Abstract
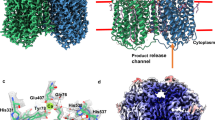
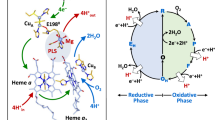
The structure of quinol-dependent nitric oxide reductase (qNOR) from G. stearothermophilus, which catalyzes the reduction of NO to produce the major ozone-depleting gas N2O, has been characterized at 2.5 Å resolution. The overall fold of qNOR is similar to that of cytochrome c–dependent NOR (cNOR), and some structural features that are characteristic of cNOR, such as the calcium binding site and hydrophilic cytochrome c domain, are observed in qNOR, even though it harbors no heme c. In contrast to cNOR, structure-based mutagenesis and molecular dynamics simulation studies of qNOR suggest that a water channel from the cytoplasm can serve as a proton transfer pathway for the catalytic reaction. Further structural comparison of qNOR with cNOR and aerobic and microaerobic respiratory oxidases elucidates their evolutionary relationship and possible functional conversions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Zumft, W.G. Cell biology and molecular basis of denitrification. Microbiol. Mol. Biol. Rev. 61, 533–616 (1997).
Zumft, W.G. Nitric oxide reductases of prokaryotes with emphasis on the respiratory, heme-copper oxidase type. J. Inorg. Biochem. 99, 194–215 (2005).
Watmough, N.J., Field, S.J., Hughes, R.J. & Richardson, D.J. The bacterial respiratory nitric oxide reductase. Biochem. Soc. Trans. 37, 392–399 (2009).
Hendriks, J. et al. Nitric oxide reductases in bacteria. Biochim. Biophys. Acta 1459, 266–273 (2000).
Stevanin, T.M., Moir, J.W. & Read, R.C. Nitric oxide detoxification systems enhance survival of Neisseria meningitidis in human macrophages and in nasopharyngeal mucosa. Infect. Immun. 73, 3322–3329 (2005).
Philippot, L. Denitrification in pathogenic bacteria: for better or worst? Trends Microbiol. 13, 191–192 (2005).
Ravishankara, A.R., Daniel, J.S. & Portmann, R.W. Nitrous oxide (N2O): the dominant ozone-depleting substance emitted in the 21st century. Science 326, 123–125 (2009).
Canfield, D.E., Glazer, A.N. & Falkowski, P.G. The evolution and future of Earth's nitrogen cycle. Science 330, 192–196 (2010).
Prather, M.J. & Hsu, J. Coupling of nitrous oxide and methane by global atmospheric chemistry. Science 330, 952–954 (2010).
Wuebbles, D.J. Atmosphere. Nitrous oxide: no laughing matter. Science 326, 56–57 (2009).
Kumita, H. et al. NO reduction by nitric-oxide reductase from denitrifying bacterium Pseudomonas aeruginosa: characterization of reaction intermediates that appear in the single turnover cycle. J. Biol. Chem. 279, 55247–55254 (2004).
Moënne-Loccoz, P. Spectroscopic characterization of heme iron-nitrosyl species and their role in NO reductase mechanisms in diiron proteins. Nat. Prod. Rep. 24, 610–620 (2007).
Yeung, N. et al. Rational design of a structural and functional nitric oxide reductase. Nature 462, 1079–1082 (2009).
Blomberg, L.M., Blomberg, M.R. & Siegbahn, P.E. Reduction of nitric oxide in bacterial nitric oxide reductase–a theoretical model study. Biochim. Biophys. Acta 1757, 240–252 (2006).
Saraste, M. & Castresana, J. Cytochrome oxidase evolved by tinkering with denitrification enzymes. FEBS Lett. 341, 1–4 (1994).
Fujiwara, T. & Fukumori, Y. Cytochrome cb-type nitric oxide reductase with cytochrome c oxidase activity from Paracoccus denitrificans ATCC 35512. J. Bacteriol. 178, 1866–1871 (1996).
Giuffrè, A. et al. The heme-copper oxidases of Thermus thermophilus catalyze the reduction of nitric oxide: evolutionary implications. Proc. Natl. Acad. Sci. USA 96, 14718–14723 (1999).
Huang, Y., Reimann, J., Lepp, H., Drici, N. & Adelroth, P. Vectorial proton transfer coupled to reduction of O2 and NO by a heme-copper oxidase. Proc. Natl. Acad. Sci. USA 105, 20257–20262 (2008).
Hayashi, T. et al. Accommodation of two diatomic molecules in cytochrome bo: insights into NO reductase activity in terminal oxidases. Biochemistry 48, 883–890 (2009).
Hendriks, J.H., Jasaitis, A., Saraste, M. & Verkhovsky, M.I. Proton and electron pathways in the bacterial nitric oxide reductase. Biochemistry 41, 2331–2340 (2002).
Reimann, J., Flock, U., Lepp, H., Honigmann, A. & Adelroth, P. A pathway for protons in nitric oxide reductase from Paracoccus denitrificans. Biochim. Biophys. Acta 1767, 362–373 (2007).
Hino, T. et al. Structural basis of biological N2O generation by bacterial nitric oxide reductase. Science 330, 1666–1670 (2010).
Iwata, S., Ostermeier, C., Ludwig, B. & Michel, H. Structure at 2.8 Å resolution of cytochrome c oxidase from Paracoccus denitrificans. Nature 376, 660–669 (1995).
Tsukihara, T. et al. The whole structure of the 13-subunit oxidized cytochrome c oxidase at 2.8 Å. Science 272, 1136–1144 (1996).
Soulimane, T. et al. Structure and mechanism of the aberrant ba3-cytochrome c oxidase from Thermus thermophilus. EMBO J. 19, 1766–1776 (2000).
Buschmann, S. et al. The structure of cbb3 cytochrome oxidase provides insights into proton pumping. Science 329, 327–330 (2010).
Flock, U., Reimann, J. & Adelroth, P. Proton transfer in bacterial nitric oxide reductase. Biochem. Soc. Trans. 34, 188–190 (2006).
Flock, U. et al. Defining the proton entry point in the bacterial respiratory nitric-oxide reductase. J. Biol. Chem. 283, 3839–3845 (2008).
Cramm, R., Pohlmann, A. & Friedrich, B. Purification and characterization of the single-component nitric oxide reductase from Ralstonia eutropha H16. FEBS Lett. 460, 6–10 (1999).
de Vries, S., Strampraad, M.J., Lu, S., Moenne-Loccoz, P. & Schroder, I. Purification and characterization of the MQH2:NO oxidoreductase from the hyperthermophilic archaeon Pyrobaculum aerophilum. J. Biol. Chem. 278, 35861–35868 (2003).
Castresana, J., Lubben, M., Saraste, M. & Higgins, D.G. Evolution of cytochrome oxidase, an enzyme older than atmospheric oxygen. EMBO J. 13, 2516–2525 (1994).
van der Oost, J. et al. Restoration of a lost metal-binding site: construction of two different copper sites into a subunit of the E. coli cytochrome o quinol oxidase complex. EMBO J. 11, 3209–3217 (1992).
Pereira, M.M., Santana, M. & Teixeira, M. A novel scenario for the evolution of haem-copper oxygen reductases. Biochim. Biophys. Acta 1505, 185–208 (2001).
Damaschun, G., Damaschun, H., Gast, K., Gernat, C. & Zirwer, D. Acid denatured apo-cytochrome c is a random coil: evidence from small-angle X-ray scattering and dynamic light scattering. Biochim. Biophys. Acta 1078, 289–295 (1991).
Butland, G., Spiro, S., Watmough, N.J. & Richardson, D.J. Two conserved glutamates in the bacterial nitric oxide reductase are essential for activity but not assembly of the enzyme. J. Bacteriol. 183, 189–199 (2001).
Thorndycroft, F.H., Butland, G., Richardson, D.J. & Watmough, N.J. A new assay for nitric oxide reductase reveals two conserved glutamate residues form the entrance to a proton-conducting channel in the bacterial enzyme. Biochem. J. 401, 111–119 (2007).
Lin, Y.W. et al. Roles of glutamates and metal ions in a rationally designed nitric oxide reductase based on myoglobin. Proc. Natl. Acad. Sci. USA 107, 8581–8586 (2010).
Olkhova, E., Hutter, M.C., Lill, M.A., Helms, V. & Michel, H. Dynamic water networks in cytochrome c oxidase from Paracoccus denitrificans investigated by molecular dynamics simulations. Biophys. J. 86, 1873–1889 (2004).
Rodrigues, M.L., Scott, K.A., Sansom, M.S., Pereira, I.A. & Archer, M. Quinol oxidation by c-type cytochromes: structural characterization of the menaquinol binding site of NrfHA. J. Mol. Biol. 381, 341–350 (2008).
Abramson, J. et al. The structure of the ubiquinol oxidase from Escherichia coli and its ubiquinone binding site. Nat. Struct. Biol. 7, 910–917 (2000).
Jasaitis, A. et al. Electrogenic reactions of cytochrome bd. Biochemistry 39, 13800–13809 (2000).
Haltia, T. et al. Crystal structure of nitrous oxide reductase from Paracoccus denitrificans at 1.6 Å resolution. Biochem. J. 369, 77–88 (2003).
Paraskevopoulos, K., Antonyuk, S.V., Sawers, R.G., Eady, R.R. & Hasnain, S.S. Insight into catalysis of nitrous oxide reductase from high-resolution structures of resting and inhibitor-bound enzyme from Achromobacter cycloclastes. J. Mol. Biol. 362, 55–65 (2006).
Ducluzeau, A.L. et al. Was nitric oxide the first deep electron sink? Trends Biochem. Sci. 34, 9–15 (2009).
Kawabata, T. MATRAS: A program for protein 3D structure comparison. Nucleic Acids Res. 31, 3367–3369 (2003).
Hemp, J. et al. Comparative genomics and site-directed mutagenesis support the existence of only one input channel for protons in the C-family (cbb3 oxidase) of heme-copper oxygen reductases. Biochemistry 46, 9963–9972 (2007).
Ettwig, K.F. et al. Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464, 543–548 (2010).
Petrek, M. et al. CAVER: a new tool to explore routes from protein clefts, pockets and cavities. BMC Bioinformatics 7, 316 (2006).
Matsuura, Y., Takano, T. & Dickerson, R.E. Structure of cytochrome c551 from Pseudomonas aeruginosa refined at 1.6 Å resolution and comparison of the two redox forms. J. Mol. Biol. 156, 389–409 (1982).
Stelter, M. et al. A novel type of monoheme cytochrome c: biochemical and structural characterization at 1.23 Å resolution of Rhodothermus marinus cytochrome c. Biochemistry 47, 11953–11963 (2008).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D Biol. Crystallogr. 59, 2023–2030 (2003).
Abrahams, J.P. & Leslie, A.G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D Biol. Crystallogr. 52, 30–42 (1996).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Brunger, A.T. Version 1.2 of the crystallography and NMR system. Nat. Protoc. 2, 2728–2733 (2007).
Larkin, M.A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
MacKerell, A.D. Jr. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Phillips, J.C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Andersen, H.C. Rattle: A “velocity” version of the shake algorithm for molecular dynamics calculations. J. Comput. Phys. 52, 24–34 (1983).
Humphrey, W., Dalke, A. & Schulten, K. VMD—Visual Molecular Dynamics. J. Mol. Graphics 14, 33–38 (1996).
Acknowledgements
We thank Y. Shimomura for support in the preparation of qNOR and the staff of the SPring-8 beamlines for their help with diffraction measurements. This work was supported by grants from the Ministry of Education, Culture, Sports, Science and Technology (21245041; T.T., H.S., S.N., A.V.P., Y. Sugita and Y. Shiro).
Author information
Authors and Affiliations
Contributions
Y.M. was responsible for cloning the qNOR gene and constructing the expression system as well as for purifying, characterizing and crystallizing the protein. T.T. carried out the enzyme assays and the ICP-AES metal analysis. T.H. purified P. aeruginosa cNOR. Y.M. and H.S. collected, processed and refined the X-ray data. A.V.P. carried out the molecular dynamics simulation. T.H., T.T., Y.M., S.N., Y. Sugita and Y. Shiro designed the study. Y.M., T.T., A.V.P., S.N. and Y. Shiro prepared the manuscript. All authors analyzed the data and discussed the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Methods (PDF 4397 kb)
Supplementary Movie 1
Dynamics in the water channel in MD simulation. Time-dependent dynamics, such as side chains fluctuations, moving water molecules, and transient hydrogen-bond networks can be seen. Water molecules in the water channel are shown in yellow, and bulk water around the channel entrance is shown as red and white lines. Hydrogen-bonds are shown as dashed green lines. (GIF 4495 kb)
Supplementary Movie 2
Motion of selected water molecules along the water channel in the MD simulation. Eight individual waters are shown as large spheres and are highlighted in different colors. Other water molecules inside the water channel and in the bulk around the channel entrance are shown as yellow sticks. The selected water molecules come into the water channel via the entry site, exchange with waters inside the channel, travel up to the binuclear active center, and eventually return to the bulk. The water molecules are highly mobile and move easily along the channel on a short timescale. There are several stable positions (“traps”) where water molecules stay near residues for a long time (typically, nanoseconds), before moving further along the channel. (GIF 4028 kb)
Rights and permissions
About this article
Cite this article
Matsumoto, Y., Tosha, T., Pisliakov, A. et al. Crystal structure of quinol-dependent nitric oxide reductase from Geobacillus stearothermophilus. Nat Struct Mol Biol 19, 238–245 (2012). https://doi.org/10.1038/nsmb.2213
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2213
This article is cited by
-
A 2.2 Å cryoEM structure of a quinol-dependent NO Reductase shows close similarity to respiratory oxidases
Nature Communications (2023)
-
Denitrifying Woodchip Bioreactors: A Microbial Solution for Nitrate in Agricultural Wastewater—A Review
Journal of Microbiology (2023)
-
Phylogenetic diversity of NO reductases, new tools for nor monitoring, and insights into N2O production in natural and engineered environments
Frontiers of Environmental Science & Engineering (2022)
-
Architecture of bacterial respiratory chains
Nature Reviews Microbiology (2021)
-
Biological nitrogen removal from wastewater using sulphur-driven autotrophic denitrification
Applied Microbiology and Biotechnology (2019)