Abstract
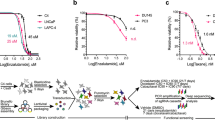
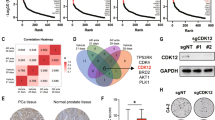
DNA repair gene mutations are frequent in castration-resistant prostate cancer (CRPC), suggesting eligibility for poly(ADP-ribose) polymerase inhibitor (PARPi) treatment. However, therapy resistance is a major clinical challenge and genes contributing to PARPi resistance are poorly understood. Using a genome-wide CRISPR-Cas9 knockout screen, this study aimed at identifying genes involved in PARPi resistance in CRPC. Based on the screen, we identified PARP1, and six novel candidates associated with olaparib resistance upon knockout. For validation, we generated multiple knockout populations/clones per gene in C4 and/or LNCaP CRPC cells, which confirmed that loss of PARP1, ARH3, YWHAE, or UBR5 caused olaparib resistance. PARP1 or ARH3 knockout caused cross-resistance to other PARPis (veliparib and niraparib). Furthermore, PARP1 or ARH3 knockout led to reduced autophagy, while pharmacological induction of autophagy partially reverted their PARPi resistant phenotype. Tumor RNA sequencing of 126 prostate cancer patients identified low ARH3 expression as an independent predictor of recurrence. Our results advance the understanding of PARPi response by identifying four novel genes that contribute to PARPi sensitivity in CRPC and suggest a new model of PARPi resistance through decreased autophagy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63:11–30.
Cornford P, van den Bergh RCN, Briers E, Van den Broeck T, Cumberbatch MG, De Santis M, et al. EAU-EANM-ESTRO-ESUR-SIOG Guidelines on Prostate Cancer. Part II—2020 Update: Treatment of Relapsing and Metastatic Prostate Cancer. Eur Urol. 2021;79:263–82.
Kirby M, Hirst C, Crawford ED. Characterising the castration-resistant prostate cancer population: a systematic review. Int J Clin Pr. 2011;65:1180–92.
Robinson D, Van Allen EM, Wu YM, Schultz N, et al. Integrative clinical genomics of advanced prostate cancer. Cell. 2015;161:1215–28.
Abida W, Armenia J, Gopalan A, Brennan R, Walsh M, Barron D, et al. Prospective Genomic Profiling of Prostate Cancer Across Disease States Reveals Germline and Somatic Alterations That May Affect Clinical Decision Making. JCO Precis Oncol. 2017;1:1–16.
Tukachinsky H, Madison RW, Chung JH, Gjoerup O, Severson EA, Dennis L, et al. Genomic analysis of circulating tumor DNA in 3,334 patients with advanced prostate cancer identifies targetable BRCA alterations and AR resistance mechanisms. Clin Cancer Res. 2021;27:3094–105.
Murai J, Huang SY, Das BB, Renaud A, Zhang Y, Doroshow JH, et al. Trapping of PARP1 and PARP2 by Clinical PARP Inhibitors. Cancer Res. 2012;72:5588–99.
Hopkins TA, Shi Y, Rodriguez LE, Solomon LR, Donawho CK, DiGiammarino EL, et al. Mechanistic Dissection of PARP1 Trapping and the Impact on In Vivo Tolerability and Efficacy of PARP Inhibitors. Mol Cancer Res. 2015;13:1465–77.
FDA. FDA approves olaparib for HRR gene-mutated metastatic castration-resistant prostate cancer FDA website: FDA website; 2021 [Available from: https://www.fda.gov/drugs/resources-information-approved-drugs/fda-approves-olaparib-hrr-gene-mutated-metastatic-castration-resistant-prostate-cancer.
de Bono J, Mateo J, Fizazi K, Saad F, Shore N, Sandhu S, et al. Olaparib for Metastatic Castration-Resistant Prostate Cancer. N Engl J Med. 2020;382:2091–102.
Hussain M, Mateo J, Fizazi K, Saad F, Shore N, Sandhu S, et al. Survival with Olaparib in Metastatic Castration-Resistant Prostate Cancer. N Engl J Med. 2020;383:2345–57.
Quigley D, Alumkal JJ, Wyatt AW, Kothari V, Foye A, Lloyd P, et al. Analysis of Circulating Cell-Free DNA Identifies Multiclonal Heterogeneity of BRCA2 Reversion Mutations Associated with Resistance to PARP Inhibitors. Cancer Disco. 2017;7:999–1005.
Goodall J, Mateo J, Yuan W, Mossop H, Porta N, Miranda S, et al. Circulating Cell-Free DNA to Guide Prostate Cancer Treatment with PARP Inhibition. Cancer Disco. 2017;7:1006–17.
Chen X, Chen Z, Zheng B, Tang W. Targeting NPRL2 to enhance the efficacy of Olaparib in castration-resistant prostate cancer. Biochem Biophys Res Commun. 2019;508:620–5.
Pettitt SJ, Krastev DB, Brandsma I, Dréan A, Song F, Aleksandrov R, et al. Genome-wide and high-density CRISPR-Cas9 screens identify point mutations in PARP1 causing PARP inhibitor resistance. Nature. Communications. 2018;9:1849.
Gogola E, Duarte AA, de Ruiter JR, Wiegant WW, Schmid JA, de Bruijn R, et al. Selective Loss of PARG Restores PARylation and Counteracts PARP Inhibitor-Mediated Synthetic Lethality. Cancer Cell. 2018;33:1078–93.
Doench JG, Fusi N, Sullender M, Hegde M, Vaimberg EW, Donovan KF, et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotechnol. 2016;34:184.
Li W, Xu H, Xiao T, Cong L, Love MI, Zhang F, et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knocout screens. Genome Biol. 2014;15:554–66.
Kolde R, Laur S, Adler P, Vilo J. Robust rank aggregation for gene list integration and meta-analysis. Bioinformatics. 2012;28:573–80.
Gibson BA, Kraus WL. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat Rev Mol Cell Biol. 2012;13:411–24.
Jirawatnotai S, Hu Y, Michowski W, Elias JE, Becks L, Bienvenu F, et al. A function for cyclin D1 in DNA repair uncovered by protein interactome analyses in human cancers. Nature 2011;474:230–4.
Hazan I, Monin J, Bouwman BAM, Crosetto N, Aqeilan RI. Activation of Oncogenic Super-Enhancers Is Coupled with DNA Repair by RAD51. Cell Rep. 2019;29:560–72.e4.
Li CG, Mahon C, Sweeney NM, Verschueren E, Kantamani V, Li D, et al. PPARγ Interaction with UBR5/ATMIN Promotes DNA Repair to Maintain Endothelial Homeostasis. Cell Rep. 2019;26:1333–43.
Pennington KL, Chan TY, Torres MP, Andersen JL. The dynamic and stress-adaptive signaling hub of 14-3-3: emerging mechanisms of regulation and context-dependent protein–protein interactions. Oncogene. 2018;37:5587–604.
Velimezi G, Robinson-Garcia L, Muñoz-Martínez F, Wiegant WW, Ferreira da Silva J, Owusu M, et al. Map of synthetic rescue interactions for the Fanconi anemia DNA repair pathway identifies USP48. Nat Commun. 2018;9:2280.
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308.
Steensel V TIDE: Tracking of Indels by DEcomposition [cited 2020 23/4]. Available from: https://tide.deskgen.com/.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2009;26:139–40.
Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucl Acids Res. 2016;44(W1):W90–7.
Chen EY, Tan CM, Kou Y, Duan Q, Wang Z, Meirelles GV, et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinforma. 2013;14:128.
Rath S, Sharma R, Gupta R, Ast T, Chan C, Durham TJ, et al. MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucl Acids Res. 2020;49(D1):D1541–7.
Lee JS, Lee H, Jang H, Woo SM, Park JB, Lee SH, et al. Targeting Oxidative Phosphorylation Reverses Drug Resistance in Cancer Cells by Blocking Autophagy Recycling. Cells. 2020;9:2013.
Eliopoulos AG, Havaki S, Gorgoulis VG. DNA Damage Response and Autophagy: A Meaningful Partnership. Front Genet. 2016;7:204.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Disco. 2012;2:401–4.
He YJ, Meghani K, Caron MC, Yang C, Ronato DA, Bian J, et al. DYNLL1 binds to MRE11 to limit DNA end resection in BRCA1-deficient cells. Nature. 2018;563:522–6.
Niere M, Kernstock S, Koch-Nolte F, Ziegler M. Functional localization of two poly(ADP-ribose)-degrading enzymes to the mitochondrial matrix. Mol Cell Biol. 2008;28:814–24.
Zhang M, Lai Y, Vasquez JL, James DI, Smith KM, Waddell ID, et al. Androgen Receptor and Poly(ADP-ribose) Glycohydrolase Inhibition Increases Efficiency of Androgen Ablation in Prostate Cancer Cells. Sci Rep. 2020;10:3836.
Cahuzac M, Langlois P, Péant B, Fleury H, Mes-Masson A-M, Saad F. Pre-activation of autophagy impacts response to olaparib in prostate cancer cells. Commun Biol. 2022;5:251.
Tai S, Sun Y, Liu N, Ding B, Hsia E, Bhuta S, et al. Combination of Rad001 (everolimus) and propachlor synergistically induces apoptosis through enhanced autophagy in prostate cancer cells. Mol Cancer Ther. 2012;11:1320–31.
Arun B, Akar U, Gutierrez-Barrera AM, Hortobagyi GN, Ozpolat B. The PARP inhibitor AZD2281 (Olaparib) induces autophagy/mitophagy in BRCA1 and BRCA2 mutant breast cancer cells. Int J Oncol. 2015;47:262–8.
Formentini L, Macchiarulo A, Cipriani G, Camaioni E, Rapizzi E, Pellicciari R, et al. Poly(ADP-ribose) Catabolism Triggers AMP-dependent Mitochondrial Energy Failure*. J Biol Chem. 2009;284:17668–76.
Virág L, Szabó C. The Therapeutic Potential of Poly(ADP-Ribose) Polymerase Inhibitors. Pharmacol Rev. 2002;54:375-429.
Lord CJ, Ashworth A. PARP inhibitors: Synthetic lethality in the clinic. Science. 2017;355:1152–8.
Valabrega G, Scotto G, Tuninetti V, Pani A, Scaglione F. Differences in PARP Inhibitors for the Treatment of Ovarian Cancer: Mechanisms of Action, Pharmacology, Safety, and Efficacy. Int J Mol Sci. 2021;22:4203.
Ryø LB, Thomsen EA, Mikkelsen JG. Production and Validation of Lentiviral Vectors for CRISPR/Cas9 Delivery. Methods Mol Biol. 2019;1961:93–109.
Acknowledgements
This work was supported by grants from The Novo Nordisk Foundation (KDS), Aase & Ejnar Danielsens Fond (MBI), Fabrikant Einar Willumsens Mindelegat (MBI), Tømrermester Jørgen Holm & Hustru Elisa F. Hansens Mindelegat (MBI), Direktør Emil C. Hertz og Hustru Inge Hertz Fond (MBI), Graduate School of Health (Aarhus University; MBI), Helge Peetz & Verner Peetz & Hustru Vilma Peetz Fond (MBI), Beckett Fonden (MBI), and NEYE Fonden (MBI). The Danish Cancer Biobank is acknowledged for providing patient samples and information on handling and storage.
Author information
Authors and Affiliations
Contributions
KDS, MBI, EMGS, JGM, and JH designed the study. MBI and EAT conducted the CRISPR-Cas9 screen. MBI and EMGS produced single gene knockout cell lines and performed all cell line experiments. MBI performed RNA-sequencing analyses for cell lines. AD performed western blots of LC3II/I. JP performed LC-MS experiments. PB supervised Seahorse assays. KDS, JF, MR, SK, MRJ, KB, BPU, and MB were responsible for collection and sequencing of patient samples. MBI and EMGS analyzed patient RNA-sequencing data. MBI, EMGS, SW, PB, JP, and KDS interpreted data. MBI, EMGS and KDS wrote the manuscript. KDS, SW, and JGM supervised the study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ipsen, M.B., Sørensen, E.M.G., Thomsen, E.A. et al. A genome-wide CRISPR-Cas9 knockout screen identifies novel PARP inhibitor resistance genes in prostate cancer. Oncogene 41, 4271–4281 (2022). https://doi.org/10.1038/s41388-022-02427-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02427-2
This article is cited by
-
Fit-Seq2.0: An Improved Software for High-Throughput Fitness Measurements Using Pooled Competition Assays
Journal of Molecular Evolution (2023)