc. natn. Acad. Sci. U.S.A. 86, 7823–7827 (1989).
Raag, R., Martinis, S.A., Sligar, S.G. & Poulos, T.L. Crystal structure of the cytochrome P450cam acive site mutant Thr252Ala. Biochemistry 20, 11420–11429 (1991).
SAXII Area Detector Software Release Notes for Version 1.21 (Siemes Analytical X-ray Instruments, Inc.).
Howard, A.J., Gilliland, G.L., Finzel, B.C., Poulos, T.L., Ohlendorf, D.H. & Salemme, F.R. The use of an imaging proportional counter in macromolecular crystallography. J. appl. Crystallogr. 20, 383–387 (1987).
Furey, W. Am. crystallogr. Assoc., 2, 18–73 (1990).
Brunger, A.T. X-PLOR Version 3.1. A system for X-ray crystallography and NMR. (New Haven, Yale University Press; 1992).
Fitzgerald, P.M.D. A package of computer programs for the determination of phases using molecular replacement. J. appl. Crystallogr. 21, 275–278 (1988).
Read, R.J. Improved fourier coefficients for maps using phases from partial structures with errors. Acta crystallogr. A42, 140–149 (1986).
Evans, P.R. Crystallographic computing 5, (eds Moras, D. et al., 136–144. Oxford University Press, 1991).
Zhang, K.Y.J. & Maine, P. The use of Sayreis equation with solvent flattening and histogram matching for phase extension and refinement of protein structues. Acta crystallogr. A46, 377–381 (1990).
Cowtan, K.D. & Maine, P. Improvement of macromolecular electron density maps by the simultaneous application of real and reciprocal space constrains. Acta crystallogr. D49, 148–157 (1993).
Cambillau, C. & Harjales, E. TOM: a FRODO subpackage for protein-ligand fitting with interactive energy minimization. J. molec. Graphics 5, 174–177 (1987).
Flinta, C., Persson, B., Jornvall, H. & von Heijne, G. Sequence determinants of cytosolic N-terminal protein processing. E. J. Biochem. 154, 193–196 (1994).
Kraulis, P. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. appl. Crystallogr. 24, 946–950 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Cupp-Vickery, J., Poulos, T. Structure of cytochrome P450eryF involved in erythromycin biosynthesis. Nat Struct Mol Biol 2, 144–153 (1995). https://doi.org/10.1038/nsb0295-144
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0295-144
This article is cited by
-
Artificial control of the multistep oxidation reactions catalyzed by the cytochrome P450 enzyme RosC
Applied Microbiology and Biotechnology (2020)
-
Hydrogen Bonds with Chalcogens: Looking Beyond the Second Row of the Periodic Table
Journal of the Indian Institute of Science (2020)
-
The hydrogen bond donor and acceptor ability of thioformic acid
Structural Chemistry (2011)
-
Understanding substrate misrecognition of hydrogen peroxide dependent cytochrome P450 from Bacillus subtilis
JBIC Journal of Biological Inorganic Chemistry (2010)
-
Substrate binding to cytochromes P450
Analytical and Bioanalytical Chemistry (2008)