Abstract
During tumourigenesis, p53 functions as 'the guardian of the genome' because p53-dependent apoptosis strongly regulates the fate of cancer cells. Therefore, p53 regulation must be sensitive and accurate. p53 activity is regulated through its ubiquitination and deubiquitination. However, the role of microRNA in ubiquitin-mediated p53 degradation has not been previously studied. Our previous studies indicated that miR-100 is required for apoptosis. In the current study, the mechanism of p53 protein ubiquitination mediated by miR-100 was characterized. An analysis of primary tumour samples from gastric cancer patients showed a significant correlation between miR-100 upregulation and primary human gastric tumourigenesis and progression. The in vivo and in vitro data indicated that miR-100 antagonism specifically induced the apoptosis of poorly differentiated gastric cancer cells but not non-cancerous gastric cells, indicating that miR-100 has a crucial role in regulating the progression of gastric tumours. In the regulation of p53-dependent apoptosis, miR-100 antagonism inhibited ubiquitin-mediated p53 protein degradation by activating RNF144B, an E3 ubiquitination ligase. Consequently, the miR-100-RNF144B-pirh2-p53-dependent pathway was initiated. Our findings highlight a novel mechanism of ubiquitin-mediated p53 protein degradation in apoptosis.
Similar content being viewed by others
Introduction
Apoptosis, or programmed cell death, can be induced by various triggers.1 By contrast, tumourigenesis results from numerous complex factors that inhibit apoptosis, which may functionally contribute to tumour growth and invasion.2 Altered tumour-suppressor gene function is thought to be the main cause of apoptosis inhibition during tumourigenesis.3, 4 The p53-triggered apoptotic pathway likely functions as a mechanism to protect organisms from impaired cells that may have malignant potential.5 Cells with a high apoptotic tendency display a combination of low growth rates, low cellular turnover and low malignant probability.
Recent studies have shown that p53 can function as a ‘guardian of the organism’ because p53-dependent apoptosis directly regulates the fate of both normal and cancer cells in vivo. The p53 protein has a negative role in tumour growth, as found in SV40 transgenic mice.6 For example, when mice were bred onto a p53-null background, the tumour proliferation rates increased dramatically, with a concordant decrease in apoptosis. Thus, the p53 protein functions as a tumour suppressor by controlling the fate of tumour cells through the regulation of the apoptotic pathway.
Owing to the critical role of p53 as a tumour-suppressor gene in human cancer, p53 regulation must be both sensitive and accurate. The p53 gene sequence has been reported to be mutated in >50% of human cancers.7 miR-125b targets the p53 gene and regulates its expression at the transcriptional level.8 Moreover, the regulation of ubiquitin-mediated p53 protein degradation is responsible for p53 inactivation in cancer,9 and ubiquitin-dependent proteolysis is the major pathway for regulating protein turnover via a mechanism that is dependent, at least in part, on activating ubiquitin-activating enzyme (E1), ubiquitin-conjugating enzyme (E2), and ubiquitin ligase (E3), which catalyse ubiquitin to protein lysine residues in an energy-dependent manner.10, 11 Some recent studies have shown that the ubiquitin-proteasome pathway regulates the stability of many onco-proteins and tumour suppressors, leading to the essential role of this pathway in cancer development and progression.12–14 These studies have indicated that p53 activation is mainly accompanied by protein stabilisation, including p53 protein ubiquitination and deubiquitination. When four or more ubiquitin monomers are targeted to the p53 protein, proteasomal degradation is triggered.
The first identified p53 E3 ubiquitin ligase was Mdm2 (murine double minute 2),15 followed by PIRH2 (ring finger and CHY zinc finger domain containing 1) and E3 ubiquitin ligase, which mediates p53 ubiquitination.16 Many p53 E3 ligases have been discovered, including HUWE1 (HECT, UBA and WWE domain containing 1), WWP1 (WW domain containing E3 ubiquitin protein ligase 1), COP1 (constitutively photomorphogenic 1) and STUB1 (STIP1 homology and U-box containing protein 1).17, 18, 19, 20 These E3 ligases have critical roles in the ubiquitin-mediated p53 protein degradation pathway, which has essential effects on cancer cell apoptosis and is regulated by many factors. However, it is still unknown precisely how microRNAs (miRNAs) mediate p53 protein ubiquitination/deubiquitination.
miRNAs, as endogenous silencers of various target genes via translational repression or mRNA degradation,21, 22 have essential roles in regulating many diseases, including cancer.23 Computational prediction studies reveal that ~50% of miRNA-coding genes are located in tumour-associated genomic regions,24, 25 thus revealing a relationship between miRNA and the development and progression of cancer. In addition, the regulation of miRNA-mediated gene expression brings new insight into the putative biological mechanisms that contribute to human cancer.26
In our previous studies, we revealed that miR-100 is required for apoptosis in shrimp and human cancers.27, 28 In this study, we characterized miR-100 to reveal the role of miRNA in ubiquitin-mediated p53 protein degradation. The results showed that miR-100 could directly trigger the degradation of p53 during the regulation of gastric cancer apoptosis. Furthermore, our study reveals a novel mechanism of p53 protein ubiquitination.
Results
High levels of miR-100 correlate with gastric tumourigenesis and progression
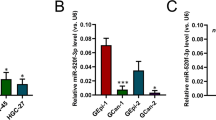
To characterize the role of miR-100 in human cancer apoptosis, we assayed miR-100 expression levels in nine cell lines derived from human liver, gastric, colorectal, and breast cancers and macrophage-derived leukaemia. We found that miR-100 was specifically upregulated in human epithelium-derived gastric cancer cells (MGC-803 and HGC-27) compared with human epithelium-derived gastric non-cancerous cells (GES-1; Figure 1a). To further evaluate miR-100 expression in gastric cancer cells, we characterized poorly differentiated gastric cancer cells (MKN-45, MGC-803 and HGC-27) and highly differentiated gastric cancer cells (AGS and N87). The results showed that miR-100 was significantly upregulated in poorly differentiated gastric cancer cells compared with GWS-1 but that it was significantly downregulated in highly differentiated gastric cancer cells (Figure 1a). Our previous study reported that miR-100 upregulation was involved in the apoptosis of breast cancer cells (SK-BR-3).28 In the present investigation, the results indicate that the miR-100 silencing led to the increase of caspase 3/7 activity in gastric cancer cells (MKN-45, MGC-803 and HGC-27; Figure 1b), suggesting that miR-100 may participate in the apoptosis regulation of gastric cancer.
High miR-100 levels correlate with gastric tumour progression. (a) miR-100 expression in gastric cancer and non-cancerous cells. miR-100 expression levels were examined in normal gastric epithelial cells (GES-1), poorly differentiated gastric cancer cells (MKN-45, MGC-803 and HGC-27), and highly differentiated gastric cancer cells (AGS and N87) using quantitative real-time PCR. (b) The influence of miR-100 silencing on the apoptosis of cancerous and non-cancerous gastric cells. GES-1, MGC-803, HGC-27 and MKN-45 cells were transfected with AMO-miR-100. At 36 h after transfection, the caspase 3/7 activities of the cells were examined. (c) miR-100 expression in tumour specimens from gastric cancer patients. A paired sample comprising cancerous tissues and corresponding normal tissues from the same patient is shown using haematoxylin-and-eosin-stained tissues sections (× 400). The expression analysis of miR-100 (scaled to 100%) is shown for ten pairs of specimens. (d) Mean expression of miR-100 in 90 paired samples. The miR-100 expression level was evaluated by quantitative real-time PCR. (e) Expression of miR-100 in gastric tumours by grade using haematoxylin-and-eosin-stained tissue sections (× 400 magnification). Samples representing grade 1 (n=19), grade 2 (n=28) and grade 3 (n=37) were analysed. The lines represent the means of miR-100 expression. Significant differences are indicated by asterisks (*P<0.05; **P<0.01).
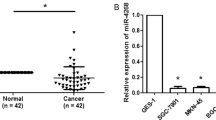
To determine whether miR-100 expression correlates with clinical outcome, we measured miR-100 expression in primary tumour samples from gastric cancer patients. Cancerous tissue and corresponding normal tissue from the same patients were examined as paired samples (Figure 1c). An expression analysis of miR-100 in 10 sample pairs showed higher miR-100 levels in cancerous tissues than in normal tissues (Figure 1c). There was a significant correlation between miR-100 expression levels and tumourigenesis.
To further address this issue, we examined 90 tumour sample pairs and found that the mean miR-100 expression levels in cancerous tissues was significantly higher than that in normal tissues (Figure 1d). Generally, gastric tumours can be classified into one of three grades (1–3) according to the degree of tumour cell differentiation, as detected by histopathology. miR-100 expression significantly increased with the increasing degree of malignancy (Figure 1e).
Taken together, these data reveal a significant correlation between augmented miR-100 expression and primary human gastric tumourigenesis and progression.
miR-100 inhibits the apoptosis of gastric cancer cells
miR-100 expression was characterized to determine whether the observed high levels of miR-100 expression correlated with apoptosis regulation in gastric cancer cells. For a comprehensive evaluation of the effect of miR-100 on gastric cancer cell apoptosis, miR-100 was silenced and overexpressed in cancer cells. miR-100 silencing or overexpression had no effect on AGS and N87 cell apoptosis (data not shown). Therefore, miR-100 was characterized in the poorly differentiated gastric cancer cells MKN-45, MGC-803 and HGC-27.
miR-100 expression was knocked down in gastric cancer cells (MKN-45, MGC-803 and HGC-27) and non-cancerous cells (GES-1) using the anti-miRNA oligonucleotide (AMO) AMO-miR-100 (Figure 2a), which specifically silences miR-100 expression (Figure 2b). The results of apoptosis and cell viability assays revealed that following cancer cell (MKN-45, MGC-803 and HGC-27) transfection with AMO-miR-100-scrambled, apoptosis remained at baseline levels and cell viability increased. However, when AMO-miR-100 was introduced, apoptosis sharply increased, and cell viability decreased (Figure 2c). In contrast, apoptosis only increased slightly due to exogenous DNA transfection into GES-1 cells, regardless of miR-100 knockdown (Figure 2c). Based on dosage assays, the apoptosis of cancer cells induced by miR-100 silencing was dose dependent (Figure 2d). In all apoptosis experiments, 50 nM AMO-miR-100 was used. The results of apoptosis detection with annexin V assays confirmed that silencing miR-100 expression led to the apoptosis of gastric cancer cells but not to that of non-cancerous gastric cells (Figure 2e). Furthermore, immunoblots of cleaved poly (ADP-ribose) polymerase 1 and caspase 3, which are key proteins in the apoptotic pathway, showed that these two proteins could only be detected in cancer cells that were transfected with AMO-miR-100, indicating that apoptosis occurred only in gastric cancer cells when miR-100 expression was inhibited (Figure 2f). These findings suggest that miR-100 has an important role in the apoptosis regulation of gastric cancer cells but not GES-1.
miR-100 inhibits the apoptosis of gastric cancer cells. (a) Nucleotide sequences and specific modifications of anti-miRNA-100 oligonucleotide (AMO-miR-100) and the control AMO-miR-100-scrambled. (b) miR-100 expression in miR-100-silenced cells. AMO-miR-100 was transfected into cells to knockdown miR-100 expression levels, which were then measured by quantitative real-time PCR. AMO-miR-100-scrambled oligonucleotide was used as a control. (c) The effects of miR-100 silencing on apoptosis and cellular viability using ApoTox-Glo Triplex assays. (d) The dose-dependent effects of miR-100 silencing on the apoptosis of cancerous and non-cancerous cells. GES-1, MGC-803, HGC-27 and MKN-45 cells were transfected with AMO-miR-100 at various concentrations. At 36 h after transfection, the caspase 3/7 activity of the cells was examined. (e) The effects of miR-100 silencing on apoptosis as determined by an annexin V assay. The apoptosis levels were measured by flow cytometry at 24 h after AMO transfection. (f) The detection of key proteins in the apoptosis of cancer cells and non-cancerous cells that were treated with AMOs. Cleaved PARP (poly ADP-ribose polymerase) and cleaved caspase 3 were detected by western blotting, and β-tubulin was used as a control. (g) The effects of miR-100 overexpression on apoptosis. miR-100 expression data were obtained by quantitative real-time PCR (left), and cell proliferation data were obtained by a specific assay (right). In all of the panels, the plotted data points refer to the mean±s.d. of triplicate assays, and asterisks represent significant differences (**P<0.01).
To evaluate the effects of miR-100 overexpression on cancer cell apoptosis, the miR-100 precursor was transfected into poorly differentiated gastric cancer cells (MKN-45, MGC-803 and HGC-27) and GES-1. The results showed that miR-100 precursor overexpression only partially stimulated the rate of cell growth in cancer cells and non-cancerous cells alike (Figure 2g). However, after miR-100 precursor transfection, the apoptosis levels were unchanged in both cancer and non-cancer cells (not shown). The data presented that miR-100 overexpression took no effect on apoptosis of gastric cancerous and non-cancerous cells.
These collective findings show that miR-100 antagonism could induce the apoptosis of poorly differentiated gastric cancer cells but not of GES-1. MiR-100 antagonism has no negative effect on normal gastric cells.
miR-100 antagonism promotes the apoptosis of gastric cancer in vivo
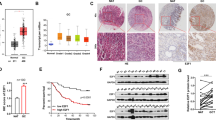
To investigate the role of miR-100 in tumour cell apoptosis in vivo, we used two approaches to establish experimental gastric cancer animal models (Figure 3a). First, we used AMO-miR-100 to inhibit miR-100 expression in gastric cancer cells (MGC-803). At 6 h after transfection, the cells were adoptively transferred into non-obese diabetic, severe combined immunodeficient mice. The tumour growth rate, weight and size (at 44 days after cell transplantation) in these miR-100-knockdown cells were significantly lower than those in control cells (AMO-miR-100-scrambled-treated cells; Figure 3b), indicating that miR-100 antagonism could inhibit tumourigenesis in vivo.
miR-100 antagonism promotes the apoptosis of gastric cancer cells in vivo. (a) A flow chart of the in vivo experiments. (b) The effects of inhibiting miR-100 expression on tumourigenicity in nude mice. MGC-803 cells were transfected with AMOs followed by the subcutaneous injection of the cells into nude mice. The mean numeral data are the product of 10 mice. The tumour weights and images were obtained 44 days after cell implantation. (c) The effects of miR-100 silencing on solid tumours in nude mice. MGC-803 cells were injected into nude mice. Next, AMOs were subcutaneously and intravenously injected into the mice. The numerical data are the means of 10 mice. The tumour weights and images were obtained 31 days after AMO injection. (d) miR-100 expression in tumour tissues as determined by quantitative real-time PCR. (e) Integration of the luciferase gene into the MGC-803 cell genome. The luciferase gene was cloned into the LV plasmid to generate the LV- EF1α-eGFP-TRE-luciferase construct. Then, this construct and the LP1, LP2 and VSV plasmids were co-transfected into the cells. The cells were subjected to PCR to confirm the integration of the EF1α-eGFP-TRE-luciferase fragment into the cell genomes. HIV LTR, HIV long terminal repeat; eGFP, enhanced green fluorescent protein; EF1α, EF1 alpha promoter; TRE, tetracycline response element. (f) In vivo luminescent imaging of apoptosis in tumour cells. To visualize tumour apoptosis after miR-100 knockdown in vivo, a doxycycline-controlled luciferase gene was integrated into the MGC-803 cell genome. Then, the cells were adoptively transferred into mice, followed by AMO injection. The apoptotic cells in mice produced positive signals after the injection of a luciferin-conjugated caspase substrate. (g) Immunohistological analysis of tumour tissue apoptosis in vivo. Tumour tissues were analysed by immunohistochemistry using antibodies against ki-67 or caspase 3. The positive signals are indicated with arrows. In all panels, the data refer to the mean±s.d. of triplicate assays (**P<0.01).
To further evaluate the effect of miR-100 on solid tumours, MGC-803 cells were adoptively transferred into nude mice followed by the repeated injection of AMO-miR-100 or AMO-miR-100-scrambled every 3 days for 1 month. The tumour growth rate in AMO-miR-100-injected mice was significantly lower than that in AMO-miR-100-scrambled-injected mice at 10 days following AMO injection (Figure 3c). At 31 days after AMO injection, the tumour sizes and weights were significantly lower in AMO-miR-100-injected mice (Figure 3c). In addition, the miR-100 levels were significantly downregulated compared with those in tumours that were treated with AMO-miR-100-scrambled. These data indicated that the downregulation of miR-100 inhibited tumour growth in vivo (Figure 3d).
Our data show that miR-100 has an important role in regulating tumour growth in vivo. To visualize tumour apoptosis after miR-100 knockdown in vivo, a doxycycline-controlled luciferase gene was integrated into the MGC-803 cell genome through viral infection (Figure 3e). The cells were adoptively transferred into mice, followed by AMO injection. By injecting the luciferin-conjugated caspase substrate, the activated caspase in apoptotic cells would catalyse the substrate to release luciferin. The released luciferin would then become a substrate for luciferase, and the signal could be detected. The in vivo imaging results showed that positive signals were produced in mice that were treated with miR-100 antagonism but were absent in control mice. This observation indicates that apoptosis occurred in tumours when miR-100 expression was inhibited, whereas the control AMO-miR-100-scrambled had no effect on tumour cell apoptosis (Figure 3f). Tumour tissue immunohistochemistry revealed that miR-100 knockdown resulted in the dampened expression of the tumour proliferation marker Ki-67 and in the enhanced protein expression of caspase 3 compared with those of the control (Figure 3g).
The above findings suggest that tumour growth inhibition in vivo was due to tumour cell apoptosis, which was caused by reduced miR-100 expression.
miR-100 antagonism inhibits ubiquitin-mediated p53 protein degradation in gastric cancer cells
The transcription factor p53 has been reportedly placed at the centre of a complex molecular network that regulates apoptosis. The p53 targets include p21, PUMA, Bax and Bid. To evaluate whether miR-100 was involved in p53-mediated gastric cancer cell apoptosis, miR-100 expression in MKN-45 and MGC-803 cells was knocked down using miR-100 antagonism (AMO-miR-100), followed by the detection of p53, p21, PUMA, Bax, Bid and cleaved caspase 3 protein expression. The sequencing analysis revealed an amino-acid mutation (from proline to arginine) at the 72 site of the p53 protein in HGC-27 cells. The HGC-27 cell line was not included in this characterisation because the miR-100 expression level in HGC-27 cells was at a level between those found in MKN-45 cells and MGC-803 cells (Figure 1a). Western blotting showed that miR-100 silencing significantly enhanced the expression of p53, p21, PUMA, Bax, Bid and cleaved caspase 3 (that is, the active form of caspase 3) in MKN-45 and MGC-803 cells (Figure 4a), showing the involvement of miR-100 in p53-mediated gastric cancer cell apoptosis. When the cells were simultaneously transfected with AMO-miR-100 and p53-siRNA, the protein levels of p21, PUMA, Bax, Bid and cleaved caspase 3 significantly decreased (Figure 4a), indicating that p53 silencing in the miR-100-silenced cells led to the downregulation of the targets of p53. These findings indicate that miR-100 negatively modulated p53 expression and p53-mediated gastric cancer cell apoptosis.
miR-100 antagonism inhibits ubiquitin-mediated p53 protein degradation in gastric cancer cells. (a) Immunoblots of p53 and its target proteins. The proteins p53, p21, PUMA, Bax, Bid, cleaved caspase 3 and β-tubulin in MKN-45 and MGC-803 cells that were treated with AMO-miR-100, AMO-miR-100-scrambled or AMO-miR-100+p53-siRNA were examined via western blotting. (b) The degradation of the p53 protein in gastric cancer cells through ubiquitination. MGC-803, MKN-45 and GES-1 cells were treated with MG-132. Non-treated cells were used as controls. At 24 h after MG-132 treatment, the cells were subjected to quantitative real-time PCR (upper panel) and western blotting (lower panel). β-tubulin expression was used as a control. (c) The effect of miR-100 on the ubiquitin-mediated degradation of p53 protein in gastric cancer cells. MGC-803, MKN-45 and GES-1 cells were transfected with AMO-miR-100 and then subjected to quantitative real-time PCR (upper panel) and western blotting (lower panel) to detect p53 mRNA and protein expression. AMO-miR-100-scrambled was included in the transfection as a control. (d) The detection of ubiquitinated p53 protein in gastric cancer and non-cancerous cells. The MGC-803, MKN-45 and GES-1 cells were transfected with AMO-miR-100. Non-treated cells were used as controls. At 24 h after transfection, the cells were assayed for the level of ubiquitinated p53 protein by western blotting with the p53-specific antibody. β-tubulin was used as a loading control. The images show p53 protein expression with different levels of ubiquitination.
The p53 sequence is mutated in most cancer cells, and its expression inhibits p53-mediated apoptosis. In this study, p53 sequencing showed that the p53 sequence mutation did not occur in gastric cancer cells (MGC-803 and MKN-45) or GES-1 (data not shown). Quantitative real-time PCR revealed that p53 was transcribed normally in MGC-803, MKN-45 and GES-1 cells (Figure 4b). However, the p53 protein was only detected in GES-1 cells (Figure 4b), suggesting that the p53 protein in gastric cancer cells (MGC-803 and MKN-45) was probably degraded by ubiquitination. To address this issue, the cells were treated with MG-132, which is a specific inhibitor of ubiquitin-conjugated protein. The p53 protein was detected in MG-132-treated MGC-803 and MKN-45 cells (Figure 4b), indicating that p53 protein degradation in gastric cancer cells depends on p53 protein ubiquitination. However, in non-cancerous cells (GES-1), ubiquitin-mediated p53 protein degradation did not occur (Figure 4b).
To evaluate whether the negative effect of miR-100 on p53 expression in gastric cancer cells resulted from p53 protein ubiquitination, MGC-803 and MKN-45 cells were transfected with AMO-miR-100, followed by a determination of the p53 mRNA and protein levels. The p53 mRNA levels in the AMO-miR-100-treated MGC-803 and MKN-45 cells were unchanged compared with those of the controls, whereas the p53 protein levels markedly increased (Figure 4c). These observations indicate that miR-100-mediated knockdown could inhibit p53 protein degradation in gastric cancer cells. However, miR-100 had no effect on p53 expression in non-cancerous cells (Figure 4c). These data reveal that miR-100 had significant effects on gastric cancer cell p53 protein degradation. We simultaneously examined ubiquitinated p53 protein expression in AMO-miR-100-treated cells. The results showed that p53 protein ubiquitination was significantly inhibited in AMO-miR-100-treated MGC-803 and MKN-45 cells (Figure 4d), indicating that miR-100 was required for p53 protein ubiquitination in gastric cancer cells.
In this context, miR-100 had a positive role in ubiquitin-mediated p53 protein degradation in poorly differentiated gastric cancer cells but not in non-cancerous gastric cells.
E3 ubiquitination ligase pirh2 participates in miR-100-mediated p53 protein ubiquitination in gastric cancer cells
We characterized p53-specific E3 ubiquitin ligases to determine the mechanism of miR-100-mediated p53 ubiquitination in gastric cancer cells. The six p53 E3 ubiquitin ligases (pirh2, COP1, WWP1, STUB1, MDM2 and HUWE1) were silenced by sequence-specific small interfering RNAs (siRNAs) in MGC-803 and MKN-45 cells (Figure 5a). The detection of the p53 protein showed that p53 specifically accumulated when pirh2 expression was knocked down by sequence-specific siRNA (Figure 5b). However, the negative control of pirh2-siRNA showed no effect on p53 expression (Figure 5b). These data indicate that pirh2 is the key E3 ubiquitin ligase of p53 ubiquitination in gastric cancer cells. As reported, COP1 is a critical negative regulator of p53 in U2-OS cells.19 In MGC-803 and MKN-45 cells, however, the expression level of COP1 protein was very low (Figure 5c). Therefore, COP1 knockdown had no effect on the p53 protein level in gastric cancer cells (MGC-803 and MKN-45; Figure 5b). To further evaluate the effects of miR-100 on pirh2 expression, miR-100 was, respectively, knocked down and overexpressed in poorly differentiated gastric cancer cells (MGC-803 and MKN-45) and non-cancerous gastric cells (GES-1), followed by an evaluation of pirh2 and p53 protein expression. miR-100 antagonism significantly decreased pirh2 expression levels and simultaneously increased p53 expression levels in MGC-803 and MKN-45 cells compared with those of the control (that is, cells without any treatment). By contrast, miR-100 overexpression had no marked effect on the expression of pirh2 and p53 compared with the control (that is, cells without any treatment; Figure 5d). However, miR-100 silencing and overexpression had no effect on the protein expression of pirh2 and p53 in GES-1 cells (Figure 5d). These results showed that miR-100 antagonism could inhibit the pirh2 expression, leading to the upregulation of p53 in gastric cancer cells but not non-cancerous cells.
The E3 ubiquitin ligase pirh2 participates in miR-100-mediated p53 protein ubiquitination in gastric cancer cells. (a) Silencing the expression of the E3 ubiquitin ligases pirh2, COP1, WWP1, MDM2, STUB1 and HUWE1 in gastric cancer cells. MGC-803 and MKN-45 cells were transfected with sequence-specific siRNAs. The negative control siRNAs (negative controls) were included in the transfections. Next, the mRNA levels of six E3 ubiquitin ligases were examined by quantitative real-time PCR. (b) Effects of expression silencing of E3 ubiquitin ligase on p53 ubiquitination in gastric cancer cells. In the siRNA-treated MGC-803 and MKN-45 cells, p53 protein was detected by western blotting. The cells that were transfected with the negative control siRNAs (negative controls) were also analysed. β-tubulin was used as a control. The genes that were treated with sequence-specific siRNA or negative control are shown on top. (c) The expression level of COP1 in gastric cancer cells. The COP1 protein in cancerous MGC-803 and MKN-45 cells and non-cancerous GES-1 cells was examined via western blotting. β-tubulin was used as a control. The antibodies that were used are indicated on the right. (d) The influence of miR-100 silencing and overexpression on pirh2 gene expression in GES-1, MGC-803 and MKN-45 cells. The cells were transfected with AMO-miR-100 or the miR-100 precursor, followed by the detection of pirh2 and p53 proteins via western blotting. (e) The effect of pirh2 silencing on miR-100-mediated p53 activation in cells. The GES-1, MGC-803 or MKN-45 cells were transfected with AMO-miR-100, miR-100 precursor or/and pirh2-siRNA. Western blotting was used to detect the pirh2 and p53 proteins. (f) The impact of pirh2 overexpression on miR-100-mediated p53 activation in GES-1, MGC-803 and MKN-45 cells. The cells were transfected with AMO-miR-100, miR-100 precursor or/and the plasmid expressing pirh2. Then, the pirh2 and p53 proteins were detected via western blotting, β-tubulin was used as a control.
To examine the involvement of pirh2 in miR-100-mediated p53 activation, pirh2 expression was knocked down or overexpressed in miR-100-silenced or miR-100-overexpressed cells. These data demonstrate that pirh2 silencing caused p53 protein accumulation in miR-100-silenced cancerous cells (GES-1 and MGC-803) (Figure 5e), whereas pirh2 overexpression decreased the p53 protein level in miR-100-silenced cancerous cells (Figure 5f). Pirh2 silencing and overexpression had no effect on the p53 protein level in non-cancerous cells (GES-1) (Figures 5e and f). These data suggest the involvement of E3 ubiquitin ligases pirh2 protein in miR-100-mediated p53 activation.
Taken together, these findings indicate that the p53 E3 ubiquitin ligase pirh2 is involved in the miR-100-mediated apoptosis pathway in gastric cancer cells but not GES-1. Based on target gene prediction, pirh2 is not a miR-100 target gene.
miR-100 triggers the p53 protein ubiquitin pathway in gastric cancer cells in vitro and in vivo
miR-100 target gene prediction showed that RNF144B, which is an E3 ubiquitin ligase, might be a miR-100 target gene (Figure 6a). To characterize the interaction between miR-100 and RNF144B, a dual-luciferase reporter assay was conducted in MGC-803 and MKN-45 cells. The results revealed that miR-100 significantly decreased the activity of the reporter luciferase that was fused with the wild-type RNF144B 3’-untranslated region (UTR) compared with the controls (Figure 6b). This observation indicated a direct interaction between miR-100 and RNF144B mRNA. To assess the interaction between miR-100 and RNF144B in vivo, miR-100 was silenced and overexpressed in MGC-803 and MKN-45 cells, respectively, after which RNF144B expression was monitored. These results demonstrate that miR-100 silencing resulted in enhanced RNF144B expression, whereas miR-100 overexpression markedly suppressed RNF144B expression levels (Figure 6c). The data indicated that miR-100 could negatively regulate RNF144B expression in gastric cancer cells. Therefore, RNF144B is a miR-100 target gene. To explore the role of RNF144B in apoptosis, RNF144B-silenced, RNF144B-overexpressed, or RNF144B-rescued MGC-803 cells were subjected to apoptosis assays. The results showed that both the overexpression and the rescue of RNF144B resulted in increased cleaved poly (ADP-ribose) polymerase 1 and cleaved caspase 3, indicating that apoptosis was activated (Figure 6d). The activation of apoptosis by RNF144B overexpression or rescue was further confirmed by the detection of caspase 3/7 activity (Figure 6e). The above data suggested that RNF144B is a target of miR-100 and is involved in the regulation of apoptosis.
miR-100 triggers p53 protein ubiquitination in gastric cancer cells in vitro and in vivo. (a) Target gene analysis of miR-100. Based on the TargetScan, miRanda, PicTar and miRInspector algorithms, RNF144B was considered a miR-100 target gene. (b) The direct interaction between miR-100 and RNF144B 3’-UTR. MGC-803 and MKN-45 cells were co-transfected with miR-100 and a luciferase reporter fused with RNF144B 3’-UTR or the RNF144B 3’-UTR mutant. Then, the firefly and Renilla luciferase activities were evaluated. (c) The interactions between miR-100 and RNF144B in vivo. MGC-803 and MKN-45 cells were transfected with AMO-miR-100 or the miR-100 precursor, and RNF144B expression was examined by quantitative real-time PCR (top panel) and western blotting (lower panel). (d) The role of RNF144B in the apoptosis of MGC-803 cells. Whole-cell extracts of cells expressing the indicated RNF144B-siRNA or/and ΔRNF144B were analysed by western blotting for the indicated proteins. (e) The detection of the caspase 3/7 activity in MGC-803 cells. The treatments were as indicated. (f) The interaction between RNF144B and pirh2 in MGC-803 cells. The cells were transiently transfected with flag-RNF144B and then co-immunoprecipitated with anti-flag. The eluted proteins were analysed by SDS-PAGE (silver staining), followed by protein identification using mass spectrometry (1st panel). To confirm the interaction between RNF144B and pirh2, flag-tagged RNF144B and/or His-tagged pirh2 were co-transfected into cells and then subjected to anti-Flag immunoprecipitation (2nd panel) and/or anti-His immunoprecipitation (3rd panel), respectively. To exclude the non-specific pull-down of His-pirh2 by flag antibody, the flag-RNF144B and His-pirh2 co-transfected cells were subjected to CoIP assays with anti-His or anti-flag (4th panel). The immunoprecipitates were analysed by western blotting using the antibodies as indicated. The total proteins of the cells (input) were included in western blotting. (g) The role of RNF144B in the p53-mediated apoptosis of gastric cancer cells. The expression of RNF144B was knocked down, overexpressed or rescued in MGC-803 and MKN-45 cells. Then, the expressions of pirh2 and p53 were examined by western blotting. For the overexpression of RNF144B, the RNF144B gene sequence was synonymously mutated at two nucleotides (underlined) to avoid recognition by RNF144B-siRNA. β-tubulin was used as a control. (h) pirh2 ubiquitination by RNF144B in MGC-803 cells. The cells were transfected with constructs expressing flag-tagged ubiquitin and/or His-pirh2 (upper panel), or flag-tagged ubiquitin, His-pirh2 and/or RNF144B-siRNA (middle panel). Whole-cell extracts were prepared and subjected to western blotting for the presence of the indicated proteins (lower panel) or precipitated with anti-flag bead pull-down and then followed for the detection of the antibody as indicated (upper panel). At the same time, the cells were incubated with MG-132 (bottom panel). Whole-cell extracts either were prepared from cells that were treated with MG-132 and analysed by western blotting directly or were immunoprecipitated with anti-RNF144B and then by western blotting with the antibodies as indicated. (i) Interactions between the RNF144B, pirh2 and p53 proteins in MGC-803 cells. Whole-cell extracts were prepared from cells that were treated with MG-132 and then incubated with anti-RNF144B antibody (upper panel), anti-p53 antibody (middle panel) or anti-pirh2 antibody (lower panel) to immunoprecipitate the associated proteins, which were then subjected to western blotting with the antibodies indicated on the right. (j) The mechanism that is responsible for miR-100-antagonism-induced apoptosis. Whole-cell extracts of cells that were transfected with His-RNF144B or AMO-miR-100 were analysed by western blotting for the indicated proteins. (k) The involvement of miR-100 in ubiquitin-mediated p53 degradation in tumour specimens from gastric cancer patients. A paired sample comprising cancerous tissue and the corresponding normal tissue from the same patient was subjected to quantitative real-time PCR (upper panel) and western blotting (lower panel). Fifteen patients were examined. The numeral data represent the mean±s.d. of triplicate assays.
To determine how RNF144B conveyed apoptotic signals from miR-100 knockdown, we overexpressed flag-tagged RNF144B in MGC-803 cells and performed co-immunoprecipitation (CoIP) with anti-flag to identify proteins that interacted with RNF144B. The results indicate that RNF144B bound to pirh2 (Figure 6f, 1st panel). Furthermore, to confirm the interaction between RNF144B and pirh2, flag-RNF144B and Histidine-tagged pirh2 (His-pirh2) were co-transfected into MGC-803 cells, followed by CoIP assays using anti-flag or anti-His antibody. The western blotting data show that RNF144B directly interacted with pirh2 (Figure 6f, 2nd and 3rd panels). In order to exclude the non-specific pull-down of His-pirh2 by the anti-flag antibody, the flag-RNF144B and His-pirh2 co-transfected MGC-803 cells were subjected to CoIP assays with anti-His or anti-flag. Western blots indicate that the CoIP assays were antibody-specific (Figure 6f, 4th panel). These findings reveal that the RNF144B protein was directly interacted with the pirh2 protein.
To investigate the role of RNF144B in p53-mediated gastric cancer cell apoptosis, the gene expression of RNF144B was silenced, overexpressed or rescued in MGC-803 and MKN-45 cells, followed by pirh2 and p53 protein detection. RNF144B overexpression markedly suppressed pirh2 expression and simultaneously induced p53 expression in gastric cancer cells (Figure 6g). However, silencing the RNF144B gene had no effect on either pirh2 or p53 expression due to the low level of RNF144B expression in gastric cancer cells (Figure 6g). The results show that rescuing RNF144B gene expression in RNF144B-silenced cells yielded similar expression profiles of pirh2 and p53 to those obtained from the overexpression treatment (Figure 6g). The above data indicate that RNF144B is involved in p53-mediated gastric cancer cell apoptosis by regulating pirh2 expression.
To determine whether pirh2 could be polyubiquitinated by RNF144B, flag-tagged ubiquitin (Flag-Ub) and His-pirh2 were co-expressed in MGC-803 cells, and anti-flag beads were used to pull-down modified proteins. The presence of polyubiquitinated pirh2 was observed as a smeared band because of the heterogeneous modification on this protein (Figure 6h), suggesting that pirh2 was polyubiquitinated by RNF144B. To confirm that this modification was mediated by RNF144B, we transfected RNF144B-siRNA into MGC-803 cells and then stained the polyubiquitinated pirh2 in the flag-Ub immunoprecipitants. RNF144B knockdown resulted in a drastic reduction of polyubiquitinated pirh2 compared with the control (Figure 6h), showing that RNF144B could mediate the ubiquitination of pirh2. MG-132 could inhibit pirh2 degradation (Figure 6h), indicating the stabilisation of the interaction between RNF144B and pirh2. These findings reveal that RNF144B is an E3 ligase for pirh2 and that RNF144B polyubiquitinates pirh2 for degradation.
To further understand how RNF144B, pirh2 and p53 associate with each other to regulate apoptosis, we examined whether there were functional interactions among these proteins. MG-132 was used to block the degradation of pirh2 and p53, and then an RNF144B CoIP was performed to examine the existence of the other two proteins. Pirh2 but not p53 was indeed detected from the RNF144B immunoprecipitates, which showed that p53 could not interact with RNF144B (Figure 6i). The reciprocal CoIP using p53 antibody demonstrated that p53 interacted with pirh2 but not with RNF144B (Figure 6i). By contrast, pirh2 interacted with both RNF144B and p53 (Figure 6i). Thus, these data indicate that RNF144B and p53 proteins exist in two mutually exclusive pirh2-containing complexes. To explore the mechanism of miR-100-induced apoptosis, we transfected His-RNF144B and AMO-miR-100 into MGC-803 cells (Figure 6j). Both ectopically expressed RNF144B and knocked down miR-100 induced pirh2 degradation because of RNF144B accumulation (Figure 6j). Owing to the lack of E3 ubiquitin ligase for its degradation, the level of p53 protein was then restored to further trigger cellular apoptosis. In this context, these data reveal that miR-100 antagonism mediates apoptosis by restoring p53 levels in cells.
To evaluate this issue in vivo, the miR-100, RNF144B, pirh2, and p53 expression levels were characterized in primary tumour samples from gastric cancer patients. Cancerous tissue and corresponding normal tissue from the same patients were examined as paired samples. The quantitative real-time PCR results showed that miR-100 had a negative effect on the expression of RNF144B in the samples (Figure 6k), indicating that miR-100 could target RNF144B in vivo. Western blotting yielded similar results (Figure 6k). Simultaneously, RNF144B negatively affected the expression of the E3 ubiquitin ligase pirh2, which led to p53 protein accumulation (Figure 6k). Taken together, miR-100 could directly trigger ubiquitin-mediated p53 protein degradation during gastric cancer apoptosis regulation in vivo.
The above data show that miR-100 has an essential role in the p53-mediated apoptosis of gastric cancer via the miR-100-RNF144B-pirh2-p53 pathway (Figure 7).
Discussion
The ability of the p53 protein to trigger cell apoptosis under environmental or oncogenic stress constitutes p53’s major function: to act as a tumour suppressor.29 The loss or gain of p53 function leads to aberrant cell growth. Therefore, both p53 gene expression and p53 protein activity are tightly regulated.30 In cells, the p53 protein has an extremely short half-life through ubiquitin-dependent proteolysis, leading to low p53 levels within cells.30 Ubiquitination provokes protein degradation and is a crucial cellular protein regulator. A protein is stable unless it is labelled with a polyubiquitin chain that marks it for degradation by the proteasome. During protein ubiquitination, E3 ubiquitin ligase critically determines substrate specificity and substrate selection.31 The ubiquitin-mediated p53 protein degradation mechanism has been extensively investigated.32 However, the role of miRNAs in the regulation of p53 protein ubiquitination is unclear. In this study, the findings reveal that miR-100 directly triggers ubiquitin-mediated p53 protein degradation during the regulation of gastric cancer apoptosis. Our study reveals a novel mechanism of p53 protein ubiquitination that is mediated by miRNA.
As reported, miR-100 is the oldest known animal miRNA33 and is widely expressed in vertebrates.34 The available findings regarding its expression profile in cancers are quite contradictory. Some investigations show that miR-100 expression is decreased in nasopharyngeal cancer,35 squamous cell carcinomas of the tongue,36 serous ovarian carcinomas,37 hepatoblastoma38 and oral cancer.39 However, miR-100 upregulation has been noted in medulloblastomas40 and in prostate41 and pancreatic42 cancers. These results indicate that the expression profile of miR-100 depends on the cancer type. To date, however, the role of miR-100 in tumourigenesis has not been extensively investigated.
Our results show that miR-100 is significantly upregulated in human gastric cancer cells. More importantly, our study reveals that miR-100 directly triggers ubiquitin-mediated p53 degradation during the regulation of gastric cancer apoptosis. The involvement of miR-100 in p53-mediated apoptosis occurs through the miR-100-RNF144B-pirh2-p53 pathway, in which miR-100 can target the RNF144B mRNA 3’-UTR. In addition, RNF144B gene depletion reportedly significantly impairs keratinocyte proliferation and differentiation by triggering p21 accumulation,43 suggesting its involvement in apoptosis regulation.
Some studies have posited that RNF144B can function as an IBR-type RING-finger E3 ubiquitin ligase that regulates Bax active form levels, inducing apoptosis in HeLa cells and in the colorectal cancer cell line HCT116.44
Our study reveals that RNF144B has an essential role in p53 protein ubiquitination in poorly differentiated gastric cancer cells, showing a novel aspect of RNF144B in cancers. In this investigation, RNF144B directly mediated pirh2 ubiquitin degradation, a p53 protein E3 ligase. As reported, pirh2 can promote p53 protein degradation in the mouse cell line BaF3/DD and the human cell line MCF-7.16 In this context, our study reveals a novel miRNA-mediated p53 protein ubiquitination pathway in cancer.
Human cancer is a complex genetic disease, and effective therapies for this condition have not yet been realized. DNA-damaging chemotherapies and radiotherapy have been the core therapeutic options for cancer for the past half-century. However, these treatments have adverse side effects. In this study, we show that miR-100 antagonism specifically induces gastric cancer cell apoptosis but not the corresponding non-cancerous cells, suggesting that the inhibition of miR-100 might represent a potential therapeutic intervention strategy for the treatment of refractory gastric cancer. Moreover, the differential expression of miR-100 in cancerous versus non-cancerous tissues that was observed in this study suggests that miR-100 might serve as a potential biomarker for gastric cancer.
Materials and methods
Cell culture
We purchased cancer cell lines (MGC-803, HGC-27 and MKN-45) from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). We purchased the gastric epithelial cell line (GES-1) from the Cell Bank of Beijing Institute for Cancer Research (Beijing, China). As reported, the MKN-45, HGC-27 and MGC-803 cell lines were poorly differentiated gastric cancer cells. The cell lines (MGC-803, HGC-27) were grown and maintained in 10% fetal bovine serum (Gibco, Grand Island, NY, USA) in RPMI 1640 (Gibco) at 37 °C in a humidified atmosphere of 5% CO2. The HGC-27 cell line was grown and maintained in 10% fetal bovine serum (Gibco) in Modified Eagle Medium and Dulbecco's Modified Eagle Medium at 37 °C in a humidified atmosphere of 5% CO2. All cell lines were profiled routinely through short-tandem repeat analysis.
Quantification of miR-100 with real-time PCR
We used TaqMan miRNA assays (Applied Biosystems, Carlsbad, CA, USA) to quantify mature miRNAs according to the supplier’s instructions, with some modifications. Total RNAs were isolated using an Ambion (Carlsbad, CA, USA) total RNA isolation kit (Ambion). Reverse transcription (RT) consisted of 100 ng of total RNAs, 50 nM stem-loop RT primer, 1 × RT buffer, dNTPs (each at 1 mM), 16.5 units of MultiScribe reverse transcriptase, and 1.36 units of RNase inhibitor (Applied Biosystems). The mixture was incubated at 16 °C for 30 min, 42 °C for 30 min and 85 °C for 5 min. The real-time PCR reaction mixture (10 μl) contained 0.5 μl of RT product, 5 μl of TaqMan 2 × Universal PCR Master Mix (Applied Biosystems), and 1 μl of TaqMan miRNA Assay (Applied Biosystems). The reactions were conducted at 95 °C for 10 min, followed by 50 cycles at 95 °C for 15 s and 60 °C for 1 min. We used the 2-(ΔΔCt) method to determine the relative individual miRNA quantities, and U6 (Applied Biosystems) was used as an internal standard for normalisation.
Pathology detection of gastric cancer patients
The gastric cancer patient tumours were examined by gastroscopy before chemotherapy administration. Then, normal and cancerous tissues were resected from gastric cancer patients and investigated by pathology. The normal and cancerous tissue samples from patients were placed in 20 ml of sterile 10% formalin in a universal container for transportation to the pathology department. Once fixed with buffered 10% formalin, the samples were dehydrated, embedded in paraffin wax and cut into 4 mm-thick sections. The sections were then stained with haematoxylin and eosin, followed by examination with white light microscopy and polarized light microscopy.
Gastric sample preparation
The cancerous tissues and normal gastric tissues near the cancerous region in gastric cancer patients were resected, immediately incubated in the RNAlater solution (Ambion) overnight and stored at −80 °C. Informed consent from patients and institutional review board approvals were obtained. We then extracted RNA from the tissues for quantitative real-time PCR. Proteins were also extracted and subjected to western blotting.
Transfection of an AMO and miRNA precursor in cells
The AMO and miRNA precursor were transfected into cells with Lipofectamine 2000 according to the manufacturer’s procedure (Life Technology, Carlsbad, CA, USA). Prior to transfection, the AMO (AMO-miR-100 or AMO-miR-100-scrambled) or miRNA precursor (miR-100 precursor or negative control) and Lipofectamine 2000 were mixed in Opti-MEM I and then incubated for 20 min at room temperature. The mixture was then added to the cells. Six hours later, fresh medium replaced the spent culture medium, and the cells were cultured. The AMO-miR-100 sequence (5′-TTCGGATCTACGGGTT-3′) was modified with locked nucleic acid (bold), 2’-O-methyl (italic) and phosphorothioate (the remaining nucleotides). The AMO-miR-100 sequence was randomly scrambled, generating AMO-miR-100-scrambled (5′-GTCGGTTCTGATGTCA-3′) with the same modifications as defined above. AMO-miR-100 and AMO-miR-100-scrambled were synthesized by Sangon Biotech Co., Ltd (Shanghai, China). The miR-100 precursor and negative control were purchased from Applied Biosystems.
Cell viability and apoptosis detection with the ApoTox-Glo triplex assay
We used the ApoTox-Glo Triplex Assay (Promega, Madison, WI, USA) to detect cell viability and apoptosis simultaneously and in the same cells according to the manufacturer’s protocols. We added 20 μl of the Viability/Cytotoxicity Reagent containing GF-AFC substrate to a final volume of 100 μl. After briefly mixing by orbital shaking at 300 g for 5 min, the mixture was incubated for 30 min at 37 °C in a humidified atmosphere of 5% CO2. We then measured fluorescence at 400 Ex/505 Em for cell viability. After measuring cell viability, we mixed 120 μl of the reaction mixture with 100 μl of Caspase-Glo 3/7 Reagent. The mixture was incubated by orbital shaking at 200 g for 1 h at room temperature. Caspase 3/7 activity was examined by luminescence to determine apoptosis.
Detection of caspase 3/7 activity
We used the Caspase-Glo 3/7 assay (Promega) to measure caspase 3 and caspase-7 activities according to the manufacturer’s instructions and mounted the cells onto a 96-well plate. The Caspase-Glo 3/7 buffer was transferred to a bottle containing the Caspase-Glo 3/7 substrate to form the Caspase-Glo 3/7 reagent. Then, 100 μl of the Caspase-Glo 3/7 reagent was added to each well of the 96-well plate. After gently mixing at 300–500 × g for 30 s, the mixture was incubated at room temperature for 2 h and underwent luminescence in a plate-reading luminometer.
Apoptosis detection with annexin V
Apoptosis assay with annexin V (Molecular Probes, Carlsbad, CA, USA) was conducted according to the manufacturer’s protocol. Cells were harvested and washed in cold phosphate-buffered saline. The cells were then resuspended in 1 × annexin-binding buffer at ~1 × 106 cells/ml, followed by the addition of 5 μl of Alexa Fluor 488 annexin V and 1 μl of 100 μg/ml PI (propidium iodide). The sample was incubated at room temperature for 15 min in the dark. Then, 400 μl of 1 × annexin-binding buffer was added to the sample and measured by flow cytometry at 530 nm and 575 nm using an excitation wavelength of 488 nm.
Cell proliferation assay
Cell proliferation was determined via an MTS [3-(4, 5-dimethylthiazol-2-yl)-5 -(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium, inner salt] assay (Promega). We added 20 μl of CellTiter 96 Aqueous One Solution Reagent into each well of the 96-well plate containing cells in 100 μl of culture medium. The plate was then incubated at 37 °C for 2 h in a humidified 5% CO2 incubator. The sample absorbance data at 450 nm were recorded using a 96-well plate reader.
Western blotting
Cells were collected by centrifugation at 300 g for 10 min and completely lysed by radio immunoprecipitation assay buffer (Beyotime, Shanghai, China) containing 2 mM phenylmethylsulfonyl fluoride. The sample was subjected to sodium dodecyl sulphate–polyacrylamide gel electrophoresis, followed by electrophoresis for 45 min at 200 V. The proteins were then electro-transferred to a polyvinylidene fluoride membrane (Millipore, Boston, MA, USA). The membrane was washed with tris buffered saline with tween (TBST) (Sangon, Shanghai, China) buffer for 5 min and then incubated with blocking buffer (4% bovine serum albumin in tris buffered saline with tween) for 1 h at room temperature. The membrane was then incubated with a primary antibody overnight at 4 °C. After three tris buffered saline with tween washes, the membrane was incubated with AP-conjugated secondary antibody (Roche, Basel, Switzerland) for 2 h at room temperature. After the tris buffered saline with tween washes, the membrane was incubated with BCIP/NBT substrate (Sangon) until the blot was visualized. The following antibodies were purchased from (Cell Signaling Technology, Boston, MA, USA): cleaved PARP (catalogue no. 5625); cleaved caspase 3 (catalogue no. 9661); caspase 3 (catalogue no. 9662); ki-67 (catalogue no. 9449); p53 (catalogue no. 9282); p21 (catalogue no. 2947); PUMA (catalogue no. 12450); bid (catalogue no. 8762); bax (catalogue no. 5023); β-tubulin (catalogue no. 2146); pirh2 (catalogue no. 5754); bcl-2 (catalogue no. 15071); RNF144B (catalogue no. sc-33017); flag (catalogue no. 14793) and his (catalogue no. 9991).
Tumourigenicity in nude mice
MGC-803 or HGC-27 cells were transfected with AMO-miR-100 or AMO-miR-100- scrambled at a final concentration of 50 nM when the cell density reached 50%. Six hours after transfection, the cells were collected and adjusted to 5 × 106 cells/ml with physiological saline. The cell suspension at 100 μl/mouse was then subcutaneously injected into seven female non-obese diabetic severe combined immunodeficient mice (4 weeks old) to induce tumour growth. The mark was hidden to exclude artificial factors during the experiments. The tumour volumes were measured every 3 days. One month later, the severe combined immunodeficient nude mice were killed, and their tumour weights were determined.
To evaluate the effect of miR-100 on solid tumours, we collected MGC-803 or HGC-27 cells and adjusted them to 5 × 106 cells/ml with physiological saline. We then subcutaneously injected a 100 μl cell suspension into nude mouse to induce tumour growth. Ten mice were used. One week later, when the tumour volume reached at ~30 mm3, the mice were subcutaneously and intravenously injected with AMO-miR-100 or AMO-miR-100-scrambled at 80 mg/kg once every 3 days. We measured the tumour volume every 3 days. Four weeks after the first injection, the mice were killed, and their tumours were weighed. Animal experiments were approved by The Animal Experiment Centre of Zhejiang University, China.
Integration of the luciferase gene into MGC-803 cells and the detection of apoptosis by in vivo imaging
The luciferase gene was cloned into the LV plasmid according to the manufacturer’s instructions (SiDanSai Stem Cell Technology Co., Ltd., Shanghai, China) to generate the LV-EF1α-eGFP-TRE-luciferase construct. Then, this construct and LP1, LP2 and VSV plasmids (SiDanSai Stem Cell Technology Co., Ltd.) were co-transfected into 293T cells for the virus package containing the luciferase gene in its genome. At 6 h after transfection, the medium was replaced with fresh medium, and the cells were cultured for 48 h. After a brief cell suspension centrifugation, the supernatant was collected and filtered through a 0.45-μm filter. Subsequently, the filtered supernatant was mixed with polybrene and added to MGC-803 cells. Six hours later, the medium was replaced with fresh medium, and the cells were cultured for 48 h. We extracted the virus-infected cell DNA and detected EF1α-eGFP-TRE-luciferase fragment integration into cell genomes using gene-specific primers (luciferase, 5′-ATGGAAGACGCCAAAAACATAAAGA-3′ and 5′-TTACAATTTGGACTTTCCGCCCTTC-3′; TRE (tetracycline response elements), 5′-CTCGAGTTTACCACTCCCTATCAGT-3′ and 5′-AGGCTGGATCGGTCCCGGTGTCTTC-3′). We then collected the MGC-803 cells containing EF1α-eGFP-TRE-luciferase and implanted them into nude mice to induce tumour growth.
One week later, the mice were subcutaneously and intravenously injected with AMO- miR-100 or AMO-miR-100-scrambled at 80 mg/kg once every 3 days. One month later, the mice were intraperitoneally injected with 100 μl of 20% urethane to anaesthetize them. Then, 100 μl of luciferase substrate [Z-Asp (O-Me)-Glu (O-Me)-Val-Asp (O-Me) aminoluciferin, sodium salt] at 65 mg/ml was intraperitoneally injected into the mice. Fifteen minutes later, the mice were examined using a molecular imaging system (NightOWL II LB 983, Berthold Technologies, Stuttgart, Germany) to detect luminescence. After in vivo luminescent imaging, the mice were killed, and the tumours were assessed by immunohistological analysis.
Immunohistology
Tumour tissue was incubated in 10% buffered formalin (pH 7.4) for up to 24 h and then cut into 3–5-μm pieces and mounted on slides. The slides were dewaxed and hydrated in 100%, 95% and 80% ethanol for 5 min each. Then, the primary antibody was incubated with slides in a humidity chamber. After washes with phosphate-buffered saline, the slides were covered with 4–5 drops of secondary antibody and incubated for 10 min at room temperature. Streptavidin peroxidase was added, followed by incubation for 10 min at room temperature. AEC buffer and AEC chromogen (Santa Cruz Biotechnology, Santa Cruz, CA, USA) were mixed and added to the slides. After incubating for 10 min at room temperature, the slides were examined under a fluorescence microscope. Human studies were approved by the local Institutional Review Board.
Detection of ubiquitinated protein
We detected the ubiquitination level of proteins using an UbiQapture-Q Kit (Enzo, Geneva, Switzerland) according to the manufacturer’s instructions. Cells were collected and lysed with radio immunoprecipitation assay lysis buffer (Beyotime). We then added 40 μl of UbiQapture-Q matrix (Beyotime) to the cell lysate. The sample was resuspended gently by inversion at 4 °C overnight to allow the ubiquitinated protein conjugates to bind to the affinity matrix. After centrifugation at 5,000 × g for 15–30 s, the matrix was washed twice in phosphate-buffered saline. The ubiquitin protein conjugates were then eluted with phosphate-buffered saline and subjected to western blotting.
Expression silencing of E3 ubiquitin ligase genes and the p53 gene
To knockdown the expression of E3 ubiquitin ligase genes in cells, the pirh2-specific siRNA (5′-AAUGUAACUUAUGCCUAGCUAUU-3′), the COP1-specific siRNA (5′-AACUGACCAAGAUAACCUUGAUU-3′), and the MDM2-specific siRNA (5′-GCCUGGCUCUGUGUGUAAUUU-3′) were transfected into cells. At 24 h after transfection, the cells were collected for subsequent assays.
To knockdown the expression level of p53 in miR-100-silenced cells, the p53-siRNA (5′-GAAGAAAAUUUCCGCAAAAUU-3′) and AMO-miR-100 were simultaneously transfected into cells, followed by culture for 24 h. Then, the cells were collected for later use.
Overexpression of the pirh2 gene in cells
To overexpress the pirh2 gene, the gene was cloned into pEASY-M1 (TransGen Biotech, Beijing, China) via PCR using the pirh2-specific primers 5′-GCCACCATGGCGGCGACGGCCCGGGAAGATG-3′ and 5′-TCATCCAGCTGTCTCCAATACCTGGTC-3′. The plasmid was confirmed by sequencing and then transfected into cells using the Attractene transfection reagent (Qiagen, Dusseldor, Germany). Thirty-six hours later, the treated cells were harvested for subsequent analysis.
Target gene prediction of miR-100
We predicted the miR-100 target genes using the TargetScan, miRanda, PicTar and miRInspector algorithms. The prediction results were ranked based on targeting efficacy as calculated using the sites' contexts+scores.
Dual-luciferase reporter assay
We performed a dual-luciferase reporter assay (Promega) to verify the interaction between miR-100 and its target gene, RNF144B. We cloned the RNF144B 3’UTR (5′-CATATAAAATACGGGTAGAGTGGTAGAG-3′) or RNF144B 3’-UTR mutant (5′-CATATAAAAGCTATACAGAGTGGTAGAG-3′) into the pmirGLO Dual-Luciferase vector (Promega), generating the RNF144B 3’-UTR and RNF144B 3’-UTR mutant constructs. Simultaneously, miR-100 (5′-AACCCGUAGAUCCGAACUUGUG-3′) was cloned into the miExpress vector (GeneCopoeia, Rockville, MD, USA). MGC-803 and MKN-45 cells were co-transfected with miExpress-miR-100 and the RNF144B 3’UTR or RNF144B 3’-UTR mutant. At 48 h after transfection, the firefly luciferase and Renilla luciferase activities were detected according to the manufacturer's protocol (Promega).
Silencing, overexpression and rescue of RNF144B gene expression
To knockdown RNF144B expression, the RNF144B-specific siRNA (5′-UCCAGAACUUGGAUAAUGACAUUUU-3′) (RNF144B-siRNA) was transfected into cells using Lipofectamine RNAiMAX reagent (Life Technology). The cells were collected 24 h after transfection. To overexpress the RNF144B gene expression, the gene was cloned into pEASY-M1 (TransGen Biotech) using the RNF144B-specific primers 5′-GCCACCATGGGCTCAGCTGGTAGGCTCCACT-3′ and 5′-TTAGGTTGTGGATGGGTCGTGCTTT-3′. To avoid recognition by RNF144B-siRNA, the RNF144B gene sequence was synonymously mutated at two nucleotides, generating the ΔRNF144B construct. The plasmid was then transfected into cells, which were collected at 24 h after infection for later use. To rescue RNF144B expression, the cells were transfected with RNF144B-siRNA to silence RNF144B gene expression. At 24 h after transfection, the ΔRNF144B plasmid was transfected into the RNF144B-silenced cells, which were harvested after culturing for 24 h.
Co-immunoprecipitation
Cells were collected and lysed with ice-cold cell lysis buffer (Beyotime). The lysate was incubated with Protein G-agarose beads (Invitrogen, Carlsbad, CA, USA) and then incubated with RNF144B antibody (Santa Cruz Biotechnology) overnight. After being washed three times with ice-cold lysis buffer, the immuno-complexes were collected and subjected to western blotting.
Statistical analyses
All biological experiments were repeated three times. Numerical data were analysed using a one-way analysis of variation. The statistical significance between treatments was assessed by one-tailed Student's t-tests or Paired Wilcoxon tests.
References
Taylor RC, Cullen SP, Martin SJ . Apoptosis: controlled demolition at the cellular level. Nat Rev Mol Cell Bio 2008; 9: 231–241.
Medina-Ramirez CM, Goswami S, Smirnova T, Bamira D, Benson B, Ferrick N et al. Apoptosis inhibitor ARC promotes breast tumorigenesis, metastasis, and chemoresistance. Cancer Res 2011; 71: 7705–7715.
Zuckerman V, Wolyniec K, Sionov RV, Haupt S, Haupt Y . Tumour suppression by p53: the importance of apoptosis and cellular senescence. J Pathol 2009; 219: 3–15.
Deng Y, Chan SS, Chang S . Telomere dysfunction and tumour suppression: the senescence connection. Nat Rev Cancer 2008; 8: 450–458.
Mercer J, Mahmoudi M, Bennett M . DNA damage, p53, apoptosis and vascular disease. Mutat Res 2007; 621: 75–86.
Donehower LA, Lozano G . 20 years studying p53 functions in genetically engineered mice. Nat Rev Cancer 2009; 9: 831–841.
Feki A, Irminger-Finger I . Mutational spectrum of p53 mutations in primary breast and ovarian tumors. Crit Rev Oncol Hematol 2004; 52: 103–116.
Le MT, Teh C, Shyh-Chang N, Xie H, Zhou B, Korzh V et al. MicroRNA-125b is a novel negative regulator of p53. Gene Dev 2009; 23: 862–876.
Wang X . p53 regulation. Cell Cycle 2011; 10: 4225–4229.
Ravid T, Hochstrasser M . Diversity of degradation signals in the ubiquitin–proteasome system. Nat Rev Mol Cell Bio 2008; 9: 679–689.
Shang F, Taylor A . Ubiquitin–proteasome pathway and cellular responses to oxidative stress. Free Radical Bio Med 2011; 51: 5–16.
Adams J . The proteasome: structure, function, and role in the cell. Cancer Treat Rev 2003; 29: 3–9.
Bernassola F, Karin M, Ciechanover A, Melino G . The HECT family of E3 ubiquitin ligases: multiple players in cancer development. Cancer Cell 2008; 14: 10–21.
Motegi A, Murakawa Y, Takeda S . The vital link between the ubiquitin–proteasome pathway and DNA repair: Impact on cancer therapy. Cancer Lett 2009; 283: 1–9.
Allende-Vega N, Saville MK . Targeting the ubiquitin–proteasome system to activate wild-type p53 for cancer therapy. Semin Cancer Biol 2010; 20: 29–39.
Leng RP, Lin Y, Ma W, Wu H, Lemmers B, Chung S et al. Pirh2, a p53-induced ubiquitin-protein ligase, promotes p53 degradation. Cell 2003; 112: 779–791.
Laine A, Ronai ZE . Regulation of p53 localization and transcription by the HECT domain E3 ligase WWP1. Oncogene 2006; 26: 1477–1483.
Liu Z, Miao D, Xia Q, Hermo L, Wing SS . Regulated expression of the ubiquitin protein ligase, E3Histone/LASU1/Mule/ARF‐BP1/HUWE1, during spermatogenesis. Dev Dyn 2007; 236: 2889–2898.
Dornan D, Wertz I, Shimizu H, Arnott D, Frantz GD, Dowd P et al. The ubiquitin ligase COP1 is a critical negative regulator of p53. Nature 2004; 429: 86–92.
Tsvetkov P, Adamovich Y, Elliott E, Shaul Y . E3 ligase STUB1/CHIP regulates NAD (P) H: quinone oxidoreductase 1 (NQO1) accumulation in aged brain, a process impaired in certain Alzheimer disease patients. J Biol Chem 2011; 286: 8839–8845.
Ambros V . The functions of animal microRNAs. Nature 2004; 431: 350–355.
Bartel DP, Chen C-Z . Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat Rev Genet 2004; 5: 396–400.
Esteller M . Non-coding RNAs in human disease. Nat Rev Genet 2011; 12: 861–874.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005; 120: 15–20.
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 2004; 101: 2999–3004.
Calin GA, Croce CM . MicroRNA signatures in human cancers. Nat Rev Cancer 2006; 6: 857–866.
Yang G, Yang L, Zhao Z, Wang J, Zhang X . Signature miRNAs involved in the innate immunity of invertebrates. PLoS One 2012; 7: e39015.
Gong Y, He T, Yang L, Yang G, Chen Y, Zhang X . The role of miR-100 in regulating apoptosis of breast cancer cells. Sci Rep 2015; 5: e11650.
Pietsch EC, Sykes SM, McMahon SB, Murphy ME . The p53 family and programmed cell death. Oncogene 2008; 27: 6507–6521.
Barboza JA, Iwakuma T, Terzian T, El-Naggar AK, Lozano G . Mdm2 and Mdm4 loss regulates distinct p53 activities. Mol Cancer Res 2008; 6: 947–954.
Isaacson MK, Ploegh HL . Ubiquitination, ubiquitin-like modifiers, and deubiquitination in viral infection. Cell Host Microbe 2009; 5: 559–570.
Chen D, Zhang J, Li M, Rayburn ER, Wang H, Zhang R . RYBP stabilizes p53 by modulating MDM2. EMBO Rep 2009; 10: 166–172.
Christodoulou F, Raible F, Tomer R, Simakov O, Trachana K, Klaus S et al. Ancient animal microRNAs and the evolution of tissue identity. Nature 2010; 463: 1084–1088.
Wienholds E, Kloosterman WP, Miska E, Alvarez-Saavedra E, Berezikov E, de Bruijn E et al. MicroRNA expression in zebrafish embryonic development. Science 2005; 309: 310–311.
Shi W, Alajez NM, Bastianutto C, Hui AB, Mocanu JD, Ito E et al. Significance of Plk1 regulation by miR‐100 in human nasopharyngeal cancer. Int J Cancer 2010; 126: 2036–2048.
Wong T-S, Liu X-B, BY-H Wong, RW-M Ng, AP-W Yuen, Wei WI . Mature miR-184 as potential oncogenic microRNA of squamous cell carcinoma of tongue. Clin Cancer Res 2008; 14: 2588–2592.
Nam EJ, Yoon H, Kim SW, Kim H, Kim YT, Kim JH et al. MicroRNA expression profiles in serous ovarian carcinoma. Clin Cancer Res 2008; 14: 2690–2695.
Cairo S, Wang Y, de Reyniès A, Duroure K, Dahan J, Redon M-J et al. Stem cell-like micro-RNA signature driven by Myc in aggressive liver cancer. Proc Natl Acad Sci USA 2010; 107: 20471–20476.
Henson BJ, Bhattacharjee S, O'Dee DM, Feingold E, Gollin SM . Decreased expression of miR‐125b and miR‐100 in oral cancer cells contributes to malignancy. Gene Chromosome Cancer 2009; 48: 569–582.
Liu W, Gong Y-H, Chao T-F, Peng X-Z, Yuan J-G, Ma Z-Y et al. Identification of differentially expressed microRNAs by microarray: a possible role for microRNAs gene in medulloblastomas. Chinese Med J 2009; 122: 2405–2411.
Leite KR, Sousa-Canavez JM, Reis ST, Tomiyama AH, Camara-Lopes LH, Sañudo A et al. Change in expression of miR-let7c, miR-100, and miR-218 from high grade localized prostate cancer to metastasis. Urol Oncol 2011; 29: 265–269.
Lee EJ, Gusev Y, Jiang J, Nuovo GJ, Lerner MR, Frankel WL et al. Expression profiling identifies microRNA signature in pancreatic cancer. Int J Cancer 2007; 120: 1046–1054.
Conforti F, Yang AL, Piro MC, Mellone M, Terrinoni A, Candi E et al. PIR2/Rnf144B regulates epithelial homeostasis by mediating degradation of p21WAF1 and p63. Oncogene 2012; 32: 4758–4765.
Benard G, Neutzner A, Peng G, Wang C, Livak F, Youle RJ et al. IBRDC2, an IBR‐type E3 ubiquitin ligase, is a regulatory factor for Bax and apoptosis activation. EMBO J 2010; 29: 1458–1471.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (41276152, 31430089) and the National Program on the Key Basic Research Project (2015CB755903).
Author contributions
GY, YG, QZW and LPW performed the experiments. XBZ, GY and YG designed the experiments and analysed the data. YG and XBZ wrote the manuscript. All authors read and approved the contents of the manuscript and its publication.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflicts of interest.
Rights and permissions
About this article
Cite this article
Yang, G., Gong, Y., Wang, Q. et al. miR-100 antagonism triggers apoptosis by inhibiting ubiquitination-mediated p53 degradation. Oncogene 36, 1023–1037 (2017). https://doi.org/10.1038/onc.2016.270
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.270
This article is cited by
-
Regulating tumor suppressor genes: post-translational modifications
Signal Transduction and Targeted Therapy (2020)
-
miR-100-3p inhibits cell proliferation and induces apoptosis in human gastric cancer through targeting to BMPR2
Cancer Cell International (2019)
-
Tumor-recruited M2 macrophages promote gastric and breast cancer metastasis via M2 macrophage-secreted CHI3L1 protein
Journal of Hematology & Oncology (2017)