Abstract
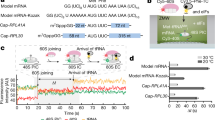
During translation elongation, the ribosome compositional factors elongation factor G (EF-G; encoded by fusA) and tRNA alternately bind to the ribosome to direct protein synthesis and regulate the conformation of the ribosome. Here, we use single-molecule fluorescence with zero-mode waveguides to directly correlate ribosome conformation and composition during multiple rounds of elongation at high factor concentrations in Escherichia coli. Our results show that EF-G bound to GTP (EF-G–GTP) continuously samples both rotational states of the ribosome, binding with higher affinity to the rotated state. Upon successful accommodation into the rotated ribosome, the EF-G–ribosome complex evolves through several rate-limiting conformational changes and the hydrolysis of GTP, which results in a transition back to the nonrotated state and in turn drives translocation and facilitates release of both EF-G–GDP and E-site tRNA. These experiments highlight the power of tracking single-molecule conformation and composition simultaneously in real time.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Blanchard, S.C., Gonzalez, R.L., Kim, H.D., Chu, S. & Puglisi, J.D. tRNA selection and kinetic proofreading in translation. Nat. Struct. Mol. Biol. 11, 1008–1014 (2004).
Blanchard, S.C., Kim, H.D., Gonzalez, R.L. Jr., Puglisi, J.D. & Chu, S. tRNA dynamics on the ribosome during translation. Proc. Natl. Acad. Sci. USA 101, 12893–12898 (2004).
Rodnina, M.V., Savelsbergh, A., Katunin, V.I. & Wintermeyer, W. Hydrolysis of GTP by elongation factor G drives tRNA movement on the ribosome. Nature 385, 37–41 (1997).
Valle, M. et al. Locking and unlocking of ribosomal motions. Cell 114, 123–134 (2003).
Spirin, A.S. A model of the functioning ribosome: locking and unlocking of the ribosome subparticles. Cold Spring Harb. Symp. Quant. Biol. 34, 197–207 (1969).
Frank, J. & Agrawal, R.K. A ratchet-like inter-subunit reorganization of the ribosome during translocation. Nature 406, 318–322 (2000).
Aitken, C.E. & Puglisi, J.D. Following the intersubunit conformation of the ribosome during translation in real time. Nat. Struct. Mol. Biol. 17, 793–800 (2010).
Agirrezabala, X. et al. Visualization of the hybrid state of tRNA binding promoted by spontaneous ratcheting of the ribosome. Mol. Cell 32, 190–197 (2008).
Munro, J.B., Altman, R.B., O'Connor, N. & Blanchard, S.C. Identification of two distinct hybrid state intermediates on the ribosome. Mol. Cell 25, 505–517 (2007).
Moazed, D. & Noller, H.F. Intermediate states in the movement of transfer RNA in the ribosome. Nature 342, 142–148 (1989).
Cornish, P.V., Ermolenko, D.N., Noller, H.F. & Ha, T. Spontaneous intersubunit rotation in single ribosomes. Mol. Cell 30, 578–588 (2008).
Fei, J., Kosuri, P., MacDougall, D.D. & Gonzalez, R.L. Jr. Coupling of ribosomal L1 stalk and tRNA dynamics during translation elongation. Mol. Cell 30, 348–359 (2008).
Fei, J. et al. Allosteric collaboration between elongation factor G and the ribosomal L1 stalk directs tRNA movements during translation. Proc. Natl. Acad. Sci. USA 106, 15702–15707 (2009).
Stark, H., Rodnina, M.V., Wieden, H.J., van Heel, M. & Wintermeyer, W. Large-scale movement of elongation factor G and extensive conformational change of the ribosome during translocation. Cell 100, 301–309 (2000).
Agrawal, R.K., Heagle, A.B., Penczek, P., Grassucci, R.A. & Frank, J. EF-G-dependent GTP hydrolysis induces translocation accompanied by large conformational changes in the 70S ribosome. Nat. Struct. Biol. 6, 643–647 (1999).
Wilden, B., Savelsbergh, A., Rodnina, M.V. & Wintermeyer, W. Role and timing of GTP binding and hydrolysis during EF-G-dependent tRNA translocation on the ribosome. Proc. Natl. Acad. Sci. USA 103, 13670–13675 (2006).
Peske, F., Savelsbergh, A., Katunin, V.I., Rodnina, M.V. & Wintermeyer, W. Conformational changes of the small ribosomal subunit during elongation factor G-dependent tRNA-mRNA translocation. J. Mol. Biol. 343, 1183–1194 (2004).
Hauryliuk, V. et al. The pretranslocation ribosome is targeted by GTP-bound EF-G in partially activated form. Proc. Natl. Acad. Sci. USA 105, 15678–15683 (2008).
Munro, J.B., Wasserman, M.R., Altman, R.B., Wang, L. & Blanchard, S.C. Correlated conformational events in EF-G and the ribosome regulate translocation. Nat. Struct. Mol. Biol. 17, 1470–1477 (2010).
Marshall, R.A., Dorywalska, M. & Puglisi, J.D. Irreversible chemical steps control intersubunit dynamics during translation. Proc. Natl. Acad. Sci. USA 105, 15364–15369 (2008).
Dorywalska, M. et al. Site-specific labeling of the ribosome for single-molecule spectroscopy. Nucleic Acids Res. 33, 182–189 (2005).
Gao, Y.G. et al. The structure of the ribosome with elongation factor G trapped in the posttranslocational state. Science 326, 694–699 (2009).
Chen, J., Tsai, A., Petrov, A. & Puglisi, J.D. Nonfluorescent quenchers to correlate single-molecule conformational and compositional dynamics. J. Am. Chem. Soc. 134, 5734–5737 (2012).
Tsai, A. et al. Heterogeneous pathways and timing of factor departure during translation initiation. Nature 487, 390–393 (2012).
Uemura, S. et al. Real-time tRNA transit on single translating ribosomes at codon resolution. Nature 464, 1012–1017 (2010).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Marshall, R.A., Aitken, C.E. & Puglisi, J.D. GTP hydrolysis by IF2 guides progression of the ribosome into elongation. Mol. Cell 35, 37–47 (2009).
Horan, L.H. & Noller, H.F. Intersubunit movement is required for ribosomal translocation. Proc. Natl. Acad. Sci. USA 104, 4881–4885 (2007).
Ratje, A.H. et al. Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites. Nature 468, 713–716 (2010).
Semenkov, Y.P., Rodnina, M.V. & Wintermeyer, W. The “allosteric three-site model” of elongation cannot be confirmed in a well-defined ribosome system from Escherichia coli. Proc. Natl. Acad. Sci. USA 93, 12183–12188 (1996).
Chen, C. et al. Allosteric vs. spontaneous exit-site (E-site) tRNA dissociation early in protein synthesis. Proc. Natl. Acad. Sci. USA 108, 16980–16985 (2011).
Wen, J.D. et al. Following translation by single ribosomes one codon at a time. Nature 452, 598–603 (2008).
Lee, T.H., Blanchard, S.C., Kim, H.D., Puglisi, J.D. & Chu, S. The role of fluctuations in tRNA selection by the ribosome. Proc. Natl. Acad. Sci. USA 104, 13661–13665 (2007).
Ermolenko, D.N. & Noller, H.F. mRNA translocation occurs during the second step of ribosomal intersubunit rotation. Nat. Struct. Mol. Biol. 18, 457–462 (2011).
Spiegel, P.C., Ermolenko, D.N. & Noller, H.F. Elongation factor G stabilizes the hybrid-state conformation of the 70S ribosome. RNA 13, 1473–1482 (2007).
Savelsbergh, A. et al. An elongation factor G-induced ribosome rearrangement precedes tRNA-mRNA translocation. Mol. Cell 11, 1517–1523 (2003).
Munro, J.B., Altman, R.B., Tung, C.S., Sanbonmatsu, K.Y. & Blanchard, S.C. A fast dynamic mode of the EF-G-bound ribosome. EMBO J. 29, 770–781 (2010).
Zavialov, A.V. & Ehrenberg, M. Peptidyl-tRNA regulates the GTPase activity of translation factors. Cell 114, 113–122 (2003).
Wang, L. et al. Allosteric control of the ribosome by small-molecule antibiotics. Nat. Struct. Mol. Biol. 19, 957–963 (2012).
Richman, N. & Bodley, J.W. Ribosomes cannot interact simultaneously with elongation factors EF Tu and EF G. Proc. Natl. Acad. Sci. USA 69, 686–689 (1972).
Ermolenko, D.N. et al. The antibiotic viomycin traps the ribosome in an intermediate state of translocation. Nat. Struct. Mol. Biol. 14, 493–497 (2007).
Stanley, R.E., Blaha, G., Grodzicki, R.L., Strickler, M.D. & Steitz, T.A. The structures of the anti-tuberculosis antibiotics viomycin and capreomycin bound to the 70S ribosome. Nat. Struct. Mol. Biol. 17, 289–293 (2010).
Chen, C. et al. Single-molecule fluorescence measurements of ribosomal translocation dynamics. Mol. Cell 42, 367–377 (2011).
Zhang, W., Dunkle, J.A. & Cate, J.H. Structures of the ribosome in intermediate states of ratcheting. Science 325, 1014–1017 (2009).
Guo, Z. & Noller, H.F. Rotation of the head of the 30S ribosomal subunit during mRNA translocation. Proc. Natl. Acad. Sci. USA 109, 20391–20394 (2012).
Fischer, N., Konevega, A.L., Wintermeyer, W., Rodnina, M.V. & Stark, H. Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy. Nature 466, 329–333 (2010).
Chen, J., Tsai, A., O'Leary, S.E., Petrov, A. & Puglisi, J.D. Unraveling the dynamics of ribosome translocation. Curr. Opin. Struct. Biol. 22, 804–814 (2012).
Li, W., Trabuco, L.G., Schulten, K. & Frank, J. Molecular dynamics of EF-G during translocation. Proteins 79, 1478–1486 (2011).
Frank, J., Gao, H., Sengupta, J., Gao, N. & Taylor, D.J. The process of mRNA-tRNA translocation. Proc. Natl. Acad. Sci. USA 104, 19671–19678 (2007).
Peske, F., Matassova, N.B., Savelsbergh, A., Rodnina, M.V. & Wintermeyer, W. Conformationally restricted elongation factor G retains GTPase activity but is inactive in translocation on the ribosome. Mol. Cell 6, 501–505 (2000).
Tsai, A. et al. The impact of aminoglycosides on the dynamics of translation elongation. Cell Rep. 3, 497–508 (2013).
Acknowledgements
We would like to thank J. Korlach (Pacific Biosciences) for providing technical support on the ZMW instrumentation. This work was supported by US National Institutes of Health grants GM51266 (J.C., A.T. and J.D.P.) and GM099687 (A.P., S.E.O. and J.D.P.) and by a Stanford Interdisciplinary Graduate Fellowship (J.C.).
Author information
Authors and Affiliations
Contributions
J.C. performed all experiments and data analysis. J.C. and J.D.P. designed the project and wrote the manuscript. J.C., A.P., A.T., S.E.O. and J.D.P. discussed results. S.E.O. assisted with protein purification and labeling.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 2063 kb)
Rights and permissions
About this article
Cite this article
Chen, J., Petrov, A., Tsai, A. et al. Coordinated conformational and compositional dynamics drive ribosome translocation. Nat Struct Mol Biol 20, 718–727 (2013). https://doi.org/10.1038/nsmb.2567
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2567
This article is cited by
-
A model for ribosome translocation based on the alternated displacement of its subunits
European Biophysics Journal (2023)
-
Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP
Nature Communications (2021)
-
2′-O-methylation in mRNA disrupts tRNA decoding during translation elongation
Nature Structural & Molecular Biology (2018)
-
tRNA tracking for direct measurements of protein synthesis kinetics in live cells
Nature Chemical Biology (2018)
-
Real-time assembly of ribonucleoprotein complexes on nascent RNA transcripts
Nature Communications (2018)