Abstract
Primary malignant melanoma of the esophagus (PMME) is an exceedingly rare disease with a poor prognosis. The etiology of PMME remains largely unknown and genetic characteristics are yet to be clarified, essential for identifying potential therapeutic targets and defining treatment guidelines. Here, we performed whole-exome sequencing on 47 formalin-fixed paraffin-embedded specimens from 18 patients with PMME, including 23 tumor samples, 6 metastatic lymph nodes, and 18 tumor-adjacent normal tissues. The genomic features of PMME were comprehensively characterized, and comparative genomic analysis was further performed between these specimens and 398 skin cutaneous melanomas (SKCM), 67 non-esophagus mucosal melanomas (NEMM), and 79 uveal melanomas (UVM). In the PMME cohort, recurrently mutated driver genes, such as MUC16, RANBP2, NRAS, TP53, PTPRT, NF1, MUC4, KMT2C, and BRAF, were identified. All RANBP2 mutations were putatively deleterious, and most affected samples had multipoint mutations. Furthermore, RANBP2 showed parallel evolution by multiregional analysis. Whole-genome doubling was an early truncal event that occurred before most driver mutations, except for in TP53. An ultraviolet radiation-related mutational signature, SBS38, was identified as specific to epithelial melanomas and could predict inferior survival outcomes in both PMME and SKCM patients. Comparing the mutational and copy number landscapes between PMME and other subtypes of melanoma revealed that PMME has a similar genomic pattern and biological characteristics to SKCM. In summary, we comprehensively defined the key genomic aberrations and mutational processes driving PMME and suggested for the first time that PMME may share similar genomic patterns with SKCM; therefore, patients with rare melanomas, such as PMME, may benefit from the current treatment used for common cutaneous melanoma.
Similar content being viewed by others
Introduction
Primary malignant melanoma of the esophagus (PMME) is an extremely rare disease, accounting for 0.1% to 0.2% of all esophageal cancers and less than 0.05% of all melanomas1,2,3,4. PMME is classified as mucosal melanoma (MM), which, together with skin cutaneous melanoma (SKCM) and acral melanoma (AM), belongs to the epithelium-associated melanoma family5,6. Unlike other melanomas, PMME is typically detected at a more advanced stage and is characterized by a high recurrence and metastasis rate, with 5-year overall survival (OS) rate ranging from 4% to 37.5%3,7. The pathogenesis and treatment options of ultraviolet (UV)-exposed SKCM have been significantly advanced. The genetic architecture of non-esophagus MMs (NEMMs) have also been examined previously, which identified several driver genes and potential therapeutic targets8,9. In contrast, due to limited progress in the exploration of PMME molecular characteristics, understanding of its pathogenesis is still lacking, and the identification of druggable targets is challenging. Additionally, the comparative genomic characteristics between PMME and other epithelium-associated melanomas remain unclear.
In our prior case report of a patient with PMME, multiregional sequencing revealed high intratumor heterogeneity (ITH) in the PMME genome10. Furthermore, oncogenic driver mutations in genes such as BRAF and KRAS and CDKN2A biallelic inactivation were found to be clonal events, while clinically actionable mutations in genes such as PIK3CA and JAK1 were subclonal10. Recently, Tsuyama et al. characterized the genetic features of 10 PMMEs by using “TodaiOncoPanel (TOP)” sequencing4, which identified mutations in NF1, SF3B1, KRAS, BRCA2, KIT, and TP53. But BRAF mutations were not detected in the Japanese cohort. They suggested that the genetic profile of PMME was similar to that of mucosal/acral melanoma but differed from the SKMM profile. However, there remains a large gap in research into the tumorigenesis and metastasis mechanisms of PMME and their relevance to genomic alterations.
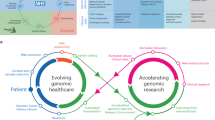
Here, we performed whole-exome sequencing (WES) on single or multiple lesions from a cohort of patients with PMME and presented a comparison of the genomic landscape of PMME to NEMM and SKCM (Fig. 1A). We comprehensively defined the key genomic aberrations and mutational processes driving PMME and compared the genomic patterns of this rare type of melanoma to those of other types of melanoma. Our results indicate that PMME has common melanoma driver genes and copy number alterations. We aimed to provide insights into potentially clinically relevant genomic aberrations and data resources for candidate therapeutic targets.
A Flow chart of the study design. B The mutation landscape of the PMME in this study. Each column represents an individual tumor or metastatic lymph node that underwent WES. Upper panel, the WGD status of each sample and clinical features, including age, sex, and risk factors (smoking and alcohol) of each patient. Middle, recurrently mutated driver genes identified in PMMEs. Bottom, the ID of patients with PMME.
Materials and methods
Sample collection
Eighteen treatment-naïve patients with PMME who underwent radical resection between January 2011 and February 2019 at Peking University Cancer Hospital (n = 15) and Nanjing Drum Tower Hospital (n = 3) were recruited for this study. Physical examination and computed tomography scans were performed on each patient and showed no evidence of skin melanoma or other MM lesions before surgery. Archived formalin-fixed paraffin-embedded (FFPE) samples of 23 tumors and 18 matched noncancerous adjacent normal tissues and 6 metastatic lymph nodes were obtained for WES. The data derived from one patient (HZY) had been previously published10 and were reanalyzed for the present study. This study was conducted according to the Declaration of Helsinki (revised in 2013) and was approved by the Internal Review Board of Peking University Cancer Hospital and the Nanjing Drum Tower Hospital. All patients provided written informed consent for the use of biological specimens for scientific research before undergoing surgery.
DNA extraction, library construction, and whole-exome sequencing
DNA was extracted from FFPE samples using the TIANamp Genomic DNA kit (Tiangen Biotech, Beijing, China) according to the manufacturer’s instructions. Primary tumor/lymph node samples and their paired normal tissues were fragmented with an ultrasonicator UCD-200 (Diagenode, Seraing, Belgium) and subsequently purified and size-selected with Ampure Beads (Beckman, MA, USA) following end-repairing, an “A” base addition, and adaptor ligation. The purity and concentration of DNA were determined using a Nanodrop 2000 spectrophotometer and Qubit 2.0 Fluorometer with the Quanti-IT dsDNA HS Assay Kit (Thermo Fisher Scientific, MA, USA). Some samples were prepared using the TruSeq Capture kit (Illumina, San Diego, CA, USA) for DNA library preparation. Whole-exome paired-end sequencing was performed on the Geneplus-2000 sequencing platform (Geneplus, Beijing, China) or Illumina HiSeq X10 (Illumina, San Diego, CA, USA) at Geneplus (Geneplus, Beijing, China) or Novegene (Novegene, Beijing, China).
Raw sequencing data were then filtered to remove low-quality reads and those containing adaptor sequences. Reads were further mapped to the reference human genome (hg19) using BWA aligner (version 0.7.10). An average target sequence depth of 150× of tumor or lymph node tissues, 100× of normal tissues, 100× coverage greater than 70%, or 50× coverage greater than 75% was adopted as the quality control for the BAM files.
WES single nucleotide variant (SNV)/insertion and deletion (indel) analysis
Somatic SNVs and small indels were called using MuTect2 packed in GATK (version 4.1.2.0). We applied the following criteria as filters to procure the reliable variants: (i) variants with allele frequency ≥0.01 were kept; (ii) mutations previously reported in public databases (1000 Genomes, gnomAD, and ExAC) with >0.001 allele frequency were removed; (iii) for patients with only one tumor sample available, variants with supported read number ≥5 were kept; (iv) for multiple samples from a single patient, if a variant was present in more than one sample, the supported read filter as mentioned in (ii) could be ignored; and (v) variants located in the coding region of the genome were defined as functional variants and kept for all analyses except for the signature decomposition.
Putative driver genes were curated by merging the Catalog of Somatic Mutations in Cancer (COSMIC) Cancer Gene Census (https://cancer.sanger.ac.uk/census) v90 and those reported by ref. 11 and ref. 12. The nonsense mutations, nonstop mutations, splice-site mutations, translation start sites, and in-frame/frame-shift small indels in the driver genes were classified as deleterious mutations. We identified putative deleterious missense mutations by using Sorting Intolerant from Tolerant (SIFT) and Polymorphism Phenotyping-2 (Polyphen2). In SIFT, the variants with a prediction score ranging from 0 to 0.05 were defined as deleterious. In Polyphen2, the variants annotated as “possibly damaging” or “probably damaging” were defined as deleterious. Missense mutations identified as deleterious by either of these two algorithms were classified as putative deleterious mutations. The number of somatic coding nonsynonymous variants (in-frame/frame-shift indels, missense mutations, nonsense mutations, nonstop mutations, splice-site mutations, and translation start sites) per megabase (Muts/Mb) of the examined genome was calculated as the tumor mutation burden (TMB).
Analysis of somatic copy number variation (sCNV)
A purity >0.2 was adopted as a filter for samples subjected to CNV analysis. The sCNV was identified by GISTIC2.0 to analyze the significantly altered copy number of the segments in samples, and a q value of 0.10 was set as the threshold of significance. Segmentation files obtained from GATK (version 4.1.2.0) were used as the inputs. For patients with multiple samples available in the PMME cohort, only one sample with the highest purity was retained to guarantee the accurate detection of frequency in the cohort. We estimated the purity of tumor cells and genome ploidy utilizing ABSOLUTE to determine the allele-specific DNA copy number and whole-genome doubling (WGD). We used the major (a2) and minor (a1) allele copy numbers to classify CNV events as duplication (a1 + a2 > 2 and a1 > 0), haploid loss of heterozygosity (LOH) (a2 = 1, a1 = 0), copy number neutral LOH (a2 = 2, a1 = 0), duplication LOH (a2 > 2, a1 = 0) and homogeneous deletion (a1 = a2 = 0). The identification of WGD was defined as when >50% of the whole genome was observed with major copy numbers of 2 or more. Additionally, the log2 ratio of segments was calculated to reflect the amplification and deletion with a threshold of 0.15 and subsequently used to gauge the CNV frequency of each cohort and to estimate the genome instability index (GII), defined as the proportion of the length of the genome with segmented copy number amplification (GIIAmp) or deletion (GIIDel). Sample level GII was calculated as the mean of multiple samples from each patient.
Inferring cancer cell fraction (CCF) and clonal status
CCFs of the mutations and CNVs were estimated using ABSOLUTE. In a single sample, mutations with a CCF higher than the 95% confidence interval ≥1.0 and a probability of clonal mutations greater than the probability of subclonal mutations were defined as clonal; otherwise, the mutations were defined as subclonal.
Reconstruction of the phylogenetic tree and the estimation of ITH
The presence or absence matrix of all nonsynonymous mutations detected in the samples was constructed for each patient. The matrices were then used as inputs for the phylogeny reconstruction by PYLIP13 with the maximum parsimony algorithm. The resulting phylogenetic trees were then manually plotted. Trunk events were defined as those shared by all the samples from the same patient, while private events were harbored by only one sample. ITH at the patient level was inferred based on the tree structure as follows: ITH = 1-#shared mutations/(#shared mutations + #private mutations).
Inference of the temporal order of variants and WGD
The temporal order of the occurrence of WGD and mutations or CNV during tumor progression was speculated using previously reported methods14,15. Briefly, subclonal events were considered post-WGD events. For clonal events, a copy number ≥1 was likely to be a pre-WGD event since it was amplified with the occurrence of WGD, whereas events with a single copy number occurred after WGD.
Analysis of the mutational signature
The Yet Another Package for Signature Analysis (YAPSA version 3.12) package (DOI: 10.18129/B9.bioc.YAPSA) was utilized to infer the trinucleotide mutational patterns and match to the signatures described in the COSMIC database to determine the possible mechanisms underlying the mutational processes and the reason for the potential carcinogen exposure.
Pathway enrichment analysis
Mutated driver genes in cohorts were compared with the Kyoto Encyclopedia of Genes and Genomes (KEGG) database to determine the altered pathways in tumors. The melanoma pathway (hsa05218) was identified, and the frequency of genes involved in this pathway was compared across different cohorts.
Published data obtained
The genetic data and clinical information of 398 SKCMs, 79 uveal melanomas (UVMs), and 67 NEMMs were collected from published databases and publications. MAF, segmentation files, and corresponding clinical data of the 398 SKCMs and 79 UVMs were downloaded from cBioPortal (http://www.cbioportal.org/). NEMM data were obtained from ref. 8, and this cohort did not include any esophageal tumors. All downloaded data were subjected to a uniform in-house filter pipeline to improve the unity of data from different origins and used for further analysis. In some analyses, the 18 PMME cases in this study were combined with the 67 published NEMM cases (termed the MM cohort).
Statistics
Continuous data were compared using the Kruskal–Wallis test or Mann–Whitney U test in GraphPad Prism (version 8.0). Spearman correlation analysis of the frequency of mutated driver genes in different cohorts was performed in Origin 2018. Overall survival (OS) time was defined as the time from the date of surgery to the date of death from any cause or the last follow-up. Relapse-free survival (RFS) time was defined as the time from the date of surgery to the date of recurrence. Disease-free survival (DFS) time was defined as the period from the date of surgery to disease recurrence, death, or the date of the last follow-up. Curves depicting the OS, RFS and DFS rates were plotted by the Kaplan–Meier method and compared through the log-rank test. Univariate and multivariate Cox proportional hazards regression models were also used to explore the potential independent prognostic factors by using R software 4.0.2. In the Cancer Genome Atlas (TCGA) SKCM cohort, OS and DFS outcomes were obtained from the clinical data. A series of cutoff points for continuous variables (SBS38, SBS6, SBS7a, SBS7b, and GIIAmp) were set within the range (the minimum value × 1.15 ~ to the maximum value × 0.85), and the optimal cutoff value was determined comprehensively considering hazard ratio (HR), P value, and the number of patients assigned to each group16. All the tests were two-sided, and a P value < 0.05 was considered statistically significant.
Results
Patients and samples
The PMME cohort included thirteen men and five women with a median age of 54.5 years (range: 37–76 years). The primary tumors were located in the middle or lower third of the esophagus in most patients (n = 16, 88.9%), while one patient had a tumor in the upper third, and one patient’s tumor was in the gastroesophageal junction. Eight patients had lymph node metastases. As of June 2021, thirteen patients had died, and five were still alive. Detailed information about each case is provided in Table S1.
The study involved WES of 47 FFPE specimens, including 23 tumor sites, 6 metastatic lymph nodes and 18 noncancerous adjacent normal tissues. Multiple tumor sites were obtained from three patients (LJP, HZY, and CXG), and metastatic lymph nodes were obtained from five patients (QYS, QXL, HZY, HXC, and SZY) (Fig. 1B). WES was covered by a median depth of 226× (range: 121–398) for tumors and lymph nodes and 267.5× (range: 163–413) for normal samples (Table S2).
Mutational and copy number landscape of PMME
A total of 3038 SNV/indels were identified across the tumor and lymph node samples. The average purity of the tumor and lymph node was estimated to be 0.62 (range: 0.20–0.87) according to ABSOLUTE, indicating that our sequencing data were robust for further analysis. Overall, the SNV/indel mutation burden (TMB) was low, with a median of 3.28 Muts/Mb (range: 0.5–9.8). The sCNV burden was represented by the fraction of the genome with CNV and was termed GII. Specifically, amplification and deletion were identified according to the log2 ratio of the segment to derive GIIAmp and GIIDel. In the 23 tumor samples, the medians (ranges) of GIIAmp and GIIDel were 24% (0–42%) and 22% (0–40%), respectively. For the lymph node samples, 20% (0–40%) GIIAmp and 18% (0–25%) GIIDel were identified (Fig. S1A).
WGD was present in 52% (12/23) of the tumors and 16.7% (1/6) of the lymph nodes (Fig. 1B). There was no significant difference in GII and TMB between patients with WGD and those without WGD (Fig. S1B–D). Comparison between the paired tumor and lymph nodes revealed no significant differences in either GII or TMB (Fig. S1E–G).
We set up a catalog including 848 putative driver genes according to the literature and the COSMIC database, as described in the methods (Table S3). The mutational landscape of these candidate driver genes was investigated in the PMME dataset. Recurrently mutated driver genes were identified, and MUC16, RANBP2, NRAS, TP53, PTPRT, NF1, MUC4, KMT2C, BRAF, and ANK1 were the most frequently mutated (Fig. 1B). All mutations targeting RANBP2, NRAS, BRAF, PTEN, SF3B1, ANK1, CALR, and EPPK1 were putatively deleterious (Fig. 1B). Nonsynonymous mutations in NRAS and BRAF were mutually exclusive (Fisher’s exact test, P < 0.001).
MUC16 is a mucin marker frequently mutated in melanoma, and mutations in this gene were highly recurrent among the PMME samples (44.4%, 8/18). Samples with MUC16 mutations had a significantly higher TMB than those with the wild-type gene in the PMME, SKCM, and MM cohorts (Fig. S2).
Several acquired genomic alterations were observed in the PMME cohort. As revealed by the GISTIC2.0 results, at the arm level, copy number gains of 1q, 6p, and 8q and copy number losses of 1p, 1q, 6q, 10p, 10q, and 11p were observed (Fig. 2A). Given that the small-scale PMME cohort was not sufficiently powered for GISTIC to characterize significant sCNVs (Fig. S3A, B), we also investigated the regions of the focal copy number aberrations using a melanoma-associated gene list (Table S4). A batch of well-recognized cancer genes was identified as concurrent focal amplification/deletion events, including the loss of SKI and CDKN2A and gain of NRAS, BRAF, TERT, MDM2, CDK4/6, GNA11/GNAQ, and FAT4 (Fig. 2B).
A The landscape of arm-level amplifications and deletions across the PMME genome determined by GISTIC2.0. B Recurrent CNVs identified per lesion in published melanoma-related genes are color-coded: light blue boxes indicate hap_LOH, dark blue boxes indicate homo_LOH, pink boxes indicate dup_LOH, red boxes indicate Dup, and gray boxes indicate cn_LOH. LOH, loss of heterozygosity; hap_LOH, haploid LOH; homo_Del, homozygous deletion; dup_LOH, duplication LOH; Dup, duplication; cn_LOH, copy neutral LOH; see Methods section for details.
The UVR-related mutational signature is identified in PMME
The C > T transition, characteristic of the UVR-induced mutational signature, was the most abundant single nucleotide substitution in PMME (median: 34.9%, range: 4.8–60.7%, Fig. 3A). A high proportion of SBS5, a signature with unknown etiology but speculated to be “clock-like”, was identified in over 94% (17/18) of PMME samples (Fig. 3B). In addition, SBS6 (defective DNA mismatch repair) and other signatures of unknown etiology (SBS1, SBS37, and SBS39) were also identified (Fig. 3C). Interestingly, SBS38 (unknown etiology, indicative of indirect UVR) was identified in 83% (15/18) of PMME samples, and the C > A transversion fraction in each sample correlated with the SBS38 contribution (Fig. 3C, D). Given the plausible UV-associated etiology and the UVR-free environment of the esophagus, the role of SBS38 and its relevance in UV damage warrant further exploration. In addition, the samples with SBS38 tended to harbor higher SNV numbers than those without SBS38, although the difference was not significant in PMME (Fig. 3E, F).
A Proportion of each mutation substitution and the fraction of transition (Ti) and transversion (Tv) categories in PMME. B Proportion of each identified mutational signature in four melanoma cohorts (PMME, SKCM, MM, and UVM). C–E Proportion of mutational signatures (C) mutation substitutions (D) and synonymous and nonsynonymous SNVs in each lesion. The samples are arranged in descending order of the SBS38 contribution. F Box plot of SNVs in melanoma samples that have evidence of SBS38 or have no evidence of this signature (Mann–Whitney U test). In each box plot, the box boundaries show the first to third quartiles, and the median is the centerline. From left to right, the cohorts are PMME, SKCM, and MM, respectively.
Comparison of the genomic features between PMME and other types of melanoma
The SKCM, NEMM, and UVM genomic data were collected to compare the genomic features with PMME. Since PMME is a subtype of MM and several previous case studies demonstrated that it presented genomic characteristics similar to those of NEMM, we merged our PMME cohort with the published NEMM cohort (n = 67) for some analyses (termed the MM cohort). Furthermore, multiple samples from one patient were merged to avoid bias.
The mutational landscape of driver genes and mutational signatures was explored across the PMME (n = 18), SKCM (n = 398), MM (n = 85, 18 PMMEs included), and UVM (n = 79) cohorts (Figs. 3A and 4A). SBS7b, a UVR-related signature, was identified in both the SKCM and MM cohorts but was absent from the other two cohorts. SBS38 was identified in 60% (239/398) of SKCMs, 83% (15/18) of PMMEs and 87% (74/85) of MMs. In contrast, SBS1, SBS6, and SBS39 were shared by UVM, MM, and PMME but not by SKCM (Fig. 3B).
The frequency of recurrently mutated driver genes in PMME exhibited a consistent pattern among the PMME and NEMM and SKCM cohorts (Fig. 4B, C). A similar trend was also observed between the SKCM and MM cohorts regarding the top 20 most frequently mutated driver genes in SKCM (Fig. 4D). Notably, although the mutation frequency of NRAS or BRAF in the PMME/MM cohort was lower than that in the SKCM cohort, these genes were still among the top 10 mutated driver genes in the PMME/MM cohorts. In the UVM cohort, only SF3B1 and MUC16 were frequently mutated, and other PMME-related driver genes were not. As expected, SKCM harbored the highest mutation burden, with a median burden of 12.4 Muts/Mb (range: 0.2–172.8), followed by PMME (median: 3.3 Muts/Mb, range: 0.5–9.8) and UVM (median: 0.33 Muts/Mb, range: 0.12–11.3). The TMB of SKCM was significantly higher than that in PMME and UVM (Fig. 4E). Consistently, the mutation burden in the MM cohort was significantly lower than that in the SKCM cohort, with a median of 1.8 Muts/Mb (range: 0.2–9.8) (Fig. 4F). We further investigated the mutation distribution across the linear protein structure of BRAF and NRAS in the PMME, MM, and SKCM cohorts. We found that most mutations of BRAF and NRAS in the three cohorts affected the same protein domains. Hotspot mutations of SKCM, such as NRAS p. Q61 and p. G12 and BRAF p. V600E were also found in the PMME cohort (Fig. S4).
A Heatmap showing the frequency of the most frequently mutated genes in the five cohorts, PMME (18 cases), SKCM (398 cases), UVM (79 cases), NEMM (67 cases), and MM (including 67 NEMMs and 18 PMMEs). B Comparison of the frequency of recurrently mutated driver genes of PMME between PMME and NEMM. C Comparison of the frequency of recurrently mutated driver genes of PMME between PMME and SKCM. D Comparison of the frequency of the top 20 mutated driver genes of SKCM between PMME and SKCM. E, F Box plot with mutation burden in the UVM, SKCM, and PMME cohorts. P values determined by a two-tailed Mann–Whitney U test, *P < 0.0001.
Genomic instability was further compared across the PMME, SKCM, and UVM cohorts. Compared with UVM, PMME and SKCM exhibited prominent amplification in chromosomes 1, 3, and 7 (Fig. S5A). Both PMME and SKCM harbored significantly amplified 1q, 6p, and 8q according to the GISTIC analysis (Fig. S5B). The amplification of 6p21.2 and deletion of 9p21.3 (CDKN2A) and 12p24.33 were significantly recurrent focal regions in both the SKCM and PMME cohorts (Fig. S5C, D). In summary, the CNV characteristics of PMME and SKCM were highly concordant at both the arm and focal levels. Additionally, copy number changes were consistent across the four SKCM subtypes, which were divided by the body site of the primary tumor, including head and neck, trunk, extremities, and ambiguous sites (Fig. S6), indicating that UVR might not be a major external factor contributing to genomic instability.
The PMME pathway is consistent with the classic melanoma pathway
The classic melanoma pathways involve the activation of Raf-MEK-ERK, PI3K-Akt, p16INK4a/cyclin-dependent kinases 4 and 6/retinoblastoma protein (p16INK4a/CDK4,6/pRb) and p14ARF/human double minute 2/p53 (p14ARF/HMD2/p53) tumor suppressor pathways (KEGG pathway database, map05218). Somatically altered genes in the PMME, SKCM, and MM cohorts were selected and compared with the determined melanoma pathways (Fig. 5). Oncogenic NRAS mutations activate both the PI3K-Akt pathway and Raf-MEK-ERK, the latter of which could also be activated by BRAF mutations. Key oncogenes in the melanoma pathways, such as NRAS, BRAF, and PIK3CA, were altered in the PMME, SKCM, and MM cohorts. Key tumor suppressor genes, such as PTEN and TP53, which promote tumor progression when functionally lost, were also frequently altered in the three groups (Fig. 5).
Multiple regional analysis revealed the genomic complexity of PMME
Trunk and branch mutations were identified using a multiregional sequencing strategy to investigate the ITH and general genome evolution of PMME. Multiple sites of primary tumor were obtained from two patients (CXG and LJP), and metastatic lymph nodes were obtained from three patients (HXC, SZY, and QXL). Of note, the phylogeny of patient HZY was reported in detail in our previous study; therefore, it was not included in the current analysis10. A phylogenetic tree was constructed for each patient based on the presence and absence of somatic mutations in each sample. Generally, the PMME genome presented typical branched rather than linear tumor evolution (Fig. S7A–E). The proportion of private mutations in each sample accounted for 21% to 68% of the mutation load (Fig. S8A). We also calculated the ITH index and genetic divergence between the paired primary tumors and lymph nodes (GDPL) derived from the number of trunk and branch mutations to evaluate the patient-level heterogeneity and found a high degree of both ITH (0.78 and 0.69 in CXG and LJP, respectively) and GDPL (range: 0.48–0.75, Fig. S8B). The clonality analysis of a single sample was performed by calculating the CCFs of the mutations. The fraction of subclonal mutations in all PMME samples ranged from 0 to 1, with a median of 0.62. The fraction of subclonal mutations in primary tumors (median: 0.75, range: 0.33–1) was higher than that in metastatic lymph node samples (median: 0.41, range: 0–0.56, P = 0.01, Mann–Whitney U test, Fig. S9A–C). The paired primary tumors and lymph nodes were compared, and the fraction of subclonal mutations in primary lesions was significantly higher than that in lymph nodes (P = 0.02, Mann–Whitney U test, Fig. S9D). These findings suggest that PMME generally presents with high ITH, and lymph node metastasis possibly occurs with clonal selection.
Driver mutations in TP53, NRAS, SF3B1, and FBXW7 were clonal mutations located in the trunk, while RANBP2, RGPD3, PTEN, and SETD2 mutations were subclonal and occurred along the branch (Fig. S7A–E).
Potential parallel tumor evolution events were observed in two patients (CXG and LJP). Two branches harbored different mutations in the same gene, suggesting that such alterations may adapt to the PMME niches and may be selected during tumor evolution. Different mutations in RANBP2, RGPD3, and PTEN were identified in the independent tumor clones in an individual (Fig. S7A). Notably, RANBP2 was the second most frequently mutated gene in PMME and had a multiple-point mutation pattern. Six samples (CXG-T1, CXG-T2, LJP-T1, LJP-T2, SFX-T1, and YGH-T) harbored RANBP2 mutations with an average mutation number of 3.5 mutations per sample (range: 1–7).
WGD is an early truncal event and may promote genomic instability
The temporal order of the occurrence of WGD and mutations or CNV during tumor progression was inferred based on clonality analysis. WGD was an early driver event and appeared mostly in the trunk (Fig. 1B, upper annotation). Many genomic aberrations following genome duplication indicated that WGD may promote genomic instability and drive tumorigenesis (Fig. S10). Most potential melanoma driver genes were mutated after WGD, such as MUC16, MUC4, RANBP2, and NF1. TP53 was the only driver gene in which mutations occurred before WGD (Table S5). We were not able to evaluate the relationship between WGD and NRAS and BRAF gene mutations due to the small sample size.
GIIAmp and mutation signature predicts inferior prognosis in PMME
We performed an association analysis between the genotype and clinical phenotypes of PMME to investigate the potential influence of genetic characteristics on clinical features. The OS and RFS data were available for all 18 patients with PMME. We found that the mutation signatures SBS38, SBS6, and GIIAmp were associated with poor prognosis in the PMME cohort. The Kaplan–Meier survival analysis and log-rank test demonstrated that patients with a high proportion of SBS38 exhibited shorter OS times (log-rank test, P = 0.0108, Fig. 6A). Patients with a high SBS6 proportion had shorter RFS durations (log-rank test, P = 0.0084, Fig. 6B). GIIAmp was a numerical variable calculated based on the fraction of the genome affected by amplification as defined previously. The patients with PMME were divided into the high GIIAmp group (5 patients) and the low GIIAmp group (13 patients). The high GIIAmp group showed a significantly shorter OS duration than the low GIIAmp group (log-rank test, P = 0.0229, Fig. 6C).
A Patients with PMME with a high proportion of SBS38 experienced shorter OS times than the remaining patients with a lower proportion of SBS38. B Patients with PMME with a high proportion of SBS6 experienced worse RFS outcomes than the remaining patients with a lower proportion of SBS6. C The patients with PMME were divided into a high GIIAmp group and a low GIIAmp group, and the high GIIAmp group showed a significantly shorter OS duration than the low GIIAmp group. D Patients with SKCM with a high proportion of SBS38 had a shorter OS duration than those without or with a low proportion of SBS38. E Patients with SKCM with high GIIAmp tended to have a shorter OS time, although the difference was not significant. A–E The number of patients (n) per group is indicated.
A multivariate Cox regression model was used to adjust for confounding factors, such as age and sex. The multivariate analysis confirmed that a high proportion of SBS38 was a potential poor prognostic factor for OS (HR: 4.657, 95% CI: 1.077–20.149, P = 0.0395). GIIAmp was also associated with OS, although the result was not significant (HR: 4.200, 95% CI: 0.765–23.047, P = 0.0985). Patients with SBS6 tended to have a shorter RFS duration (HR: 7.381, 95% CI: 1.537–35.436, P = 0.0125).
The SKCM cohort was used to validate the above observations. Consistent with the results that we demonstrated in PMME, the SKCM patients with a high proportion of SBS38 had shorter OS times than those without or with a low proportion of SBS38 (log-rank test, P = 0.0419, Fig. 6D). Patients with SKCM with a high GIIAmp tended to have a shorter OS, although only marginal significance was obtained (log-rank test, P = 0.0791, Fig. 6E).
Identification of actionable (potentially influencing therapy) mutations in PMME
Due to the rarity of PMME, there is little knowledge regarding the rate of patient response to targeted therapy, even among those with MM. Previously, we established a pipeline to identify clinically actionable mutations by annotating genomic variants using the American College of Medical Genetics and Genomics/Association for Molecular Pathology (ACMG/AMP) 2015 and AMP/American Society of Clinical Oncology/College of American Pathologists (AMP/ASCO/AP) 2017 Guidelines and Standards. Here, we used this self-established pipeline to identify potentially exploitable mutations responsive to known targeted therapy drugs. Somatic aberrations identified in PMME were compatible with a range of treatment options. Of the 18 tested PMME patients, only 6 patients had no potential therapeutic target; over 66.6% (12/18) of patients with PMME carried at least one mutation that potentially corresponds to a targeted drug (Table S6).
Discussion
In this study, we depicted the global genomic profiles of a PMME cohort. To the best of our knowledge, this is not only the first study to thoroughly compare the mutational and CNV landscape between PMME and other subtypes of melanoma but also the first to point out potential indicators of clinical outcomes and actionable mutations in a relatively large PMME cohort. Although the sampling of rare tumors is challenging, we performed multiple samplings from several patients and conducted a phylogeny and clonality analysis to trace the clonal evolution trajectory of the key driver events involved in PMME progression. According to our observations, genomic features such as SBS38, SBS6, and genomic amplification might be potential indicators of clinical outcomes. Moreover, actionable mutations were identified in the PMME samples, which warrants further investigation, e.g., in clinical trials.
UVR-induced DNA damage has been identified as the key environmental driver of SKCM17. SBS7 is regarded as a single signature composed of C > T at CCN and TCN trinucleotides, together with fewer T > N mutations. SBS7 is dominant in UV-related melanoma and likely results from UV light-induced formation of pyrimidine dimers followed by translation DNA synthesis by error-prone polymerases, predominantly inserting A opposite to the damaged cytosines18. As a UVR-related signature, SBS7 was also identified in a subset of MM and AM6,8,9,19. As Newell et al. observed, this signature occurred predominantly in MM samples from upper body sites but was absent in those from lower mucosal body sites8. Mundra et al. identified SBS7 in a subset of MMs and found that MMs with SBS7 have similar genetic patterns to SKCM6. Our previous study described a PMME genome with significant shifts in the mutational signatures during tumorigenesis, and the decreased proportion of the C > T transversion was accompanied by an increase in the C > A transition over time10. In this study, high proportions of C > T and C > A alterations were observed in the PMME cohort. SBS7 was identified in both the MM and SKCM cohorts, but it was absent from the PMME and UVM cohorts. These observations are consistent with those of previous studies.
Of note, we found that another UVR-related signature, SBS38, existed extensively in the PMME genome. According to the literature and COSMIC database (https://cancer.sanger.ac.uk/signatures/sbs/sbs38/), SBS38 is found only in UV light-associated melanomas; therefore, it is postulated to be the result of indirect damage from UV light18. The etiology of SBS38 may be associated with oxidative stress, as the signature has been reported to cluster with other oxidation damage repair-related signatures, when using the simple probabilistic model DNA Repair FootPrint (RePrint) to represent each signature20. SBS38 was found to occur exclusively with other UVR signatures in AM, indicating that it plays an independent role in causing mutations19. We further interrogated the clinical role of SBS38 and found that a high proportion of SBS38 indicated an inferior OS outcome in both the PMME and SKCM cohorts. PMME occurs in the esophagus, an environment without UVR exposure; therefore, it is not clear how UV light generates an indirect effect on melanocytes arising from the esophageal mucosa. SBS38 is present only in epithelium-associated melanoma despite the status of UVR exposure, indicating that it may not be necessarily induced by UV light but by an unknown mechanism resulting from the indirect effect of UVR.
SBS38 was associated with the mutation burden in both SKCM and MM. The median mutation burden of PMME samples with SB38 was also higher than samples without SBS38, although there was no significance due to the limited sample size (Fig. 3F). Moreover, a high proportion of SBS38 predicted poor OS outcomes in the SKCM and PMME cohorts (Fig. 6A, D). In contrast to SBS38, the well-known UVR signature SBS7 is also correlated with the TMB in melanoma6; however, patients with a high proportion of SBS7 had better OS outcomes than those without or with a low proportion of SBS7 in the SKCM cohort (Fig. S11). Our analysis suggested that multiple mutational processes were involved in melanoma tumorigenesis, that UVR exposure represented by SBS7 initiated somatic mutation accumulation, and as the tumor progressed, an unknown exogenous or endogenous exposure associated with SBS38 may drive the melanoma to deteriorate further. Our data suggest that SBS38 might play an underappreciated role in the pathogenesis of melanoma and may be generated by the microenvironment of epithelium-associated originating melanocytes.
Mucosal melanoma has long been considered distinct from cutaneous melanoma; thus, patients might not benefit from treatments currently used for common cutaneous melanoma. There is no standard TNM staging system for PMME owing to its scarcity. The lack of systematic studies has impeded the effective treatment of this disease. In this study, we comprehensively described the whole-exome landscape of PMME and compared the genetic features between melanomas from different primary sites. MUC16 was the most frequently mutated gene in our PMME cohort (44%), and its mutation rates in MM and SKCM were 18% and 73%, respectively. Several recent studies have revealed that MUC16 mutations are associated with the TMB in cutaneous melanoma and several other solid tumors; furthermore, researchers provided evidence that MUC16 mutation could predict immunotherapy efficacy and suggested that it could be a surrogate biomarker for the global TMB21,22,23,24,25. In our data, the TMB was also significantly higher in patients with PMME with MUC16 mutations, and most of the MUC16 mutations were passenger mutations (Fig. 1B). We speculated that MUC16 might not play a role as a driver in melanoma and, for this reason, could be used to measure the overall mutational burden compared with other regions of the genome.
NRAS, BRAF, NF1, TP53, PTEN, and SF3B1 mutations and other melanoma driver mutations were observed in PMME samples. We compared the frequency of recurrently mutated genes in PMME across different melanoma cohorts. Our data showed that the prevalence of most mutated genes was comparable across SKCM, MM, and PMME. As reported by others4,26,27, NRAS mutations were frequent in PMME or MM, while BRAF mutations were frequent in SKCM. In our study, we observed a high prevalence of mutations in both NRAS and BRAF in PMME. The prevalence of BRAF mutations was 17% (rank: 3) and 16% (rank: 3) in the PMME and MM cohorts (combination of PMME and MM reported by Newell et al., Fig. 4A), respectively. Although the absolute prevalence of BRAF mutations was lower in PMME/MM than in SKCM (53%), its prevalence was top-ranked, as seen in SKCM (rank: 2). The overall mutation burden is significantly higher in SKCM than in other melanomas due to the exposure to UVR; therefore, it is not appropriate to directly compare the absolute frequency between cohorts. Therefore, we used Spearman’s rank correlation to evaluate the mutation frequency distribution of critical gene sets in SKCM and other melanomas with low mutation rates (Fig. 4B, C) and found globally concordant patterns. SF3B1 was another frequently mutated gene in PMME (rank: 4) and MM (rank: 4); this gene is the most commonly mutated gene in UVM but rarely mutated in SKCM. We also observed a high consistency of copy number landscape alternations between SKCM and PMME, especially when compared with UVM (Fig. S5). At the focal level, typical genomic alterations, such as the amplification of 6p21.2 and deletion of 9p21.3 containing CDKN2A, occurred in both the PMME and SKCM genomes. We also demonstrated the frequent copy number gain of MDM2, CDK4, CDK6, and TERT in PMME; however, segments covering these genes were not identified as significantly altered regions by GISTIC analysis, possibly due to the small sample size. Overall, the pathways affecting SKCM were also involved in PMME, such as Raf-MEK-ERK. We hypothesized that in different types of epithelium-associated melanoma, the source of mutagenesis might vary at the initiation stage. However, as mutations accumulate, the microenvironment generated by epithelial melanocytes may be similar and shape the melanoma genome by selecting for a tumor clone with the same genetic defects, conferring a fitness advantage to the surviving clone. Therefore, our analysis suggested that PMME might have a genomic pattern and biological characteristics similar to its common counterpart. In addition, patients might benefit from the successful therapeutic approaches used for common melanomas.
Several studies have focused on the ITH and genomic evolution of MM. We have described a PMME case by using multiregional WES and observed complex genomic characteristics such as chromothripsis and parallel evolution of chromosome 4 in the PMME genome10. BRAF and KRAS mutations, as well as CDKN2A biallelic inactivation, were truncal alterations, while PIK3CA and PTEN deletions occurred in the branch. In this study, we assessed genomic events during PMME progression in a larger PMME cohort using comparative multiple lesion sequencing and clonality analysis. Whole-genome duplication (WGD) was an early truncal event concerning most driver mutations (MUC16, MUC4, RANBP2, and NF1 mutations), but it was a late event relative to TP53 driver mutations, as others reported previously28. Birkeland et al. found that mutated BRAF and NRAS and low-level gains of mutated BRAF alleles, occurred earlier than WGD. Unfortunately, the number of samples with WGD and concurrent BRAF or NRAS mutations was too small to perform this analysis; therefore, we could not evaluate their timing. BRAF mutations were found in normal skin and were suggested to result from positive selection during melanoma pathogenesis and from TP53 mutations29. TP53 mutations are found in histologically normal tissues, such as skin, esophageal epithelium, and breast luminal cells29,30,31,32. It has been suggested that the carcinogen-exposed epithelium is similar to a polyclonal quilt of driver mutations subjected to selection and that TP53 acts as a guardian in precancers by switching cells from the exponential division resembling self-renewal to a linear, asymmetric division. TP53 controls the genomic stability of stem cells, regulation of cell differentiation, and maintenance of structural organization in human brain organoids33. However, the interplay between TP53 and genome instability remains unclear. It is believed that WGD can be selected to mitigate the irreversible, ratchet-like accumulation of deleterious somatic alterations34. Some studies have suggested that WGD is a macroevolutionary event that shapes tumor evolution; it is highly common among cancers and is associated with poor prognosis in several cancers14. Our data indicate that both TP53 mutations and WGD are early genomic events in PMME, and TP53 mutations occurred before WGD, consistent with previous reports14,35. The TP53 mutant subclone may expand under WGD conditions. Both TP53 mutations and WGD may contribute to genomic instability and further cause the accumulation of genetic alterations. The GII was used as a measurement of genomic instability (see the Materials and Methods section), and could be further divided into segmented copy number amplification (GIIAmp) and deletion (GIIDel). We found that GIIAmp was associated with a poor prognosis in the PMME cohort (Fig. 6C). Our findings reveal the genomic complexity of PMME and provide new evidence for tumor evolution in epithelial melanoma.
A less noticeable gene, RANBP2, was recurrently mutated in the PMME cohort. Moreover, multiple mutations and a parallel evolution model were also observed in this gene, highlighting the importance of RANBP2 in PMME pathogenesis. RANBP2 was one of the most recurrently mutated genes, with an average mutation number of 3.5 mutations per PMME sample. Furthermore, all of the identified RANBP2 mutations were predicted to be putative deleterious missense mutations in BRAF and NRAS. According to the tumor evolutionary analysis, RANBP2 aberrations preferentially occurred in independent evolutionary clades (Fig. S7A, B), and this feature fits the parallel evolutionary theory of clonal evolution of cancer. Based on the above findings, tumor cells with RANBP2 alterations may have growth advantages and adapt well to the PMME niche. RANBP2 is a RanGTPase-binding protein that directly interacts with the E2 enzyme UBC9 and strongly enhances SUMO1 transfer from UBC9 to the SUMO1 target SP10036,37. Some studies have suggested that it could regulate the Mdm2-p53 signaling axis38. The frequency of RANBP2 mutation in SKCM was approximately 9%; however, it has not been previously suggested as an important driver gene in melanoma. Further studies are required to confirm the potential role of RANBP2 in melanoma tumorigenesis.
Treatment, especially immunotherapy, and melanoma patient survival have dramatically improved in the last 10 years39; however, patients with MM, including PMME, have limited benefits compared with those with cutaneous melanoma40. As reported in previous studies9,41, CDK4/6 amplification is common in PMME, which could be inhibited by palbociclib or other inhibitors42. We further performed an analysis to evaluate the potential benefits of targeted therapy in patients with PMME. Our data suggested several altered genes that may be amenable to precision medicine. Among the 18 tested patients, only 5 had no druggable mutations, while the rest had at least one targeted therapy option (Table S2). The driver mutations in the MAPK pathway, including mutations in BRAF, PTEN, NF1, SF3B1, TP53, SPRED1, and PIK3CA, in 55% (16/29) of the studied samples, indicated that MEK/MAPK inhibition might be suitable for a large proportion PMME patients.
We acknowledge that the limited sample size prevented the acquisition of a broad picture of the disease that a large-scale study could obtain. With only two multilesion sampling cases (two lesions of each were available) and three pairs of primary tumors and lymph nodes subjected to phylogenetic construction, we could not identify more recurrent events, such as lymph node-specific features or trunk-unique characteristics, than those observed in RANBP2 and WGD. Since samples with concurrent WGD and driver events were rare, inference of the temporal order of the driver mutations in BRAF or NRAS relative to WGD was not possible, as mentioned above. The statistical power suffered attrition. For example, when GISTIC2.0 was adopted to determine the significantly altered regions, the discovery was limited to only one amplified region identified. Similarly, multivariate regression analysis may require more samples to support the statistical integrity. Characterizing features associated with the clinical outcomes would also dictate a larger dataset than the current PMME cohort, although validation in a large TCGA SKCM cohort was performed. There were also limitations related to the external nature of the data, as samples were collected from nonprimary sites, and some of the research subjects were not treatment-naïve. Consortium work and multicenter collaborations are warranted to help enlarge the scale of studies and increase the number of samples of this rare disease.
In conclusion, different types of epithelium-associated melanoma may have a similar genomic pattern despite the differences in the initiation of the mutational process. Based on the genetic characteristics of PMME, patients might benefit from the current treatment used for common cutaneous melanoma. Further clinical studies regarding this issue are worthwhile.
Data availability
The sequence reported in this paper has been deposited in the CNGBdb (China National GeneBank database, https://www.cngb.org) under accession number CNP0001947. The remaining data are available within the supplementary files or from the authors upon request.
References
Wallis, G., Sehgal, V., Haider, A., Bridgewater, J., Novelli, M., Dawas, K. et al. Primary malignant melanoma of the esophagus. Endoscopy 47 Suppl 1 UCTN, E81–82 (2015).
Spencer, K. R. & Mehnert, J. M. Mucosal Melanoma: Epidemiology, Biology and Treatment. Cancer Treat Res 167, 295–320 (2016).
Lasota, J., Kowalik, A., Felisiak-Golabek, A., Zieba, S., Waloszczyk, P., Masiuk, M. et al. Primary malignant melanoma of esophagus: clinicopathologic characterization of 20 cases including molecular genetic profiling of 15 tumors. Mod Pathol 32, 957–966 (2019).
Tsuyama, S., Kohsaka, S., Hayashi, T., Suehara, Y., Hashimoto, T., Kajiyama, Y. et al. Comprehensive clinicopathological and molecular analysis of primary malignant melanoma of the oesophagus. Histopathology 78, 240–251 (2021).
Bastian, B. C. The molecular pathology of melanoma: an integrated taxonomy of melanocytic neoplasia. Annu Rev Pathol 9, 239–271 (2014).
Mundra, P. A., Dhomen, N., Rodrigues, M., Mikkelsen, L. H., Cassoux, N., Brooks, K. et al. Ultraviolet radiation drives mutations in a subset of mucosal melanomas. Nat Commun 12, 259 (2021).
Wang, X., Kong, Y., Chi, Z., Sheng, X., Cui, C., Mao, L. et al. Primary malignant melanoma of the esophagus: A retrospective analysis of clinical features, management, and survival of 76 patients. Thorac Cancer 10, 950–956 (2019).
Newell, F., Kong, Y., Wilmott, J. S., Johansson, P. A., Ferguson, P. M., Cui, C. et al. Whole-genome landscape of mucosal melanoma reveals diverse drivers and therapeutic targets. Nat Commun 10, 3163 (2019).
Zhou, R., Shi, C., Tao, W., Li, J., Wu, J., Han, Y. et al. Analysis of Mucosal Melanoma Whole-Genome Landscapes Reveals Clinically Relevant Genomic Aberrations. Clin Cancer Res 25, 3548–3560 (2019).
Li, J., Yan, S., Liu, Z., Zhou, Y., Pan, Y., Yuan, W. et al. Multiregional sequencing reveals genomic alterations and clonal dynamics in primary malignant melanoma of the esophagus. Cancer Res 78, 338–347 (2018).
Bailey, M. H., Tokheim, C., Porta-Pardo, E., Sengupta, S., Bertrand, D., Weerasinghe, A. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385 e318 (2018).
Vogelstein, B., Papadopoulos, N., Velculescu, V. E., Zhou, S., Diaz, L. A., Jr. & Kinzler, K. W. Cancer genome landscapes. Science (New York, N.Y.) 339, 1546–1558 (2013).
Hu, Z., Ding, J., Ma, Z., Sun, R., Seoane, J. A., Scott Shaffer, J. et al. Quantitative evidence for early metastatic seeding in colorectal cancer. Nat Genet 51, 1113–1122 (2019).
Bielski, C. M., Zehir, A., Penson, A. V., Donoghue, M. T. A., Chatila, W., Armenia, J. et al. Genome doubling shapes the evolution and prognosis of advanced cancers. Nat Genet 50, 1189–1195 (2018).
Nahar, R., Zhai, W., Zhang, T., Takano, A., Khng, A. J., Lee, Y. Y. et al. Elucidating the genomic architecture of Asian EGFR-mutant lung adenocarcinoma through multi-region exome sequencing. Nat Commun 9, 216 (2018).
Wang, Z., Duan, J., Cai, S., Han, M., Dong, H., Zhao, J. et al. Assessment of blood tumor mutational burden as a potential biomarker for immunotherapy in patients with non-small cell lung cancer with use of a next-generation sequencing cancer gene panel. JAMA Oncol 5, 696–702 (2019).
Cancer Genome Atlas, N. Genomic Classification of Cutaneous Melanoma. Cell 161, 1681–1696 (2015).
Alexandrov, L. B., Kim, J., Haradhvala, N. J., Huang, M. N., Tian Ng, A. W., Wu, Y. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Newell, F., Wilmott, J. S., Johansson, P. A., Nones, K., Addala, V., Mukhopadhyay, P. et al. Whole-genome sequencing of acral melanoma reveals genomic complexity and diversity. Nat Commun 11, 5259 (2020).
Wojtowicz, D., Leiserson, M. D. M., Sharan, R. & Przytycka, T. M. DNA Repair Footprint Uncovers Contribution of DNA Repair Mechanism to Mutational Signatures. Pac Symp Biocomput 25, 262–273 (2020).
Wang, Q., Yang, Y., Yang, M., Li, X. & Chen, K. High mutation load, immune-activated microenvironment, favorable outcome, and better immunotherapeutic efficacy in melanoma patients harboring MUC16/CA125 mutations. Aging (Albany NY) 12, 10827–10843 (2020).
Wang, X., Yu, X., Krauthammer, M., Hugo, W., Duan, C., Kanetsky, P. A. et al. The Association of MUC16 Mutation with Tumor Mutation Burden and Its Prognostic Implications in Cutaneous Melanoma. Cancer Epidemiol Biomarkers Prev 29, 1792–1799 (2020).
Yang, Y., Zhang, J., Chen, Y., Xu, R., Zhao, Q. & Guo, W. MUC4, MUC16, and TTN genes mutation correlated with prognosis, and predicted tumor mutation burden and immunotherapy efficacy in gastric cancer and pan-cancer. Clin Transl Med 10, e155 (2020).
Yu, Y., Lin, D., Li, A., Chen, Y., Ou, Q., Hu, H. et al. Association of immune checkpoint inhibitor therapy with survival in patients with cancers with MUC16 variants. JAMA Netw Open 3, e205837 (2020).
Zhang, L., Han, X. & Shi, Y. Association of MUC16 mutation with response to immune checkpoint inhibitors in solid tumors. JAMA Netw Open 3, e2013201 (2020).
Sekine, S., Nakanishi, Y., Ogawa, R., Kouda, S. & Kanai, Y. Esophageal melanomas harbor frequent NRAS mutations unlike melanomas of other mucosal sites. Virchows Archiv Int J Pathol 454, 513–517 (2009).
Langer, R., Becker, K., Feith, M., Friess, H., Hofler, H. & Keller, G. Genetic aberrations in primary esophageal melanomas: molecular analysis of c-KIT, PDGFR, KRAS, NRAS and BRAF in a series of 10 cases. Mod Pathol 24, 495–501 (2011).
Birkeland, E., Zhang, S., Poduval, D., Geisler, J., Nakken, S., Vodak, D. et al. Patterns of genomic evolution in advanced melanoma. Nat Commun 9 (2018).
Martincorena, I., Roshan, A., Gerstung, M., Ellis, P., Van Loo, P., McLaren, S. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015).
Wang, X., El-Halaby, A. A., Zhang, H., Yang, Q., Laughlin, T. S., Rothberg, P. G. et al. p53 alteration in morphologically normal/benign breast luminal cells in BRCA carriers with or without history of breast cancer. Hum Pathol 68, 22–25 (2017).
Martincorena, I., Fowler, J. C., Wabik, A., Lawson, A. R. J., Abascal, F., Hall, M. W. J. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Yokoyama, A., Kakiuchi, N., Yoshizato, T., Nannya, Y., Suzuki, H., Takeuchi, Y. et al. Age-related remodelling of oesophageal epithelia by mutated cancer drivers. Nature 565, 312–317 (2019).
Marin Navarro, A., Pronk, R. J., van der Geest, A. T., Oliynyk, G., Nordgren, A., Arsenian-Henriksson, M. et al. p53 controls genomic stability and temporal differentiation of human neural stem cells and affects neural organization in human brain organoids. Cell Death Dis 11, 52 (2020).
Lopez, S., Lim, E. L., Horswell, S., Haase, K., Huebner, A., Dietzen, M. et al. Interplay between whole-genome doubling and the accumulation of deleterious alterations in cancer evolution. Nat Genet 52, 283–293 (2020).
McGranahan, N., Favero, F., de Bruin, E. C., Birkbak, N. J., Szallasi, Z. & Swanton, C. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci Transl Med 7, 283ra254 (2015).
Navarro, M. S. & Bachant, J. RanBP2: a tumor suppressor with a new twist on TopoII, SUMO, and centromeres. Cancer Cell 13, 293–295 (2008).
Dawlaty, M. M., Malureanu, L., Jeganathan, K. B., Kao, E., Sustmann, C., Tahk, S. et al. Resolution of sister centromeres requires RanBP2-mediated SUMOylation of topoisomerase IIalpha. Cell 133, 103–115 (2008).
Blondel-Tepaz, E., Leverve, M., Sokrat, B., Paradis, J. S., Kosic, M., Saha, K. et al. The RanBP2/RanGAP1-SUMO complex gates beta-arrestin2 nuclear entry to regulate the Mdm2-p53 signaling axis. Oncogene 40, 2243–2257 (2021).
Curti, B. D. & Faries, M. B. Recent Advances in the Treatment of Melanoma. N Engl J Med 384, 2229–2240 (2021).
Mao, L., Qi, Z., Zhang, L., Guo, J. & Si, L. Immunotherapy in acral and mucosal melanoma: current status and future directions. Front Immunol 12, 680407 (2021).
Lyu, J., Song, Z., Chen, J., Shepard, M. J., Song, H., Ren, G. et al. Whole-exome sequencing of oral mucosal melanoma reveals mutational profile and therapeutic targets. J Pathol 244, 358–366 (2018).
Garutti, M., Targato, G., Buriolla, S., Palmero, L., Minisini, A. M. & Puglisi, F. CDK4/6 inhibitors in melanoma: a comprehensive review. Cells 10 (2021).
Acknowledgements
We thank all the participants in this study. We would also like to thank Isabella Chao, Diana Hu, Professor Hong Cai and Professor Wenqing Li for their language suggestions.
Funding
This study was supported by the National Key R&D Program of China (No. 2018YFC0910700), National Natural Science Foundation of China (81902508, 81972842), Beijing Natural Science Foundation (No. 7192036), Special Fund of Beijing Municipal Administration of Hospitals Clinical Medicine Development (No. XMLX201841) and Beijing Municipal Administration of Hospital’s Ascent Plan (No. DFL20191101).
Author information
Authors and Affiliations
Contributions
Conceptualization: J.J.L., B.L., and N.W.; Methodology: J.J.L., Q.Y., X.X., S.Y., C.X.W., S.L.L., and Y.Q.W.; Data analysis: J.J.L., B.L., X.X., C.X.W., and Z.C.Y.; Investigation: J.J.L., B.L., Z.X.T., and S.M.P.; Resources: W.Y.G., L.H., and Y.Y.M.; Writing and visualization: J.J.L., B.L., Q.Y., and X.X.; Supervision: N.W.; Funding acquisition: J.J.L. and N.W. The authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This study was approved by the Ethics Committee of Peking University Cancer Hospital & Institute (Institutional Review Board No. 2019KT59).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Li, J., Liu, B., Ye, Q. et al. Comprehensive genomic analysis of primary malignant melanoma of the esophagus reveals similar genetic patterns compared with epithelium-associated melanomas. Mod Pathol 35, 1596–1608 (2022). https://doi.org/10.1038/s41379-022-01116-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-022-01116-5