Abstract
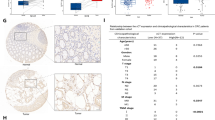
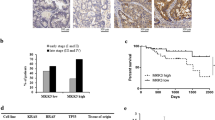
BEST4 is a member of the bestrophin protein family that plays a critical role in human intestinal epithelial cells. However, its role and mechanism in colorectal cancer (CRC) remain largely elusive. Here, we investigated the role and clinical significance of BEST4 in CRC. Our results demonstrate that BEST4 expression is upregulated in clinical CRC samples and its high-level expression correlates with advanced TNM (tumor, lymph nodes, distant metastasis) stage, LNM (lymph node metastasis), and poor survival. Functional studies revealed that ectopic expression of BEST4 promoted CRC cell proliferation and metastasis, whereas the depletion of BEST4 had the opposite effect both in vitro and in vivo. Mechanistically, BEST4 binds to the p85α regulatory subunit of phosphatidylinositol-3-kinase (PI3K) and promotes p110 kinase activity; this leads to activation of Akt signaling and expression of MYC and CCND1, which are critical regulators of cell proliferation and metastasis. In clinical samples, the expression of BEST4 is positively associated with the expression of phosphorylated Akt, MYC and CCND1. Pharmacological inhibition of Akt activity markedly repressed BEST4-mediated Akt signaling and proliferation and metastasis of CRC cells. Importantly, the interaction between BEST4 and p85α was also enhanced by epidermal growth factor (EGF) in CRC cells. Therapeutically, BEST4 suppression effectively sensitized CRC cells to gefitinib treatment in vivo. Taken together, our findings indicate the oncogenic potential of BEST4 in colorectal carcinogenesis and metastasis by modulating BEST4/PI3K/Akt signaling, highlighting a potential strategy for CRC therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Engelman JA, Luo J, Cantley LC. The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat Rev Genet. 2006;7:606–19.
Martini M, De Santis MC, Braccini L, Gulluni F, Hirsch E. PI3K/AKT signaling pathway and cancer: an updated review. Ann Med. 2014;46:372–83.
Noorolyai S, Shajari N, Baghbani E, Sadreddini S, Baradaran B. The relation between PI3K/AKT signalling pathway and cancer. Gene. 2019;698:120–8.
Stemke-Hale K, Gonzalez-Angulo AM, Lluch A, Neve RM, Kuo WL, Davies M, et al. Integrative genomic and proteomic analysis of PIK3CA, PTEN, and AKT mutations in breast cancer. Cancer Res. 2008;68:6084–91.
Samuels Y, Wang ZH, Bardelli A, Silliman N, Ptak J, Szabo S, et al. High frequency of mutations of the PIK3CA gene in human cancers. Science. 2004;304:554–554.
Levine DA, Bogomolniy F, Yee CJ, Lash A, Barakat RR, Borgen PI, et al. Frequent mutation of the PIK3CA gene in ovarian and breast cancers. Clin Cancer Res. 2005;11:2875–8.
Lee JW, Soung YH, Kim SY, Lee HW, Park WS, Nam SW, et al. PIK3CA gene is frequently mutated in breast carcinomas and hepatocellular carcinomas. Oncogene. 2005;24:1477–80.
Triscott J, Rubin MA. Prostate power play: does Pik3ca accelerate Pten-deficient cancer progression? Cancer Discov. 2018;8:682–5.
Cheung LWT, Hennessy BT, Li J, Yu SX, Myers AP, Djordjevic B, et al. High frequency of PIK3R1 and PIK3R2 mutations in endometrial cancer elucidates a novel mechanism for regulation of PTEN protein stability. Cancer Discov. 2011;1:170–85.
Manning BD, Cantley LC. AKT/PKB signaling: navigating downstream. Cell. 2007;129:1261–74.
Stemke-Hale K, Gonzalez-Angulo AM, Lluch A, Neve RM, Kuo WL, Davies M, et al. An integrative genomic and proteomic analysis of PIK3CA, PTEN, and AKT mutations in breast cancer. Cancer Res. 2008;68:6084–91.
Hennessy BT, Smith DL, Ram PT, Lu Y, Mills GB. Exploiting the PI3K/AKT pathway for cancer drug discovery. Nat Rev Drug Discov. 2005;4:988–1004.
Stambolic V, Suzuki A, de la Pompa JL, Brothers GM, Mirtsos C, Sasaki T, et al. Negative regulation of PKB/Akt-dependent cell survival by the tumor suppressor PTEN. Cell. 1998;95:29–39.
Papa A, Wan LX, Bonora M, Salmena L, Song MS, Hobbs RM, et al. Cancer-associated PTEN mutants act in a dominant-negative manner to suppress PTEN protein function. Cell. 2014;157:595–610.
Zhou H, Liu W, Su Y, Wei Z, Liu J, Kolluri SK, et al. NSAID sulindac and its analog bind RXRalpha and inhibit RXRalpha-dependent AKT signaling. Cancer Cell. 2010;17:560–73.
Yan TD, Wu H, Zhang HP, Lu N, Ye P, Yu FH, et al. Oncogenic potential of retinoic acid receptor-gamma in hepatocellular carcinoma. Cancer Res. 2010;70:2285–95.
He XS, Guo LC, Du MZ, Huang S, Huang RP, Zhan SH, et al. The long non-coding RNA NONHSAT062994 inhibits colorectal cancer by inactivating Akt signaling. Oncotarget. 2017;8:68696–706.
Tsunenari T, Sun H, Williams J, Cahill H, Smallwood P, Yau KW, et al. Structure-function analysis of the bestrophin family of anion channels. J Biol Chem. 2003;278:41114–25.
Petrukhin K, Koisti MJ, Bakall B, Li W, Xie GC, Marknell T, et al. Identification of the gene responsible for Best macular dystrophy. Nat Genet. 1998;19:241–7.
Tsunenari T, Nathans J, Yau KW. Ca2+-activated Cl- current from human bestrophin-4 in excised membrane patches. J Gen Physiol. 2006;127:749–54.
Sun H, Tsunenari T, Yau KW, Nathans J. The vitelliform macular dystrophy protein defines a new family of chloride channels. Proc Natl Acad Sci USA. 2002;99:4008–13.
Stohr H, Marquardt A, Nanda I, Schmid M, Weber BHF. Three novel human VMD2-like genes are members of the evolutionary highly conserved RFP-TM family. Eur J Hum Genet. 2002;10:281–4.
Ji C, Li Y, Kittredge A, Hopiavuori A, Ward N, Yao P, et al. Investigation and restoration of BEST1 activity in patient-derived RPEs with dominant mutations. Sci Rep. 2019;9:19026.
Yu K, Lujan R, Marmorstein A, Gabriel S, Hartzell HC. Bestrophin-2 mediates bicarbonate transport by goblet cells in mouse colon. J Clin Investig. 2010;120:1722–35.
Kramer F, Stohr H, Weber BHF. Cloning and characterization of the murine Vmd2 RFP-TM gene family. Cytogenet Genome Res. 2004;105:107–14.
Wu L, Sun Y, Ma L, Zhu J, Zhang B, Pan Q, et al. A C-terminally truncated mouse Best3 splice variant targets and alters the ion balance in lysosome-endosome hybrids and the endoplasmic reticulum. Sci Rep. 2016;6:27332.
Ito G, Okamoto R, Murano T, Shimizu H, Fujii S, Nakata T, et al. Lineage-specific expression of bestrophin-2 and bestrophin-4 in human intestinal epithelial cells. PLoS ONE. 2013;8:e79693.
Yu J, Zhang Y, McIlroy J, Rordorf-Nikolic T, Orr GA, Backer JM. Regulation of the p85/p110 phosphatidylinositol 3’-kinase: stabilization and inhibition of the p110alpha catalytic subunit by the p85 regulatory subunit. Mol Cell Biol. 1998;18:1379–87.
Thorpe LM, Spangle JM, Ohlson CE, Cheng HL, Roberts TM, Cantley LC, et al. PI3K-p110 alpha mediates the oncogenic activity induced by loss of the novel tumor suppressor PI3K-p85 alpha. Proc Natl Acad Sci USA. 2017;114:7095–7100.
Liu Y, Liu Z, Wang K. The Ca(2+)-activated chloride channel ANO1/TMEM16A: an emerging therapeutic target for epithelium-originated diseases? Acta Pharm Sin B. 2021;11:1412–33.
Scaltriti M, Baselga J. The epidermal growth factor receptor pathway: a model for targeted therapy. Clin Cancer Res. 2006;12:5268–72.
Jacobsen K, Bertran-Alamillo J, Molina MA, Teixido C, Karachaliou N, Pedersen MH, et al. Convergent Akt activation drives acquired EGFR inhibitor resistance in lung cancer. Nat Commun. 2017;8:410.
Yang J, Nie J, Ma XL, Wei YQ, Peng Y, Wei XW. Targeting PI3K in cancer: mechanisms and advances in clinical trials. Mol Cancer. 2019;18:1–28.
Manning BD, Cantley LC. AKT/PKB signaling: navigating downstream. Cell. 2007;129:1261–74.
Dieterle AM, Bohler P, Keppeler H, Alers S, Berleth N, Driessen S, et al. PDK1 controls upstream PI3K expression and PIP3 generation. Oncogene. 2014;33:3043–53.
Kaneko T, Li L, Li SSC. The SH3 domain - a family of versatile peptide- and protein-recognition module. Front Biosci. 2008;13:4938–52.
Yang JL, Qu XJ, Russell PJ, Goldstein D. Regulation of epidermal growth factor receptor in human colon cancer cell lines by interferon alpha. Gut. 2004;53:123–9.
Rajagopal S, Huang S, Moskal TL, Lee BN, el-Naggar AK, Chakrabarty S. Epidermal growth factor expression in human colon and colon carcinomas: anti-sense epidermal growth factor receptor RNA down-regulates the proliferation of human colon cancer cells. Int J Cancer. 1995;62:661–7.
Guo PD, Lu XX, Gan WJ, Li XM, He XS, Zhang S, et al. RAR gamma downregulation contributes to colorectal tumorigenesis and metastasis by derepressing the Hippo-Yap pathway. Cancer Res. 2016;76:3813–25.
Xing C, Lu XX, Guo PD, Shen T, Zhang S, He XS, et al. Ubiquitin-specific protease 4-mediated deubiquitination and stabilization of PRL-3 is required for potentiating colorectal oncogenesis. Cancer Res. 2016;76:83–95.
Wu H, Li XM, Wang JR, Gan WJ, Jiang FQ, Liu Y, et al. NUR77 exerts a protective effect against inflammatory bowel disease by negatively regulating the TRAF6/TLR-IL-1R signalling axis. J Pathol. 2016;238:457–69.
Wang JR, Gan WJ, Li XM, Zhao YY, Li Y, Lu XX, et al. Orphan nuclear receptor Nur77 promotes colorectal cancer invasion and metastasis by regulating MMP-9 and E-cadherin. Carcinogenesis. 2014;35:2474–84.
Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D, et al. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 2015;44:e71–e71.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2009;26:139–40.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci. 2005;102:15545–50.
Acknowledgements
This work was supported by the Natural Science Foundation of Jiangsu Province (BK20190042, BK20181434, and BK20190182), National Natural Science Foundation of China (82022050, 81972601, and 81772541), and the Science and Technology Foundation of Suzhou (SYS2019034, SKJY2021070). This work was also supported by the Fujian Provincial Key Laboratory of Innovative Drug Target Research, the Tang Scholar Funds, and the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Contributions
XH and HW contributed to the study conception and design. XH, WY, YY, RZ, and YZ performed the experiments. XY performed the bioinformatics analysis. FL, JW, XD, HB, and WG collected and evaluated the clinical samples. YZ and LG provided technical support. XH, XY, and HW wrote the manuscript. HW supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
He, XS., Ye, WL., Zhang, YJ. et al. Oncogenic potential of BEST4 in colorectal cancer via activation of PI3K/Akt signaling. Oncogene 41, 1166–1177 (2022). https://doi.org/10.1038/s41388-021-02160-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-02160-2
This article is cited by
-
High VSX1 expression promotes the aggressiveness of clear cell renal cell carcinoma by transcriptionally regulating FKBP10
Journal of Translational Medicine (2022)