Abstract
Schizophrenia is a severe, complex mental disorder characterized by a combination of positive symptoms, negative symptoms, and impaired cognitive function. Schizophrenia is highly heritable (~80%) with multifactorial etiology and complex polygenic genetic architecture. Despite the large number of genetic variants associated with schizophrenia, few causal variants have been established. Gaining insight into the mechanistic influences of these genetic variants may facilitate our ability to apply these findings to prevention and treatment. Though there have been more than 300 studies of gene expression in schizophrenia over the past 15 years, none of the studies have yielded consistent evidence for specific genes that contribute to schizophrenia risk. The aim of this work is to conduct a systematic review and synthesis of case–control studies of genome-wide gene expression in schizophrenia. Comprehensive literature searches were completed in PubMed, EmBase, and Web of Science, and after a systematic review of the studies, data were extracted from those that met the following inclusion criteria: human case–control studies comparing the genome-wide transcriptome of individuals diagnosed with schizophrenia to healthy controls published between January 1, 2000 and June 30, 2020 in the English language. Genes differentially expressed in cases were extracted from these studies, and overlapping genes were compared to previous research findings from the genome-wide association, structural variation, and tissue-expression studies. The transcriptome-wide analysis identified different genes than those previously reported in genome-wide association, exome sequencing, and structural variation studies of schizophrenia. Only one gene, GBP2, was replicated in five studies. Previous work has shown that this gene may play a role in immune function in the etiology of schizophrenia, which in turn could have implications for risk profiling, prevention, and treatment. This review highlights the methodological inconsistencies that impede valid meta-analyses and synthesis across studies. Standardization of the use of covariates, gene nomenclature, and methods for reporting results could enhance our understanding of the potential mechanisms through which genes exert their influence on the etiology of schizophrenia. Although these results are promising, collaborative efforts with harmonization of methodology will facilitate the identification of the role of genes underlying schizophrenia.
Similar content being viewed by others
Introduction
Schizophrenia is a severe, complex mental disorder characterized by a combination of positive symptoms (hallucinations and delusions, and psychotic symptoms in which there is a loss of contact with reality), negative symptoms (paucity of spontaneous speech, social withdrawal, and amotivation), and impaired cognitive function [1]. Overall, the sex ratio in prevalence is approximately equal, but with greater severity and earlier onset in males than in females [2]. Genetic epidemiologic studies have shown that schizophrenia is highly heritable (~80%) but with a multifactorial etiology and complex polygenic genetic architecture. There is evidence that both common and rare genetic variants, as well as diverse environmental factors contribute to its etiology [1]. Specific copy number variants (CNVs) have also been shown to be associated with schizophrenia [3].
Despite the large number of genetic variants that have been associated with schizophrenia, few causal variants have been established, and most genetic associations have not imparted useful clinical applications [1]. Moreover, many of the genome-wide associated variants are not found in genes, possibly indicating that they may have a regulatory role in modifying gene expression or are expression quantitative trait loci (eQTLs) [4]. Information on gene expression overall and the up- or downregulation of specific genes in cases as compared with unaffected controls, may provide novel information on the pathogenesis of schizophrenia. In addition, by examining gene expression in peripheral tissues, biomarkers for use in diagnosis or treatment may be found. Such peripheral biomarkers may be particularly valuable as the primary organ of interest, i.e., the brain, cannot be sampled in vivo [5].
Gene expression, the connection between the information encoded within a gene and the final functional protein, is the central concept of molecular biology. This complex, multistaged, highly regulated process involves the transcription of genes into mRNA followed by their translation into proteins [6, 7]. This process is influenced by both genetic and environmental factors. CNVs can modify gene expression through multiple potential mechanisms including the impact of gene duplication or deletion on dosage-sensitive genes, modification of regulatory elements, position effects, or regulation of normal flanking genes [8]. Differences in gene expression can be a biomarker or highlight a potential therapeutic target. While these changes may not always translate into consequential biological activity or be independently interpretable, this information can be combined with other information to highlight areas for additional research [9]. Altered gene expression has been associated with multiple disorders, such as Alzheimer’s disease [10], cancer [11], susceptibility to infection [12], and psychiatric disorders [13, 14], including schizophrenia [15]. There are also gene expression differences across tissues, sexes, ages, and genetic ancestry [16].
Techniques for assessing gene expression or transcript abundance have changed rapidly over the last 10–15 years. Advances in technologies characterizing transcript abundance levels, such as sequencing with RNAseq, and imputed expression based on tissue databases such as the Genotype-Tissue Expression (GTEx) project [17] have gradually supplanted the less expensive and less comprehensive microarray technology. Imputed gene expression has several advantages over microarray methods and RNAseq including larger sample sizes than directly measured expression, ability to target the tissue of interest for a particular disease or condition, and obviated need to control for environmental impacts, e.g., medication use or smoking.
There have been more than three hundred studies of gene expression in schizophrenia over the past 15 years, but to date there is no consistent evidence for clearly implicated genes from these findings. Moreover, though there have been several reviews of gene expression studies in schizophrenia (e.g., Bray [18], Harrison and Weinberger [19], Iwamoto and Kato [20], Kumarasinghe et al. [15], Mirnics et al. 2001 [21]), the majority were completed more than 10 years ago, and to date there has been no synthesis of the results across tissue sources and gene expression measurement technologies. Other limitations of many of the earlier studies include the use of candidate gene rather than genome-wide approaches; lack of unaffected controls; and failure to consider potential confounding factors, such as sex, age, and genetic ancestry [16]. Therefore, the overarching aim of this report is to complete a systematic review and synthesis of case–control studies of genome-wide gene expression in schizophrenia.
Methods
Scope of the review and PROSPERO registration
The scope of this systematic review is to identify genes that are differentially expressed in individuals with schizophrenia. The review was limited to noninterventional human studies that screen the genome in an unbiased manner. The review protocol was registered with PROSPERO [22], the International prospective register of systematic reviews (https://www.crd.york.ac.uk/PROSPERO/). The PRISMA checklist for this study is included in the supplementary materials (Supplementary Table 4).
Stages of literature review
To maximize the number of potential studies included in the review, the initial searches were completed in three electronic bibliographic databases: (1) PubMed (https://pubmed.ncbi.nlm.nih.gov/); (2) EMBASE (https://www.embase.com/); and (3) Web of Science (https://login.webofknowledge.com/). The searches were limited to human case–control studies comparing the genome-wide transcriptome of individuals diagnosed with schizophrenia to healthy controls published between January 1, 2000 and June 30, 2020 in the English language. The schizophrenia case definition included patients with schizophrenia as recognized using any standard diagnostic criteria (Diagnostic and Statistical Manual of Mental Disorders (DSM), International Classification of Diseases (ICD), or clinician diagnosis). Study participants were considered healthy controls if they did not have a diagnosed mental disorder. The search syntax was customized for each database: PubMed: “Schizophrenia” [MeSH] and (“Transcriptome” [MeSH] or “Transcription” [MeSH] or “Sequence Analysis, RNA” [MeSH] or “Microarray Analysis” [MeSH]) and “humans” [MeSH Terms] and “case–control studies” [MeSH Terms]) and “humans” [MeSH Terms]; EmBase: (“schizophrenia”/exp OR schizophrenia) and (“transcriptome”/exp or transcriptome or “microarray analysis”/exp or “microarray analysis” or “RNA sequence”/exp or “RNA sequence” or “gene expression”/exp or “gene expression”) and (“case control study”/exp or “case control study”) and “human”/de and “article”/it; Web of Science: TS = (Schizophr* AND (Transcriptome or Transcription or “Microarray Analysis” or “RNA sequence”))) and LANGUAGE: (English) and DOCUMENT TYPES: (Article) Refined by: TOPIC: (case control) Timespan: 2000–2020.
After the literature searches were completed, the references were downloaded to the Rayyan systematic review web app [23] where duplicate references were identified and removed. Next, the titles and abstracts of the references were screened by two reviewers who assessed whether they met the inclusion and exclusion criteria. Any discrepancy in inclusion decisions was resolved by a third researcher.
Articles that met the inclusion criteria after the title and abstract review were then reviewed in full by three blind reviewers and assessed for inclusion in data extraction and analysis. Inclusion discrepancies were resolved through discussion and consensus of involved researchers.
Data were extracted from each study that met the a priori inclusion criteria. Specifically, sample size, including the number of cases and controls; diagnostic methodology; effect size, type, and statistical significance per gene; type of tissue in which gene expression was measured; technology used to quantify gene expression; demographic and other covariates, including sex, age, genetic ancestry, smoking status; and major findings from the study were extracted into a standardized data collection form.
Two of the reviewers assessed the risk of bias and quality in the included studies by considering the guidelines provided by the adapted Systematic Omics Analysis Review (SOAR) assessment [24]. Only a subset of the SOAR criteria applied to the studies presented here, notably, the “Preliminary Questions”, “Experimental Design”, “Human Subjects”, and “Microarray Data” sections. All studies provided human subjects information, but to our knowledge, none provided raw data files, and the data do not appear to be available in public repositories. However, data files may be available if requested from the authors.
Data integration and analysis
To harmonize the data, gene nomenclature was standardized to HUGO Gene Nomenclature (https://www.genenames.org/) for each study. Each gene was classified as being up- or downregulated in cases as compared to controls, and statistically significant or not as defined by the authors of the study.
The data from each study were then combined into a single analytic file. The frequency of each statistically significant gene was calculated, and whether it was consistently up or downregulated in all studies was noted. Genes found to be differentially expressed in multiple studies were then compared to findings from the largest schizophrenia genome-wide association study (GWAS) to date (90 cohorts including 67,390 cases and 94,015 controls [25]) and to well-established schizophrenia-associated recurrent and non-recurrent CNVs [26]. We also queried genes found to be differentially expressed in multiple studies in the schizophrenia exome meta-analysis consortium database (SCHEMA, https://schema.broadinstitute.org/). SCHEMA aggregates high-throughput sequencing data of patients with schizophrenia and recorded whether these genes were found to be associated with specific classes of loss-of-function variants [27].
To examine differentially expressed genes in a biological context, we utilized the GENE2FUNC tool on the Functional Mapping and Annotation of Genome-Wide Association Studies (FUMA GWAS, https://fuma.ctglab.nl/) website [28]. This tool aggregates data from the GTEx project and allows the user to visualize the expression of a gene in various body tissues. Here, we obtained the average expression value per tissue type for each gene and the average of normalized expression per tissue type.
Defining statistical significance
Each study tested a different number of transcripts, and several different statistical significance or false-discovery rate thresholds were applied. In many cases, the exact number of transcripts tested was not specified, making it difficult to rigorously evaluate the likelihood of differential transcription being reported for the same transcript by multiple studies. For purposes of estimation, we assumed testing of on the order of 20,000 transcripts in each study and we varied significance thresholds from a nominal 0.01 to a stricter 0.001. We then used a binomial model to assess the probability of a transcript being reported significant by three or more studies under the null model of no association with schizophrenia.
Results
As shown in the PRISMA flow diagram (Fig. 1), 415 records were identified through database searching. Of the 415, 11 were duplicate articles and were excluded before further consideration. Next, a total of 404 titles and abstracts were screened by two researchers to check if they met the study inclusion criteria (i.e., human case–control, noninterventional studies in English comparing the transcriptome of individuals diagnosed with schizophrenia to healthy controls). 366 abstracts did not meet the study inclusion criteria and were excluded. Articles were often excluded for more than one reason, and most excluded articles were candidate gene studies or not genome-wide (n = 334). Others were excluded because they did not meet the study criteria of examining a case–control comparison (n = 190); were completed in a non-schizophrenia sample (e.g., bipolar disorder patients) (n = 172); or did not actually examine gene expression (n = 26). Then, 12 of the 38 full-text articles that were assessed for data extraction eligibility did not meet study criteria and were excluded from further consideration. Specifically, five did not report a direct case–control comparison (Ellis et al. [29], Hagihara et al. [30], Kakiuchi et al. [31], Manchia et al. [32], Viana et al. [33]), one included previously reported data (Haroutunian et al. [34]), three were not genome-wide (Huang et al. [35], Katsel et al. [36], van Beveren et al. [37]), two did not report individual genes (Narla et al. [38], Yu et al. [39]) and one only reported results for bipolar disorder (Konradi et al. [40]). The remaining 26 articles were included in the data extraction stage, and were considered for synthesis, and qualitative review.
Descriptive synthesis
Table 1 itemizes the studies included in our qualitative review. All studies reported in this review met the SOAR criteria for assessing quality and bias in omics studies reported above. A total of 18 studies assessed gene expression using microarrays, six utilized RNAseq and two used imputed gene expression based on genotype. In most of the studies (n = 12), gene expression was quantified from postmortem brain tissue (n = 2 amygdala; n = 4 dorsolateral prefrontal cortex (DLPFC); n = 3 hippocampus; n = 2 left hemisphere superior temporal cortex (STC); and n = 1 gyrus (STG)). Other studies measured transcription in blood samples (n = 10), induced pluripotent stem cell (IPSC)-derived neurons (n = 1), lymphoblastoid cell lines (LCLs) (n = 2), or skin fibroblasts (n = 1). One article (Collado-Torres et al. [41]) used data from the GTEx project to impute expression levels in two brain regions, so we have included these analyses as separate studies. The format for reporting effect size varied across studies, including beta, fold change or ratio, and fragments per kilobase per million mapped reads (FPKM). Correction for multiple comparisons also varied across studies, including a conservative Bonferroni, false-discovery rate (FDR), or no stated correction.
The mean age of participants in each study ranged from 14 to 69 years in schizophrenia cases and 14 to 65 years in healthy controls. Included studies ranged from 0 to 75% female for both schizophrenia cases and healthy controls. The sample size varied substantially from as few as three cases and three controls [42] to over 40,000 cases and 65,000 controls [43]. In general, most studies had case/control samples that were well matched on age and sex, but there were a few studies where there was greater than the 10-year age difference between cases and controls (e.g., Collado-Torres et al. [41] and Wu et al. [44]). In a few instances, there was a substantial difference in the sex ratio between cases and controls, for example in the work presented by Schmitt et al. [45] and Sellmann et al. [46] the controls were 20% female, while the cases were 50% female.
Replicated genes and direction of effect
Six articles that met full-text inclusion criteria were later excluded from data synthesis because the data could not be standardized or were unique to the individual study (Bousman et al. [47]; Chen et al. [42]; Gardiner et al. [48]; Wei et al. [49]; Yu et al. [39]; Zhang et al. [50]). Specifically, two studies completed pathway analyses, examining genes in aggregate instead of studying individual genes. Unfortunately, each of these studies used different pathway nomenclature; Bousman et al. [47] examined Ingenuity pathways, the only study to do so, while Gardiner [48] examined KEGG pathways. Thus, it was unclear whether findings overlapped between studies. Four other studies interrogated long non-coding RNAs (lncRNA, Chen et al. [42]) or microRNAs (miRNA, Wei et al. [49]; Yu et al. [39]; Zhang et al. [50]). Despite the growing evidence for their role in the regulation of gene expression [51, 52], and the potential use of lncRNAs as biomarkers for neuropsychiatric disorders [53], the studies that examined these variants were not included in the data synthesis presented here, because they used unique transcript identifiers that could not be directly mapped onto gene symbols or to one another.
Of the 20 remaining articles (21 studies), a total of 6771 unique genes were reported as statistically significantly differentially expressed in schizophrenia cases; 5566 reported in one study, 1045 in two, 138 reported in three, 21 in four, and one in five studies (Supplementary Table 1). Three studies reported that no genes reached statistical significance after correction for multiple comparisons (Schmitt et al. [45]; Sellmann et al. [46]; Tian et al. [54]). To narrow the focus of the results presented here, we chose to concentrate our analyses on the 160 genes that were reported three or more times across studies, as itemized in Table 2.
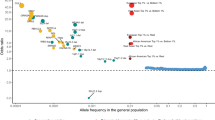
Of the top 160 genes, the majority of replicated findings were inconsistent in their reported direction of effect (n = 108 genes). This finding did not appear to follow a pattern based on the origin tissue (as shown in Fig. 2 and Supplementary Fig. 1) or the expression measurement technology employed (Supplementary Fig. 2). The GBP2 gene, which appeared in five studies, was reported to be upregulated in individuals with schizophrenia in four studies and downregulated in one. Of the 21 genes reported as significant by four studies, 19 had inconsistent directions of effect. Of the 138 genes that appeared in three studies, 88 had the inconsistent direction of effect. Of the 50 that had a consistent direction of effect across studies, most genes (n = 28) were upregulated.
Genes that were reported as having different expression levels in individuals with and without schizophrenia in four (blue) or five (purple) studies are shown, with the lighter color representing consistent direction of effect and the darker color representing inconsistent directions of effect across the indicated studies. The first panel shows the studies that measured expression in blood samples, the middle panel shows studies in which expression was measured in brain tissue, and the third panel shows studies completed in other tissue types (skin fibroblasts, lymphoblastoid cell lines).
The finding of an association of gene expression of the GBP2 gene in five studies is greater than expected by chance. Assuming independence between the 21 sets of results reported in the 20 articles, in the absence of any true associations, the probability under a binomial model of three or more studies reporting the same transcript as significant would be 0.0012 for a permissive significance threshold of 0.01, and 1.3 × 10-6 at a stricter P value of 0.001. Given testing of ~20,000 genes, this results in an expectation of 24 or <1 such transcript significant by chance at the respective thresholds. The probability of four or more studies reporting the same transcript at a significance threshold of 0.001 is 5.9 × 10−9. Collectively, more genes are reported by multiple studies than would be expected by chance assuming no association.
Overlap with previous research
Of the genes that were found in three or more studies, six have been previously reported in the largest schizophrenia case/control GWAS to date: [25] BNIP3L, CD47, CHRNA2, GATAD2A, TCF4, and TYW5. Eight genes reported as differentially expressed by three or more studies were in regions that overlap with schizophrenia-associated CNVs. PPP1R2 is in the 3q29 deletion region; HSPB1 is in the 7q11.23 duplication region; two genes (INO80E and YPEL3) are in the 16p11.2 duplication region; DHRS11 is in 17q12 deletion region; and three genes (SEPT5, RTN4R, SLC2A11) are in the 22q11 deletion region. Generally, the genes in duplication regions were reported as upregulated, and genes in deletion regions were downregulated. The exception is YPEL3, which was more often downregulated in the studies reviewed here despite being in the 16p11.2 duplication region.
Of the 160 genes reported as significantly differentially expressed in three or more studies, none showed associations with rare variants in the SCHEMA data after correction for multiple comparisons. However, six genes had rare variants nominally associated with schizophrenia with meta-analysis P values <0.05 (ATP1B1, CDKN1A, FAM8A1, FBXO32, PDGFRB, SPHK2), while only one, SPHK2, demonstrated a meta-analysis p-value less than 0.01 (P = 0.00801). There were five protein-truncating or putatively loss-of-function variants (PTVs): ATP1B1, CDKN1A, FAM8A1, FBXO32, and SPHK2; and three damaging missense variants: CDKN1A, PDGFRB, and SPHK2.
Figure 3 shows the collective expression patterns of all 160 genes in 54 different tissue types using GTEx v8 data. Tissues with significant enrichment at Bonferroni corrected P value ≤0.05 are shown in red. This correction is completed separately for up-, down-, and both-sided sets. Here, we can see that when considering both up- and downregulated genes, our list of genes reported in schizophrenia gene expression studies is shown to be enriched for genes differentially expressed in the brain as compared to other tissues. The list is also enriched for genes differentially expressed in the pancreas, subcutaneous adipose tissue, whole blood, liver, and lymphocytes.
The -log 10 P value is provided for each of the 54 tissue types of the Genotype-Tissue Expression (GTEx) project v8 data, indicating whether the list of 160 genes reported as differentially expressed in individuals with schizophrenia is enriched for genes expressed in that tissue. Tissues with significant enrichment at a Bonferroni corrected P value < 0.05 are shown in red. The top panel shows upregulated differentially expressed genes (DEG), the middle panel shows downregulated DEG, and the bottom panel shows DEG in both directions.
Discussion
This review summarizes the literature on gene expression in schizophrenia and demonstrates the surprisingly small overlap in the genes reported across studies. Only 26 studies met our a priori inclusion criteria and were described here. The results of this review were unexpected, in that few genes were found in more than three studies, and the reported direction of effect was so variable. It was hoped that gene expression would help to explain the large number of genome-wide associated variants that are not found in genes and are theorized to be regulatory. With some exceptions described below, gene expression does not implicate the same genes that have been found by GWAS, CNV studies, or exome studies via SCHEMA. Yet, we did find that some of these top genes recapitulated findings from other types of studies in schizophrenia, as described below. Given the limited overlap with previous work, it appears that gene expression is providing us with new and different information about schizophrenia rather than bolstering previous work. This highlights the importance of the standardization of additional studies to clarify the associations between gene expression and schizophrenia.
The gene that was reported most often as differentially expressed in case–control studies of schizophrenia, GBP2, was only reported in five studies and was shown to be upregulated in four studies and downregulated in one. GBP2, guanylate-binding protein 2, is in the guanine-binding protein family, and is involved in interferon-gamma signaling and the innate immune system [55]. GBP2 has been associated with migraine [56], breast [57], and other cancers [58, 59], and HIV-1 infection [60], but has not been previously associated with psychiatric or brain/behavior phenotypes. The recent finding implicating GBP2 as a potential prognostic biomarker for pancreatic adenocarcinoma [58], could provide a model for future inquiry into its role in schizophrenia. Guanylate-binding proteins (GBPs) more broadly have been implicated in the body’s defense against infections, specifically viruses [58], and the major histocompatibility complex (MHC) has been implicated in GWAS of schizophrenia [61]. Based on maternal infection as a well-established risk factor for schizophrenia [62], a potential mechanism for the role of GBP2 in schizophrenia could be its role in reactivity to viral exposures, perhaps through epigenetic effects. This suggests that investigation of immune-related phenotypes may be useful.
Two genes that were found in four studies to have a differential expression with a consistent direction of effect, Interferon-induced transmembrane protein 3 (IFITM3) and KRAS Proto-Oncogene, GTPase (KRAS), involve immune-related function. IFITM3 has been implicated in immune function, specifically in viral restriction [63], and has been associated with cortical immune activation and inflammation, as well as neurodevelopment [64,65,66]. KRAS codes for K-Ras which is involved in cell proliferation, maturation, and differentiation [67]. KRAS is also involved in signal transduction and has been associated with a host of cancers [68,69,70] and Noonan Syndrome [71].
Several of the genes that were found to be differentially expressed in our review have also been found in the largest GWAS of schizophrenia: BNIP3L, CD47, CHRNA2, GATAD2A, TCF4, and TYW5, and have also been associated with other conditions. For example, all of these genes except TYW5 have been associated with various cancers [72,73,74,75,76,77], BNIP3L and CD47 have been associated with immune function [78] or infection [79], and CHRNA2 and TCF4 have been associated with other neuropsychiatric conditions and substance use [80, 81]. GATAD2A has been associated with cardiometabolic disease and type 2 diabetes [82, 83], which is pertinent because of the increased risk of metabolic syndrome secondary to the use of atypical antipsychotic medications in patients with schizophrenia [84].
The CNV findings in this study also may have important etiologic relevance for both schizophrenia as well as its core features and associations with other conditions. Five schizophrenia-associated CNV regions contained genes that were found to be differentially expressed in this study: PPP1R2 in 3q29, HSPB1 in 7q11.23, INO80E and YPEL3 in 16p11.2, DHRS11 in 17q12, and SEPT5, RTN4R, and SLC2A11 in 22q11.2. Though these CNVs have wide-ranging phenotypic presentations, CNVs in these regions are all associated with developmental delays and intellectual disability. They are also associated with neuropsychiatric phenotypes in addition to schizophrenia, notably anxiety (3q29, 7q11.23, 17q12), autism spectrum disorder (ASD; 3q29, 7q11.23, 16p11.2, 17q12, 22q), attention-deficit/hyperactivity disorder (ADHD; 7q11.23, 22q), and bipolar disorder (3q29, 7q11.23, 17q12). The physical manifestations of these CNVs also share some commonalities: 3q29, 7q11.23, 16p11.2 and 22q are all associated with cardiac problems and varying craniofacial dysmorphology, while 17q12 and 22q are both associated with immune system dysfunction. Studies have examined some of these comorbid conditions explicitly such as the demonstration of pleiotropic effects of the INO80E gene in 16p11.2 on both schizophrenia and cardiometabolic disease [82]. The most well-known CNV associated with an increased risk of schizophrenia is the 22q deletion syndrome, where ~25% of cases go on to develop schizophrenia [85]. The 22q deletion syndrome is one of the strongest risk factors for schizophrenia because of its high prevalence, thereby warranting emphasis in future studies [86].
After querying the 160 differentially expressed genes in SCHEMA, only one gene, SPHK2, showed suggestive evidence of association in the SCHEMA database at P < 0.01. The associated variant is classified in the database as a protein-truncating variant (PTV; stop-gained, frameshift, and essential splice donor or acceptor variants), which suggests that the effect on schizophrenia most likely tracks with decreasing expression of the gene [87]. SPHK2 mediates cellular migration, proliferation, and apoptosis, and may be responsible for promoting angiogenesis and tumorigenesis in some cancers. Multiple isoforms of SPHK2 have been observed from alternatively spliced transcript variants [88].
As we would expect, the genes found in this study appear to be differentially expressed in the brain when compared to other body tissues. However, there are some unexpected results; notably, differential expression in the pancreas, subcutaneous adipose tissue, whole blood, liver, and lymphocytes. It is possible that the differential expression in blood, brain, and lymphocytes is due in some part to these being the tissues assayed in the transcriptome abundance studies summarized here. However, the finding of differential expression in the pancreas, adipose tissue, and liver is interesting given that metabolic syndrome is often associated with schizophrenia [84]. It would be valuable to investigate whether this effect was secondary to medication, but only one of the included studies covaried for medication [89]. Although several studies have used FUMA [28] to annotate genetic variants with functional information from GWAS summary statistics (e.g., Liu et al. [82] and Wendt et al. [90]) we did not find that others had used the GENE2FUNC module to examine tissue expression in schizophrenia.
Recommendations for future research
Greater consensus on standard formats for reporting data and results
One of the greatest challenges in completing the work presented in this review was the inconsistency in the way the results of gene expression studies were reported. A great deal of effort was devoted to convert the results to a standard nomenclature before the data could be aggregated for analysis. Different statistical significance thresholds and corrections for multiple comparisons were applied, and at times the number of transcripts evaluated was not reported such that it could not be determined if the correction for multiple comparisons was appropriate. These manuscripts were published over several years, and there has not yet been a coalescence of standards of reporting in the field. Efforts to standardize methods and reporting, similar to those for GWAS [91], would help to enhance the interpretation and discernment of common findings in future studies. It would also be beneficial for the field to adopt standard naming conventions and a well-defined statistical significance threshold for the transcriptome.
Better definition of covariates
There are several factors that may impact gene expression [16] that should be included as covariate in gene expression studies, notably, age, sex, race, or genetic ancestry principal components, medication usage, smoking status, and batch. In addition, for the postmortem brain studies, the cause of death and postmortem interval may be relevant. Few of the studies reviewed considered all of these factors (as shown in Supplementary Table 2). Only one study [92] considered medication use and smoking status as covariates. Smoking status is an important covariate in the present context given that patients with schizophrenia are more likely to smoke than are healthy controls (>60% versus ~17%) [93], and gene expression is known to differ between those who smoke and those who do not [94]. When the top 160 genes from this study were compared to the 1270 genes differentially expressed between current and never smokers [95], 24 were common to both studies, including GBP2 (Supplementary Table 3). This may mean that had these studies controlled for smoking status, the results presented, and conclusions drawn from the work may have been different.
Case/control definition
The definition of cases ranged from a statement that the participant population included “schizophrenia cases” to more stringent definitions based on multiple clinician assessments of ICD or DSM criteria. The selection of the control samples was also highly variable and often not specified. Many of the studies did not state the source of controls, nor whether they were screened or interviewed to rule out schizophrenia or another major mental disorder. In future meta-analytic studies, it would also be interesting to examine longitudinal changes in gene expression between the first episode and long-standing schizophrenia (as in Ota et al. [96]), as well as the specificity of the findings with respect to schizophrenia versus other mental disorders such as bipolar disorder.
An additional consideration in examining this body of literature is the challenge of differentiating whether the participants were included in more than one study, especially in the research utilizing postmortem brain tissue. For example, at least two studies [43, 97] used postmortem brain tissue from the CommonMind consortium [98]. There are several other brain banks that are commonly used, for example, the Maryland Brain Collection [99], the Stanley Neuropathology Consortium (SNC [92]), and the Stanley Medical Research Institute (SMRI [100]). Though there does not appear to be sample overlap in the studies reviewed here, it is likely that other studies do use these data.
Expression technology
Some of the variability in the results reported here may be the result of the use of different gene expression measurement technologies. Of course, some of these will change over time, but standardization of RNA-seq data processing pipelines (for the studies that use this technology) may lead to a greater coalescence of the reported results [101]. In addition, it has been reported that there is bias in the publication of gene expression studies; notably, that genes are more likely to appear in the literature if they are over-expressed, found to be associated in multiple diseases, and are more “popular” in the biomedical literature [9]. Moreover, one might expect greater overlap in the studies that used imputed data, as this approach implicitly controls for environmental sources of variation. However, there was a lack of synergy across the studies that used imputed gene expression.
Standardized reporting of pathways
The pathway analytic results reported in the studies reviewed here were not synthesized due to the number of different approaches employed. Although articles such as Bousman et al. [47] and Wei et al. [49] identified pathways that were differentially expressed in patients with schizophrenia, their use of different pathway nomenclature made comparisons across these studies difficult. Unfortunately, Gene Ontology (http://geneontology.org/), Ingenuity (www.ingenuity.com), and KEGG (https://www.genome.jp/kegg/) pathway analyses were completed and the results from each of these are not directly comparable to one another [102]. In addition, the lack of open-source programs that can convert between gene pathway nomenclature made it nearly impossible to standardize data from multiple authors. Of course, the differences between KEGG, Ingenuity, Gene Ontology, and Reactome are not purely ones of nomenclature but also due to the different data underlying pathway definitions, e.g., whether they are based on biological pathways, ontological gene-set pathways, or statistically associated entities. Therefore, the pathway definitions will subtly differ across these databases, so reporting the results from multiple pathway approaches in the same manuscript may be a better approach to facilitating comparisons across studies. Researchers could also include information about which genes in the pathway have the strongest signal, potentially allowing for gene-level comparisons in ancillary work derived from pathway analysis. Nomenclature standardization and reporting results across techniques would allow larger meta-analyses to occur and offer a clearer picture of the differentially expressed gene pathways in patients with schizophrenia.
Public repositories
One area for future consideration for schizophrenia gene expression reporting is the lack of publicly available results repositories. Given the wide variety of GWAS repositories (e.g., the database of Genotypes and Phenotypes (dbGaP), https://www.ncbi.nlm.nih.gov/gap/; the NHGRI-EBI GWAS Catalog, https://www.ebi.ac.uk/gwas/), it was interesting to note that transcriptome-wide association study (TWAS) repositories such as the Gene Expression database of Normal and Tumor tissues (GENT2, http://gent2.appex.kr/gent2/) and the Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/) contained solely experimental data but not results. The creation of publicly shared gene expression result repositories would facilitate widespread sharing of information between lab groups and allow for much easier meta-analyses. Similarly, such a repository would help lead to a standardized format of reporting results.
Strengths and limitations
The main strengths of this work are our exhaustive search of multiple databases, and interpretation of the replicated results across different types of genetic associations (i.e., CNVs, GWAS, exome sequencing). Additional strengths include the examination of concordance of the results across tissues and assay methods. The primary limitation is that we were unable to complete a meta-analysis of gene expression in schizophrenia due to differences in the study methodology (i.e., selection of cases, convenience versus selected controls), analytic methods (i.e., quantification of case–control differences, control for multiple testing, inclusion of confounders, statistical significance thresholds), and gene expression methodology (i.e., gene nomenclature, measures of gene expression, different tissue types, etc.).
Conclusion
To our knowledge, this is the first systematic review and synthesis of controlled studies of gene expression in schizophrenia. This work adds to the body of knowledge that demonstrates differential gene expression between schizophrenia cases and controls. Although case–control differences were reported in numerous studies, only one gene, GBP2, was replicated in five studies. This suggests that further inquiry into the potential role of GBP2 could have promise in advancing our understanding of the genetic architecture of schizophrenia. These findings also highlight the methodological inconsistencies that impede valid meta-analyses and synthesis across studies. Standardization of the use of covariates, gene nomenclature, and methods for reporting results could enhance our understanding of the potential mechanisms through which genes exert their influence on the etiology of schizophrenia.
References
Owen MJ, Sawa A, Mortensen PB. Schizophrenia. Lancet. 2016;388:86–97.
Merikangas AK, Almasy L. Using the tools of genetic epidemiology to understand sex differences in neuropsychiatric disorders. Genes Brain Behav. 2020;19:e12660.
Rees E, Kirov G. Copy number variation and neuropsychiatric illness. Curr Opin Genet Dev. 2021;68:57–63.
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7.
Hayashi-Takagi A, Vawter MP, Iwamoto K. Peripheral biomarkers revisited: integrative profiling of peripheral samples for psychiatric research. Biol Psychiatry. 2014;75:920–8.
Blackstock JC. Gene expression. In: Blackstock JC, editor. Guide to biochemistry. Oxford: Butterworth-Heinemann; 1989. p. 208–21.
Volgin DV. Gene Expression: analysis and quantitation. In: Verma AS, Singh A, editors. Animal biotechnology. San Diego: Academic Press; 2014. p. 307–25.
Gamazon ER, Stranger BE. The impact of human copy number variation on gene expression. Brief Funct Genomics. 2015;14:352–7.
Rodriguez-Esteban R, Jiang X. Differential gene expression in disease: a comparison between high-throughput studies and the literature. BMC Med Genomics. 2017;10:59.
Swarbrick S, Wragg N, Ghosh S, Stolzing A. Systematic review of miRNA as biomarkers in Alzheimer’s disease. Mol Neurobiol. 2019;56:6156–67.
Blok EJ, Bastiaannet E, van den Hout WB, Liefers GJ, Smit V, Kroep JR, et al. Systematic review of the clinical and economic value of gene expression profiles for invasive early breast cancer available in Europe. Cancer Treat Rev. 2018;62:74–90.
Maertzdorf J, Repsilber D, Parida SK, Stanley K, Roberts T, Black G, et al. Human gene expression profiles of susceptibility and resistance in tuberculosis. Genes Immun. 2011;12:15–22.
Darby MM, Yolken RH, Sabunciyan S. Consistently altered expression of gene sets in postmortem brains of individuals with major psychiatric disorders. Transl Psychiatry. 2016;6:e890.
Ramaker RC, Bowling KM, Lasseigne BN, Hagenauer MH, Hardigan AA, Davis NS, et al. Post-mortem molecular profiling of three psychiatric disorders. Genome Med. 2017;9:72.
Kumarasinghe N, Tooney PA, Schall U. Finding the needle in the haystack: a review of microarray gene expression research into schizophrenia. Aust N. Z J Psychiatry. 2012;46:598–610.
Mele M, Ferreira PG, Reverter F, DeLuca DS, Monlong J, Sammeth M, et al. Human transcriptome across tissues and individuals. Science. 2015;348:660–5.
GTEx Consortium. The genotype-tissue expression (GTEx) project. Nat Genet. 2013;45:580–5.
Bray NJ. Gene expression in the etiology of schizophrenia. Schizophr Bull. 2008;34:412–8.
Harrison PJ, Weinberger DR. Schizophrenia genes, gene expression, and neuropathology: on the matter of their convergence. Mol Psychiatry. 2005;10:40–68.
Iwamoto K, Kato T. Gene expression profiling in schizophrenia and related mental disorders. Neuroscientist. 2006;12:349–61.
Mirnics K, Middleton FA, Lewis DA, Levitt P. Analysis of complex brain disorders with gene expression microarrays: schizophrenia as a disease of the synapse. Trends Neurosci. 2001;24:479–86.
What genes are differentially expressed in individuals with schizophrenia? PROSPERO 2020 CRD42020132433. 2020. https://www.crd.york.ac.uk/prospero/display_record.php?ID=CRD42020132433. Accessed 27 July 2021.
Ouzzani M, Hammady H, Fedorowicz Z, Elmagarmid A. Rayyan-a web and mobile app for systematic reviews. Syst Rev. 2016;5:210.
McConnell ER, Bell SM, Cote I, Wang RL, Perkins EJ, Garcia-Reyero N, et al. Systematic Omics Analysis Review (SOAR) tool to support risk assessment. PLoS ONE. 2014;9:e110379.
Schizophrenia Working Group of the Psychiatric Genomics Consortium, Ripke S, Walters JT, O’Donovan MC. Mapping genomic loci prioritises genes and implicates synaptic biology in schizophrenia. medRxiv:2020.2009.2012.20192922 [Preprint]. 2020. Available from: https://www.medrxiv.org/content/10.1101/2020.09.12.20192922v1.
Rees E, O’Donovan MC, Owen MJ. Genetics of schizophrenia. Curr Opin Behav Sci. 2015;2:8–14.
Singh T, Neale BM, Daly MJ, Consortium obotSEM-A. Exome sequencing identifies rare coding variants in 10 genes which confer substantial risk for schizophrenia. medRxiv: 2020.2009.2018.20192815. [Preprint]. 2020. Available from: https://www.medrxiv.org/content/10.1101/2020.09.18.20192815v1.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Ellis SE, Panitch R, West AB, Arking DE. Transcriptome analysis of cortical tissue reveals shared sets of downregulated genes in autism and schizophrenia. Transl Psychiatry. 2016;6:e817.
Hagihara H, Ohira K, Takao K, Miyakawa T. Transcriptomic evidence for immaturity of the prefrontal cortex in patients with schizophrenia. Mol Brain. 2014;7:41.
Kakiuchi C, Ishiwata M, Nanko S, Ozaki N, Iwata N, Umekage T, et al. Up-regulation of ADM and SEPX1 in the lymphoblastoid cells of patients in monozygotic twins discordant for schizophrenia. Am J Med Genet B Neuropsychiatr Genet. 2008;147B:557–64.
Manchia M, Piras IS, Huentelman MJ, Pinna F, Zai CC, Kennedy JL, et al. Pattern of gene expression in different stages of schizophrenia: down-regulation of NPTX2 gene revealed by a meta-analysis of microarray datasets. Eur Neuropsychopharmacol. 2017;27:1054–63.
Viana J, Pidsley R, Troakes C, Spiers H, Wong CC, Al-Sarraj S, et al. Epigenomic and transcriptomic signatures of a Klinefelter syndrome (47,XXY) karyotype in the brain. Epigenetics. 2014;9:587–99.
Haroutunian V, Katsel P, Dracheva S, Stewart DG, Davis KL. Variations in oligodendrocyte-related gene expression across multiple cortical regions: implications for the pathophysiology of schizophrenia. Int J Neuropsychopharmacol. 2007;10:565–73.
Huang KC, Yang KC, Lin H, Tsao TT, Lee SA. Transcriptome alterations of mitochondrial and coagulation function in schizophrenia by cortical sequencing analysis. BMC Genomics. 2014;15:S6.
Katsel P, Davis KL, Gorman JM, Haroutunian V. Variations in differential gene expression patterns across multiple brain regions in schizophrenia. Schizophr Res. 2005;77:241–52.
van Beveren NJ, Buitendijk GH, Swagemakers S, Krab LC, Roder C, de Haan L, et al. Marked reduction of AKT1 expression and deregulation of AKT1-associated pathways in peripheral blood mononuclear cells of schizophrenia patients. PLoS ONE. 2012;7:e32618.
Narla ST, Lee YW, Benson CA, Sarder P, Brennand KJ, Stachowiak EK, et al. Common developmental genome deprogramming in schizophrenia—role of Integrative Nuclear FGFR1 Signaling (INFS). Schizophr Res. 2017;185:17–32.
Yu HC, Wu J, Zhang HX, Zhang GL, Sui J, Tong WW, et al. Alterations of miR-132 are novel diagnostic biomarkers in peripheral blood of schizophrenia patients. Prog Neuropsychopharmacol Biol Psychiatry. 2015;63:23–29.
Konradi C, Eaton M, MacDonald ML, Walsh J, Benes FM, Heckers S. Molecular evidence for mitochondrial dysfunction in bipolar disorder. Arch Gen Psychiatry. 2004;61:300–8.
Collado-Torres L, Burke EE, Peterson A, Shin J, Straub RE, Rajpurohit A, et al. Regional heterogeneity in gene expression, regulation, and coherence in the frontal cortex and hippocampus across development and schizophrenia. Neuron. 2019;103:203–16. e208
Chen S, Sun X, Niu W, Kong L, He M, Li W, et al. Aberrant expression of long non-coding RNAs in schizophrenia patients. Med Sci Monit. 2016;22:3340–51.
Huckins LM, Dobbyn A, Ruderfer DM, Hoffman G, Wang W, Pardinas AF, et al. Gene expression imputation across multiple brain regions provides insights into schizophrenia risk. Nat Genet. 2019;51:659–74.
Wu JQ, Green MJ, Gardiner EJ, Tooney PA, Scott RJ, Carr VJ, et al. Altered neural signaling and immune pathways in peripheral blood mononuclear cells of schizophrenia patients with cognitive impairment: a transcriptome analysis. Brain Behav Immun. 2016;53:194–206.
Schmitt A, Leonardi-Essmann F, Durrenberger PF, Wichert SP, Spanagel R, Arzberger T, et al. Structural synaptic elements are differentially regulated in superior temporal cortex of schizophrenia patients. Eur Arch Psychiatry Clin Neurosci. 2012;262:565–77.
Sellmann C, Villarin Pildain L, Schmitt A, Leonardi-Essmann F, Durrenberger PF, Spanagel R, et al. Gene expression in superior temporal cortex of schizophrenia patients. Eur Arch Psychiatry Clin Neurosci. 2014;264:297–309.
Bousman CA, Chana G, Glatt SJ, Chandler SD, Lucero GR, Tatro E, et al. Preliminary evidence of ubiquitin proteasome system dysregulation in schizophrenia and bipolar disorder: convergent pathway analysis findings from two independent samples. Am J Med Genet B Neuropsychiatr Genet. 2010;153B:494–502.
Gardiner E, Beveridge NJ, Wu JQ, Carr V, Scott RJ, Tooney PA, et al. Imprinted DLK1-DIO3 region of 14q32 defines a schizophrenia-associated miRNA signature in peripheral blood mononuclear cells. Mol Psychiatry. 2012;17:827–40.
Wei H, Yuan Y, Liu S, Wang C, Yang F, Lu Z, et al. Detection of circulating miRNA levels in schizophrenia. Am J Psychiatry. 2015;172:1141–7.
Zhang F, Xu Y, Shugart YY, Yue W, Qi G, Yuan G, et al. Converging evidence implicates the abnormal microRNA system in schizophrenia. Schizophr Bull. 2015;41:728–35.
Corcoran R, Mercer G, Frith CD. Schizophrenia, symptomatology and social inference: investigating “theory of mind” in people with schizophrenia. Schizophr Res. 1995;17:5–13.
Liu B, Li J, Cairns MJ. Identifying miRNAs, targets and functions. Brief Bioinform. 2014;15:1–19.
van den Berg MMJ, Krauskopf J, Ramaekers JG, Kleinjans JCS, Prickaerts J, Briede JJ. Circulating microRNAs as potential biomarkers for psychiatric and neurodegenerative disorders. Prog Neurobiol. 2020;185:101732.
Tian T, Wei Z, Chang X, Liu Y, Gur RE, Sleiman PMA, et al. The long noncoding RNA landscape in amygdala tissues from schizophrenia patients. EBioMedicine. 2018;34:171–81.
GeneCards: GBP2 Gene. https://www.genecards.org/cgi-bin/carddisp.pl?gene=GBP2&keywords=GBP2, 1996–2021. Accessed 27 July 2021.
Jiang Y, Wu R, Chen C, You ZF, Luo X, Wang XP. Six novel rare non-synonymous mutations for migraine without aura identified by exome sequencing. J Neurogenet. 2015;29:188–94.
Godoy P, Cadenas C, Hellwig B, Marchan R, Stewart J, Reif R, et al. Interferon-inducible guanylate binding protein (GBP2) is associated with better prognosis in breast cancer and indicates an efficient T cell response. Breast Cancer. 2014;21:491–9.
Liu B, Huang R, Fu T, He P, Du C, Zhou W, et al. GBP2 as a potential prognostic biomarker in pancreatic adenocarcinoma. PeerJ. 2021;9:e11423.
Song LB, Luan JC, Zhang QJ, Chen L, Wang HY, Cao XC, et al. The identification and validation of a robust immune-associated gene signature in cutaneous melanoma. J Immunol Res. 2021;2021:6686284.
Srinivasachar Badarinarayan S, Shcherbakova I, Langer S, Koepke L, Preising A, Hotter D, et al. HIV-1 infection activates endogenous retroviral promoters regulating antiviral gene expression. Nucleic Acids Res. 2020;48:10890–908.
Mokhtari R, Lachman HM. The major histocompatibility complex (MHC) in schizophrenia: a review. J Clin Cell Immunol. 2016;7:479–85.
Brown AS. Exposure to prenatal infection and risk of schizophrenia. Front Psychiatry. 2011;2:63.
GeneCards: IFITM3 Gene. https://www.genecards.org/cgi-bin/carddisp.pl?gene=IFITM3, 1996-2021. Accessed 27 July 2021.
Lanz TA, Reinhart V, Sheehan MJ, Rizzo SJS, Bove SE, James LC, et al. Postmortem transcriptional profiling reveals widespread increase in inflammation in schizophrenia: a comparison of prefrontal cortex, striatum, and hippocampus among matched tetrads of controls with subjects diagnosed with schizophrenia, bipolar or major depressive disorder. Transl Psychiatry. 2019;9:151.
Yamada S, Itoh N, Nagai T, Nakai T, Ibi D, Nakajima A, et al. Innate immune activation of astrocytes impairs neurodevelopment via upregulation of follistatin-like 1 and interferon-induced transmembrane protein 3. J Neuroinflammation. 2018;15:295.
Volk DW, Chitrapu A, Edelson JR, Roman KM, Moroco AE, Lewis DA. Molecular mechanisms and timing of cortical immune activation in schizophrenia. Am J Psychiatry. 2015;172:1112–21.
GeneCards: KRAS Gene. https://www.genecards.org/cgi-bin/carddisp.pl?gene=KRAS, 1996-2021. Accessed 27 July 2021.
Chiosea SI, Sherer CK, Jelic T, Dacic S. KRAS mutant allele-specific imbalance in lung adenocarcinoma. Mod Pathol. 2011;24:1571–7.
Hartman DJ, Davison JM, Foxwell TJ, Nikiforova MN, Chiosea SI. Mutant allele-specific imbalance modulates prognostic impact of KRAS mutations in colorectal adenocarcinoma and is associated with worse overall survival. Int J Cancer. 2012;131:1810–7.
Krasinskas AM, Moser AJ, Saka B, Adsay NV, Chiosea SI. KRAS mutant allele-specific imbalance is associated with worse prognosis in pancreatic cancer and progression to undifferentiated carcinoma of the pancreas. Mod Pathol. 2013;26:1346–54.
Schubbert S, Zenker M, Rowe SL, Boll S, Klein C, Bollag G, et al. Germline KRAS mutations cause Noonan syndrome. Nat Genet. 2006;38:331–6.
Du J, Zhang L. Pathway deviation-based biomarker and multi-effect target identification in asbestos-related squamous cell carcinoma of the lung. Int J Mol Med. 2017;39:579–86.
Hayat SMG, Bianconi V, Pirro M, Jaafari MR, Hatamipour M, Sahebkar A. CD47: role in the immune system and application to cancer therapy. Cell Oncol. 2020;43:19–30.
Hellwig M, Lauffer MC, Bockmayr M, Spohn M, Merk DJ, Harrison L, et al. TCF4 (E2-2) harbors tumor suppressive functions in SHH medulloblastoma. Acta Neuropathol. 2019;137:657–73.
Lee GT, Rosenfeld JA, Kim WT, Kwon YS, Palapattu G, Mehra R, et al. TCF4 induces enzalutamide resistance via neuroendocrine differentiation in prostate cancer. PLoS ONE. 2019;14:e0213488.
McKay JD, Hung RJ, Han Y, Zong X, Carreras-Torres R, Christiani DC, et al. Large-scale association analysis identifies new lung cancer susceptibility loci and heterogeneity in genetic susceptibility across histological subtypes. Nat Genet. 2017;49:1126–32.
Kar SP, Beesley J, Amin Al Olama A, Michailidou K, Tyrer J, Kote-Jarai Z, et al. Genome-wide meta-analyses of breast, ovarian, and prostate cancer association studies identify multiple new susceptibility loci shared by at least two cancer types. Cancer Discov. 2016;6:1052–67.
Xu Y, Shen J, Ran Z. Emerging views of mitophagy in immunity and autoimmune diseases. Autophagy. 2020;16:3–17.
Deretic V. Autophagy in infection. Curr Opin Cell Biol. 2010;22:252–62.
Mossakowska-Wojcik J, Orzechowska A, Talarowska M, Szemraj J, Galecki P. The importance of TCF4 gene in the etiology of recurrent depressive disorders. Prog Neuropsychopharmacol Biol Psychiatry. 2018;80:304–8.
Demontis D, Rajagopal VM, Thorgeirsson TE, Als TD, Grove J, Leppala K, et al. Genome-wide association study implicates CHRNA2 in cannabis use disorder. Nat Neurosci. 2019;22:1066–74.
Liu H, Sun Y, Zhang X, Li S, Hu D, Xiao L, et al. Integrated analysis of summary statistics to identify pleiotropic genes and pathways for the comorbidity of schizophrenia and cardiometabolic disease. Front Psychiatry. 2020;11:256.
Saxena R, Elbers CC, Guo Y, Peter I, Gaunt TR, Mega JL, et al. Large-scale gene-centric meta-analysis across 39 studies identifies type 2 diabetes loci. Am J Hum Genet. 2012;90:410–25.
Hasnain M, Fredrickson SK, Vieweg WV, Pandurangi AK. Metabolic syndrome associated with schizophrenia and atypical antipsychotics. Curr Diab Rep. 2010;10:209–16.
Rees E, Walters JT, Georgieva L, Isles AR, Chambert KD, Richards AL, et al. Analysis of copy number variations at 15 schizophrenia-associated loci. Br J Psychiatry. 2014;204:108–14.
Owen MJ, Doherty JL. What can we learn from the high rates of schizophrenia in people with 22q11.2 deletion syndrome? World Psychiatry. 2016;15:23–25.
Rivas MA, Pirinen M, Conrad DF, Lek M, Tsang EK, Karczewski KJ, et al. Human genomics. Effect of predicted protein-truncating genetic variants on the human transcriptome. Science. 2015;348:666–9.
NCBI Gene: SPHK2 sphingosine kinase 2. https://www.ncbi.nlm.nih.gov/gene?cmd=retrieve&list_uids=56848, 2021. Accessed 27 July 2021.
Harris LW, Wayland M, Lan M, Ryan M, Giger T, Lockstone H, et al. The cerebral microvasculature in schizophrenia: a laser capture microdissection study. PLoS ONE. 2008;3:e3964.
Wendt FR, Pathak GA, Levey DF, Nunez YZ, Overstreet C, Tyrrell C, et al. Sex-stratified gene-by-environment genome-wide interaction study of trauma, posttraumatic-stress, and suicidality. Neurobiol Stress. 2021;14:100309.
Barsh GS, Copenhaver GP, Gibson G, Williams SM. Guidelines for genome-wide association studies. PLoS Genet. 2012;8:e1002812.
Hwang Y, Kim J, Shin JY, Kim JI, Seo JS, Webster MJ, et al. Gene expression profiling by mRNA sequencing reveals increased expression of immune/inflammation-related genes in the hippocampus of individuals with schizophrenia. Transl Psychiatry. 2013;3:e321.
Peterson RE, Bigdeli TB, Ripke S, Bacanu SA, Gejman PV, Levinson DF, et al. Genome-wide analyses of smoking behaviors in schizophrenia: findings from the Psychiatric Genomics Consortium. J Psychiatr Res. 2021;137:215–24.
Charlesworth JC, Curran JE, Johnson MP, Goring HH, Dyer TD, Diego VP, et al. Transcriptomic epidemiology of smoking: the effect of smoking on gene expression in lymphocytes. BMC Med Genomics. 2010;3:29.
Huan T, Joehanes R, Schurmann C, Schramm K, Pilling LC, Peters MJ, et al. A whole-blood transcriptome meta-analysis identifies gene expression signatures of cigarette smoking. Hum Mol Genet. 2016;25:4611–23.
Ota VK, Moretti PN, Santoro ML, Talarico F, Spindola LM, Xavier G, et al. Gene expression over the course of schizophrenia: from clinical high-risk for psychosis to chronic stages. NPJ Schizophr. 2019;5:5.
Hauberg ME, Fullard JF, Zhu L, Cohain AT, Giambartolomei C, Misir R, et al. Differential activity of transcribed enhancers in the prefrontal cortex of 537 cases with schizophrenia and controls. Mol Psychiatry. 2019;24:1685–95.
Hoffman GE, Bendl J, Voloudakis G, Montgomery KS, Sloofman L, Wang YC, et al. CommonMind consortium provides transcriptomic and epigenomic data for schizophrenia and bipolar disorder. Sci Data. 2019;6:180.
Mudge J, Miller NA, Khrebtukova I, Lindquist IE, May GD, Huntley JJ, et al. Genomic convergence analysis of schizophrenia: mRNA sequencing reveals altered synaptic vesicular transport in post-mortem cerebellum. PLoS ONE. 2008;3:e3625.
Altar CA, Jurata LW, Charles V, Lemire A, Liu P, Bukhman Y, et al. Deficient hippocampal neuron expression of proteasome, ubiquitin, and mitochondrial genes in multiple schizophrenia cohorts. Biol Psychiatry. 2005;58:85–96.
Arora S, Pattwell SS, Holland EC, Bolouri H. Variability in estimated gene expression among commonly used RNA-seq pipelines. Sci Rep. 2020;10:2734.
Tarca AL, Bhatti G, Romero R. A comparison of gene set analysis methods in terms of sensitivity, prioritization and specificity. PLoS ONE. 2013;8:e79217.
Kuzman MR, Medved V, Terzic J, Krainc D. Genome-wide expression analysis of peripheral blood identifies candidate biomarkers for schizophrenia. J Psychiatr Res. 2009;43:1073–7.
Lee J, Goh LK, Chen G, Verma S, Tan CH, Lee TS. Analysis of blood-based gene expression signature in first-episode psychosis. Psychiatry Res. 2012;200:52–54.
Leirer DJ, Iyegbe CO, Di Forti M, Patel H, Carra E, Fraietta S, et al. Differential gene expression analysis in blood of first episode psychosis patients. Schizophr Res. 2019;209:88–97.
Liu Y, Chang X, Hahn CG, Gur RE, Sleiman PAM, Hakonarson H. Non-coding RNA dysregulation in the amygdala region of schizophrenia patients contributes to the pathogenesis of the disease. Transl Psychiatry. 2018;8:44.
Wu JQ, Wang X, Beveridge NJ, Tooney PA, Scott RJ, Carr VJ, et al. Transcriptome sequencing revealed significant alteration of cortical promoter usage and splicing in schizophrenia. PLoS ONE. 2012;7:e36351.
Cattane N, Minelli A, Milanesi E, Maj C, Bignotti S, Bortolomasi M, et al. Altered gene expression in schizophrenia: findings from transcriptional signatures in fibroblasts and blood. PLoS ONE. 2015;10:e0116686.
Lin M, Pedrosa E, Hrabovsky A, Chen J, Puliafito BR, Gilbert SR, et al. Integrative transcriptome network analysis of iPSC-derived neurons from schizophrenia and schizoaffective disorder patients with 22q11.2 deletion. BMC Syst Biol. 2016;10:105.
Sanders AR, Drigalenko EI, Duan J, Moy W, Freda J, Goring HHH, et al. Transcriptome sequencing study implicates immune-related genes differentially expressed in schizophrenia: new data and a meta-analysis. Transl Psychiatry. 2017;7:e1093.
Sanders AR, Goring HH, Duan J, Drigalenko EI, Moy W, Freda J, et al. Transcriptome study of differential expression in schizophrenia. Hum Mol Genet. 2013;22:5001–14.
Acknowledgements
Maylene (Kefeng) Qiu, M.A., Systematic Review Coordinator and Clinical Liaison Librarian, Biomedical Library, University of Pennsylvania. Hannah L. Bocchinfuso, Resource Coordinator for the Almasy Lab. Support for this project was provided by the Penn Undergraduate Research Mentoring Program, and the CHOP Research Institute Summer Scholars Program, and NIH U01 MH119690.
Author information
Authors and Affiliations
Contributions
LA and AKM designed the study. All authors were involved in reviewing the titles, abstracts, and full-text articles included in the review. AKM, MS, AK, NK, and NT extracted the review data. AKM performed the data synthesis, and AKM and MS performed the data analysis. AKM and LA drafted the manuscript. All authors discussed the results, commented on them, and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41380_2021_1420_MOESM1_ESM.docx
What Genes are Differentially Expressed in Individuals with Schizophrenia? A systematic review. Supplementary information.
41380_2021_1420_MOESM2_ESM.docx
Supplementary table 1: 6771 unique genes reported as statistically significantly differentially expressed in schizophrenia cases
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Merikangas, A.K., Shelly, M., Knighton, A. et al. What genes are differentially expressed in individuals with schizophrenia? A systematic review. Mol Psychiatry 27, 1373–1383 (2022). https://doi.org/10.1038/s41380-021-01420-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-021-01420-7
This article is cited by
-
Three patterns link brain organization to genes in health and disease
Nature Neuroscience (2024)
-
Disclosing common biological signatures and predicting new therapeutic targets in schizophrenia and obsessive–compulsive disorder by integrated bioinformatics analysis
BMC Psychiatry (2023)
-
Patients with schizophrenia and bipolar disorder display a similar global gene expression signature in whole blood that reflects elevated proportion of immature neutrophil cells with association to lipid changes
Translational Psychiatry (2023)
-
Molecular mapping of a core transcriptional signature of microglia-specific genes in schizophrenia
Translational Psychiatry (2023)
-
Reply to “Comment on: What genes are differentially expressed in individuals with schizophrenia? A systematic review”
Molecular Psychiatry (2023)