Abstract
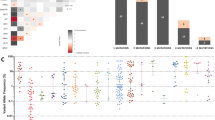
The implementation of next-generation sequencing (NGS) has influenced diagnostic, prognostic, and therapeutic decisions in myeloid malignancies. However, the clinical relevance of serial molecular annotation in patients with myelodysplastic syndrome (MDS) undergoing active treatment is unknown. MDS or secondary acute myeloid leukemia (sAML) patients who had at least two NGS assessments were identified. Outcomes according to mutation clearance (NGS-) on serial assessment were investigated. Univariate and multivariate Cox regression models were used to evaluate the prognostic impact of NGS trajectory on overall survival (OS). A total of 157 patients (MDS [n = 95]; sAML [n = 52]; CMML [n = 10]) were identified, with 93% of patients receiving treatment between NGS assessments. Magnitude of VAF delta from baseline was significantly associated with quality of response to treatment. Patients achieving NGS- had significantly improved OS compared to patients with mutation persistence (median OS not reached vs. 18.5 months; P = 0.002), which was confirmed in multivariate analysis (HR,0.14; 95%CI = 0.03–0.56; P = 0.0064). Serial TP53 VAF evaluation predicts outcomes with TP53 clearance representing an independent covariate for superior OS (HR,0.22; 95%CI = 0.05–0.99; P = 0.048). Collectively, our study highlights the clinical value of serial NGS during treatment and warrants prospective validation of NGS negativity as a biomarker for treatment outcome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Tefferi A, Vardiman JW. Myelodysplastic syndromes. N Engl J Med. 2009;361:1872–85.
Löwenberg B, Downing JR, Burnett A. Acute myeloid leukemia. N Engl J Med. 1999;341:1051–62.
Kayser S, Döhner K, Krauter J, Köhne CH, Horst HA, Held G, et al. The impact of therapy-related acute myeloid leukemia (AML) on outcome in 2853 adult patients with newly diagnosed AML. Blood. 2011;117:2137–45.
Hulegårdh E, Nilsson C, Lazarevic V, Garelius H, Antunovic P, Rangert Derolf Å, et al. Characterization and prognostic features of secondary acute myeloid leukemia in a population-based setting: a report from the Swedish Acute Leukemia Registry. Am J Hematol. 2015;90:208–14.
Renaud L, Nibourel O, Berthon C, Roumier C, Rodriguez C, Frimat C, et al. De Novo and secondary acute myeloid leukemia, real world data on outcomes from the french nord-pas-de-calais picardie acute myeloid leukemia observatory. Blood. 2016;128:4013.
Döhner H, Estey E, Grimwade D, Amadori S, Appelbaum FR, Büchner T, et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood. 2017;129:424–47.
Bejar R, Stevenson K, Abdel-Wahab O, Galili N, Nilsson B, Garcia-Manero G, et al. Clinical effect of point mutations in myelodysplastic syndromes. N Engl J Med. 2011;364:2496–506.
Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P, et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood. 2013;122:3616–27. quiz 3699.
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND, et al. Genomic classification and prognosis in acute myeloid leukemia. N Engl J Med. 2016;374:2209–21.
Bally C, Ades L, Renneville A, Sebert M, Eclache V, Preudhomme C, et al. Prognostic value of TP53 gene mutations in myelodysplastic syndromes and acute myeloid leukemia treated with azacitidine. Leuk Res. 2014;38:751–5.
Bejar R, Lord A, Stevenson K, Bar-Natan M, Perez-Ladaga A, Zaneveld J, et al. TET2 mutations predict response to hypomethylating agents in myelodysplastic syndrome patients. Blood. 2014;124:2705–12.
Bejar R, Stevenson KE, Caughey B, Lindsley RC, Mar BG, Stojanov P, et al. Somatic mutations predict poor outcome in patients with myelodysplastic syndrome after hematopoietic stem-cell transplantation. J Clin Oncol. 2014;32:2691–8.
Della Porta MG, Galli A, Bacigalupo A, Zibellini S, Bernardi M, Rizzo E, et al. Clinical effects of driver somatic mutations on the outcomes of patients with myelodysplastic syndromes treated with allogeneic hematopoietic stem-cell transplantation. J Clin Oncol. 2016;34:3627–37.
Lindsley RC, Saber W, Mar BG, Redd R, Wang T, Haagenson MD, et al. Prognostic mutations in myelodysplastic syndrome after stem-cell transplantation. N Engl J Med. 2017;376:536–47.
Yoshizato T, Nannya Y, Atsuta Y, Shiozawa Y, Iijima-Yamashita Y, Yoshida K, et al. Genetic abnormalities in myelodysplasia and secondary acute myeloid leukemia: impact on outcome of stem cell transplantation. Blood. 2017;129:2347–58.
Takahashi K, Patel K, Bueso-Ramos C, Zhang J, Gumbs C, Jabbour E, et al. Clinical implications of TP53 mutations in myelodysplastic syndromes treated with hypomethylating agents. Oncotarget. 2016;7:14172–87.
Sallman DA, Komrokji R, Vaupel C, Cluzeau T, Geyer SM, McGraw KL, et al. Impact of TP53 mutation variant allele frequency on phenotype and outcomes in myelodysplastic syndromes. Leukemia. 2016;30:666–73.
Bejar R, Papaemmanuil E, Haferlach T, Garcia-Manero G, Maciejewski JP, Sekeres MA, et al. TP53 mutation status divides MDS patients with complex karyotypes into distinct prognostic risk groups: analysis of combined datasets from the international working group for MDS-molecular prognosis committee. Blood. 2014;124:532.
Belickova M, Vesela J, Jonasova A, Pejsova B, Votavova H, Merkerova MD, et al. TP53 mutation variant allele frequency is a potential predictor for clinical outcome of patients with lower-risk myelodysplastic syndromes. Oncotarget. 2016;7:36266–79.
Makishima H, Yoshizato T, Yoshida K, Sekeres MA, Radivoyevitch T, Suzuki H, et al. Dynamics of clonal evolution in myelodysplastic syndromes. Nat Genet. 2017;49:204–12.
da Silva-Coelho P, Kroeze LI, Yoshida K, Koorenhof-Scheele TN, Knops R, van de Locht LT, et al. Clonal evolution myelodysplastic syndromes. Nat Commun. 2017;8:15099.
Uy GL, Duncavage EJ, Chang GS, Jacoby MA, Miller CA, Shao J, et al. Dynamic changes in the clonal structure of MDS and AML in response to epigenetic therapy. Leukemia. 2017;31:872–81.
Mossner M, Jann J-C, Wittig J, Nolte F, Fey S, Nowak V, et al. Mutational hierarchies in myelodysplastic syndromes dynamically adapt and evolve upon therapy response and failure. Blood. 2016;128:1246–59.
Pellagatti A, Roy S, Di Genua C, Burns A, McGraw K, Valletta S, et al. Targeted resequencing analysis of 31 genes commonly mutated in myeloid disorders in serial samples from myelodysplastic syndrome patients showing disease progression. Leukemia. 2016;30:247–50.
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127:2391–405.
Cheson BD, Bennett JM, Kantarjian H, Pinto A, Schiffer CA, Nimer SD, et al. Report of an international working group to standardize response criteria for myelodysplastic syndromes. Blood. 2000;96:3671–4.
Greenberg PL, Tuechler H, Schanz J, Sanz G, Garcia-Manero G, Solé F, et al. Revised international prognostic scoring system for myelodysplastic syndromes. Blood. 2012;120:2454–65.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–23.
Komrokji RS, DeZern AE, Zell K, Al Ali NH, Estling C, Zimmerman C, et al. Validation of International Working Group (IWG) Response Criteria in Higher-Risk Myelodysplastic Syndromes (MDS): A Report on Behalf of the MDS Clinical Research Consortium (MDS CRC). Blood. 2015;126:909.
Duncavage EJ, Jacoby MA, Chang GS, Miller CA, Edwin N, Shao J. et al. Mutation Clearance after Transplantation for Myelodysplastic Syndrome. N Engl J Med. 2018;379:1028–41.
Ganzel C, Sun Z, Gonen M, Patel JP, Abdel-Wahab O, Fernandez HF, et al. Minimal residual disease assessment by flow cytometry in AML Is an independant prognostic factor even after adjusting for cytogenetic/molecular abnormalities. Blood. 2014;124:1016.
Minetto P, Guolo F, Clavio M, Kunkl A, Ballerini F, Colombo N, et al. Minimal residual disease assessment may drive post remission therapy in acute myeloid leukemia. It’s time for MRD-driven therapy. Blood. 2016;128:2895.
Sebert M, Vidal V, Eclache V, Thepot S, Braun T, Gardin C, et al. Impact cytogenetics cytogenetic response outcome MDS treat azacitidine (AZA). Blood. 2012;120:2807.
Ivey A, Hills RK, Simpson MA, Jovanovic JV, Gilkes A, Grech A, et al. Assessment of minimal residual disease in standard-risk AML. N. Engl J Med. 2016;374:422–33.
Balsat M, Renneville A, Thomas X, de Botton S, Caillot D, Marceau A, et al. Postinduction minimal residual disease predicts outcome and benefit from allogeneic stem cell transplantation in acute myeloid leukemia with npm1 mutation: a study by the acute leukemia french association group. J Clin Oncol. 2017;35:185–93.
Freeman SD, Hills RK, Virgo P, Khan N, Couzens S, Dillon R, et al. Measurable residual disease at induction redefines partial response in acute myeloid leukemia and stratifies outcomes in patients at standard risk without NPM1 mutations. J Clin Oncol. 2018;36:1486–97.
Jongen-Lavrencic M, Grob T, Hanekamp D, Kavelaars FG, al Hinai A, Zeilemaker A, et al. Molecular minimal residual disease in acute myeloid leukemia. N. Engl J Med. 2018;378:1189–99.
Morita K, Kantarjian HM, Wang F, Yan Y, Bueso-Ramos C, Sasaki K, et al. Clearance of somatic mutations at remission and the risk of relapse in acute myeloid leukemia. J Clin Oncol. 2018;36:1788–97.
Welch JS, Petti AA, Miller CA, Fronick CC, O’Laughlin M, Fulton RS, et al. TP53 and decitabine in acute myeloid leukemia and myelodysplastic syndromes. N Engl J Med. 2016;375:2023–36.
Sallman DA, DeZern AE, Garcia-Manero G, Steensma DP, Roboz GJ, et al. Phase 1b/2 combination study of APR-246 and azacitidine (AZA) in patients with TP53 mutant myelodysplastic syndromes (MDS) and acute myeloid. Leuk (AML). Blood. 2019;132(Suppl 1):676.
Acknowledgements
This work was supported in part by a research grant from the Graduate Medical Education (GME) at the University of South Florida (SY), Research Training Award for Fellow (RTAF) from the American Society of Hematology (SY), Scholar Award from the American Society of Hematology (SY), NIH grant K08 CA237627 (SY), MDS Foundation Young Investigator Grant (DAS), Early Career Award of the Dresner Foundation (DAS), and Edward P. Evans Foundation Award (DAS).
Author information
Authors and Affiliations
Contributions
Conception and design: SY, DAS. Collection and assembly of data: SY, NHA, MH, KS, JL, JH, AL, EP, RK, DAS. Data analysis and interpretation: SY, SMG, JS, CV, DAS. Manuscript writing: SY, NHA, MH, KS, SMG, JL, JH, AL, EP, RK, DAS.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Yun, S., Geyer, S.M., Komrokji, R.S. et al. Prognostic significance of serial molecular annotation in myelodysplastic syndromes (MDS) and secondary acute myeloid leukemia (sAML). Leukemia 35, 1145–1155 (2021). https://doi.org/10.1038/s41375-020-0997-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-020-0997-4
This article is cited by
-
Vemurafenib induces senescence in acute myeloid leukemia and myelodysplastic syndrome by activating the HIPPO signaling pathway: implications for potential targeted therapy
Biology Direct (2024)
-
Comparing Arsenic-Containing Qinghuang Powder and Low-Intensity Chemotherapy in Elderly Patients with Acute Myeloid Leukemia
Chinese Journal of Integrative Medicine (2023)