Abstract
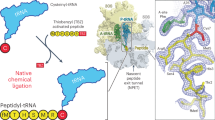
Ribosomes catalyze the formation of peptide bonds between aminoacyl esters of transfer RNAs within a catalytic center composed of ribosomal RNA only. Here we show that the reaction of P-site formylmethionine (fMet)-tRNAfMet with a modified A-site tRNA substrate, Phelac-tRNAPhe, in which the nucleophilic amino group is replaced with a hydroxyl group, does not show the pH dependence observed with small substrate analogs such as puromycin and hydroxypuromycin. This indicates that acid-base catalysis by ribosomal residues is not important in the reaction with the full-size substrate. Rather, the ribosome catalyzes peptide bond formation by positioning the tRNAs, or their 3′ termini, through interactions with rRNA that induce and/or stabilize a pH-insensitive conformation of the active site and provide a preorganized environment facilitating the reaction. The rate of peptide bond formation with unmodified Phe-tRNAPhe is estimated to be >300 s−1.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Noller, H.F., Hoffarth, V. & Zimniak, L. Unusual resistance of peptidyl transferase to protein extraction procedures. Science 256, 1416–1419 (1992).
Ban, N., Nissen, P., Hansen, J., Moore, P.B. & Steitz, T.A. The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution. Science 289, 905–920 (2000).
Nissen, P., Hansen, J., Ban, N., Moore, P.B. & Steitz, T.A. The structural basis of ribosome activity in peptide bond synthesis. Science 289, 920–930 (2000).
Harms, J. et al. High resolution structure of the large ribosomal subunit from a mesophilic eubacterium. Cell 107, 679–688 (2001).
Satterthwait, A.C. & Jencks, W.P. The mechanism of the aminolysis of acetate esters. J. Am. Chem. Soc. 96, 7018–7031 (1974).
Bevilacqua, P.C., Brown, T.S., Nakano, S. & Yajima, R. Catalytic roles for proton transfer and protonation in ribozymes. Biopolymers 73, 90–109 (2004).
Muth, G.W., Chen, L., Kosek, A.B. & Strobel, S.A. pH-dependent conformational flexibility within the ribosomal peptidyl transferase center. RNA 7, 1403–1415 (2001).
Krayevsky, A.A. & Kukhanova, M.K. The peptidyltransferase center of ribosomes. Prog. Nucleic Acid Res. Mol. Biol. 23, 1–51 (1979).
Nierhaus, K.H., Schulze, H. & Cooperman, B.S. Molecular mechanisms of the ribosomal peptidyltransferase center. Biochem. Int. 1, 185–192 (1980).
Schmeing, T.M., Huang, K.S., Kitchen, D.E., Strobel, S.A. & Steitz, T.A. Structural insights into the roles of water and the 2′ hydroxyl of the P site tRNA in the peptidyl transferase reaction. Mol. Cell 20, 437–448 (2005).
Sharma, P.K., Xiang, Y., Kato, M. & Warshel, A. What are the roles of substrate-assisted catalysis and proximity effects in peptide bond formation by the ribosome? Biochemistry 44, 11307–11314 (2005).
Trobro, S. & Aqvist, J. Mechanism of peptide bond synthesis on the ribosome. Proc. Natl. Acad. Sci. USA 102, 12395–12400 (2005).
Seila, A.C., Okuda, K., Nunez, S., Seila, A.F. & Strobel, S.A. Kinetic isotope effect analysis of the ribosomal peptidyl transferase reaction. Biochemistry 44, 4018–4027 (2005).
Sievers, A., Beringer, M., Rodnina, M.V. & Wolfenden, R. The ribosome as an entropy trap. Proc. Natl. Acad. Sci. USA 101, 7897–7901 (2004).
Hansen, J.L., Schmeing, T.M., Moore, P.B. & Steitz, T.A. Structural insights into peptide bond formation. Proc. Natl. Acad. Sci. USA 99, 11670–11675 (2002).
Yusupov, M.M. et al. Crystal structure of the ribosome at 5.5 Å resolution. Science 292, 883–896 (2001).
Bashan, A. et al. Structural basis of the ribosomal machinery for peptide bond formation, translocation, and nascent chain progression. Mol. Cell 11, 91–102 (2003).
Kim, D.F. & Green, R. Base-pairing between 23S rRNA and tRNA in the ribosomal A site. Mol. Cell 4, 859–864 (1999).
Pape, T., Wintermeyer, W. & Rodnina, M.V. Complete kinetic mechanism of elongation factor Tu-dependent binding of aminoacyl-tRNA to the A site of the E.coli ribosome. EMBO J. 17, 7490–7497 (1998).
Katunin, V.I., Muth, G.W., Strobel, S.A., Wintermeyer, W. & Rodnina, M.V. Important contribution to catalysis of peptide bond formation by a single ionizing group within the ribosome. Mol. Cell 10, 339–346 (2002).
Youngman, E.M., Brunelle, J.L., Kochaniak, A.B. & Green, R. The active site of the ribosome is composed of two layers of conserved nucleotides with distinct roles in peptide bond formation and peptide release. Cell 117, 589–599 (2004).
Wolfenden, R. The mechanism of hydrolysis of amino acyl RNA. Biochemistry 338, 1090–1092 (1963).
Fahnestock, S. & Rich, A. Synthesis by ribosomes of viral coat protein containing ester linkages. Nat. New Biol. 229, 8–10 (1971).
Fahnestock, S., Neumann, H., Shashoua, V. & Rich, A. Ribosome-catalyzed ester formation. Biochemistry 9, 2477–2483 (1970).
Schmeing, T.M., Huang, K.S., Strobel, S.A. & Steitz, T.A. An induced-fit mechanism to promote peptide bond formation and exclude hydrolysis of peptidyl-tRNA. Nature 438, 520–524 (2005).
Derwenskus, K.H. & Sprinzl, M. Interaction of cinnamyl-tRNAPhe with Escherichia coli elongation factor Tu. FEBS Lett. 151, 143–147 (1983).
Moazed, D. & Noller, H.F. Intermediate states in the movement of transfer RNA in the ribosome. Nature 342, 142–148 (1989).
Bevilacqua, P.C. Mechanistic considerations for general acid-base catalysis by RNA: revisiting the mechanism of the hairpin ribozyme. Biochemistry 42, 2259–2265 (2003).
Fedor, M.J. & Williamson, J.R. The catalytic diversity of RNAs. Nat. Rev. Mol. Cell Biol. 6, 399–412 (2005).
Okuda, K., Seila, A.C. & Strobel, S.A. Uncovering the enzymatic pKa of the ribosomal peptidyl transferase reaction utilizing a fluorinated puromycin derivative. Biochemistry 44, 6675–6684 (2005).
Muth, G.W., Ortoleva-Donnelly, L. & Strobel, S.A. A single adenosine with a neutral pKa in the ribosomal peptidyl transferase center. Science 289, 947–950 (2000).
Xiong, L., Polacek, N., Sander, P., Bottger, E.C. & Mankin, A. pKa of adenine 2451 in the ribosomal peptidyl transferase center remains elusive. RNA 7, 1365–1369 (2001).
Bayfield, M.A., Dahlberg, A.E., Schulmeister, U., Dorner, S. & Barta, A. A conformational change in the ribosomal peptidyl transferase center upon active/inactive transition. Proc. Natl. Acad. Sci. USA 98, 10096–10101 (2001).
Brunelle, J.L., Youngman, E.M., Sharma, D. & Green, R. The interaction between C75 of tRNA and the A loop of the ribosome stimulates peptidyl transferase activity. RNA 12, 33–39 (2006).
Hesslein, A.E. et al. Exploration of the conserved A+C wobble pair within the ribosomal peptidyl transferase center using affinity purified mutant ribosomes. Nucleic Acids Res. 32, 3760–3770 (2004).
Beringer, M. et al. Essential mechanisms in the catalysis of peptide bond formation on the ribosome. J. Biol. Chem. 280, 36065–36072 (2005).
Weinger, J.S., Parnell, K.M., Dorner, S., Green, R. & Strobel, S.A. Substrate-assisted catalysis of peptide bond formation by the ribosome. Nat. Struct. Mol. Biol. 11, 1101–1106 (2004).
Das, G.K., Bhattacharyya, D. & Burma, D.P. A possible mechanism of peptide bond formation on ribosome without mediation of peptidyl transferase. J. Theor. Biol. 200, 193–205 (1999).
Radzicka, A. & Wolfenden, R. A proficient enzyme. Science 267, 90–93 (1995).
Rodnina, M.V. & Wintermeyer, W. Fidelity of aminoacyl-tRNA selection on the ribosome: kinetic and structural mechanisms. Annu. Rev. Biochem. 70, 415–435 (2001).
Rodnina, M.V. & Wintermeyer, W. GTP consumption of elongation factor Tu during translation of heteropolymeric mRNAs. Proc. Natl. Acad. Sci. USA 92, 1945–1949 (1995).
Rodnina, M.V. et al. Thiostrepton inhibits turnover but not GTP hydrolysis by elongation factor G on the ribosome. Proc. Natl. Acad. Sci. USA 96, 9586–9590 (1999).
Johnson, A.E., Adkins, H.J., Matthews, E.A. & Cantor, C.R. Distance moved by transfer RNA during translocation from the A site to the P site on the ribosome. J. Mol. Biol. 156, 113–140 (1982).
Fahnestock, S., Neumann, H. & Rich, A. Assay of ester and polyester formation by the ribosomal peptidyltransferase. Methods Enzymol. 30, 489–497 (1974).
Stern, S., Moazed, D. & Noller, H.F. Structural analysis of RNA using chemical and enzymatic probing monitored by primer extension. Methods Enzymol. 164, 481–489 (1988).
Acknowledgements
We thank W. Wintermeyer for discussion and valuable comments on the manuscript, H.-J. Wieden for fMet-tRNAfMet(Flu) and Phe-tRNAPhe(QSY), Y.P. Semenkov and V.I. Katunin (Petersburg Nuclear Physics Institute) for generous gifts of tRNAs, D. Rodnin for ribosome preparations and A. Böhm, P. Striebeck, C. Schillings and S. Möbitz for expert technical assistance. The work was supported by the Deutsche Forschungsgemeinschaft, the European Union, the Alfried Krupp von Bohlen und Halbach-Stiftung and the Fonds der Chemischen Industrie.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Bieling, P., Beringer, M., Adio, S. et al. Peptide bond formation does not involve acid-base catalysis by ribosomal residues. Nat Struct Mol Biol 13, 423–428 (2006). https://doi.org/10.1038/nsmb1091
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1091
This article is cited by
-
A proton wire to couple aminoacyl-tRNA accommodation and peptide-bond formation on the ribosome
Nature Structural & Molecular Biology (2014)
-
Novel base triples in RNA structures revealed by graph theoretical searching methods
BMC Bioinformatics (2011)
-
Different substrate-dependent transition states in the active site of the ribosome
Nature (2011)
-
A two-step chemical mechanism for ribosome-catalysed peptide bond formation
Nature (2011)
-
The 2′-OH group of the peptidyl-tRNA stabilizes an active conformation of the ribosomal PTC
The EMBO Journal (2011)