Abstract
Colorectal cancer (CRC) is one of the most frequently occurring malignancy tumors with a high morbidity additionally, CRC patients may develop liver metastasis, which is the major cause of death. Despite significant advances in diagnostic and therapeutic techniques, the survival rate of colorectal liver metastasis (CRLM) patients remains very low. CRLM, as a complex cascade reaction process involving multiple factors and procedures, has complex and diverse molecular mechanisms. In this review, we summarize the mechanisms/pathophysiology, diagnosis, treatment of CRLM. We also focus on an overview of the recent advances in understanding the molecular basis of CRLM with a special emphasis on tumor microenvironment and promise of newer targeted therapies for CRLM, further improving the prognosis of CRLM patients.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is one of the three most common cancers and the fourth most common cause of cancer deaths worldwide.1 In 2020, 1.9 million cases were newly diagnosed with CRC and 935,000 cases with CRC past away.2 The global incidence of CRC has been rising with annual increases of 3.2%, beginning with 783,000 cases in 1999 and increasing to 1.8 million in 20202,3,4,5,6,7; this trend is likely to continue. Accordingly, the global CRC burden is increasing mainly owing to the growth of human development index—The incidence rates in developing countries is a quarter of that in developed countries.2
Metastasis of CRC remains a major problem after curative treatment and is the critical cause of CRC-related death.8 The liver is the most common organ of distant metastasis in CRC.9 Liver metastasis of CRC may be associated with the following factors: the portal vein system directly connecting the colorectal and liver, which is associated with abundant blood supply; location and histological type of primary tumor.10,11,12 Curative resection and chemotherapy are the standard methods treatment in patients with colorectal liver metastasis (CRLM),13 However, due to factors such as the location and size of the tumor, unresectable disease, presence of extrahepatic disease, or patients’ comorbidities, surgery is only applicable in 10–20% of cases, with a 5-year survival rate as low as 30%.14,15 Furthermore, those who are not eligible for surgery have an even worse prognosis. Although, significant progress in the development of new chemotherapeutic drugs has been made, CRLM patients that receive fluorouracil and platinum chemotherapy initially, will eventually develop chemotherapy resistance due to inherent or acquired resistance. For CRLM, it is necessary to find more effective targeted therapies. Understanding of the molecular mechanisms underlined in this process could accelerate the achievement of this goal.
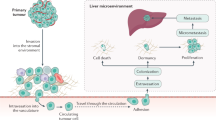
A small subset of CRC cells acquires a capacity to evade from the primary CRC, in part by morphological changes such as epithelial-to-mesenchymal transition (EMT), migration through the extracellular matrix (ECM), and invasion into the neighboring tissues, intravasation, survival in the circulation, extravasation and finally colonization to distant liver forming more aggressive secondary CRLM.
Epidemiology
Nearly 50% of patients with CRC will develop liver metastases during the disease.16 The incidence of stage IV left colon cancer liver metastases is high.17,18 However, once liver metastasis occurred in right colon cancer, the range of liver metastasis was wider than that in left colon cancer.19 The probability of liver metastasis in CRC is also associated with gender. Males are more likely to be a risk factor of CRC, with higher illness burden and earlier onset age2,4,20 and 25–50% of these patients develop liver metastasis21,22, of which ~30% are diagnosed with CRC at the same time. Notably, ethnicity was likely to be correlated with the incidence of development of CRLM. Research in the United States alone reported that non-Hispanic black patients have the highest incidence of liver metastasis, followed by American Indian/Alaska Native, non-Hispanic white, Hispanic and Asian/Pacific Islanders.23 That is, the variations in genes might contribute to the CRLM. Prior works have suggested that BRAF, KRAS, NRAS, PI3KCA, TP53, NRAS, CDK12, EBF1 might be risk genes of CRLM.24,25,26,27 Furthermore, the types of gene mutations were also likely to be correlated with the prognosis27—the highest cure rates appeared in patients with NOTCH1 and PIK3C2B mutations, and the lowest in those with SMAD3 mutations.
Mechanisms/pathophysiology
In 1889, Paget proposed the concepts of “seed” (tumor cells) and “soil” (specific organs) for tumor metastasis.28 The liver metastatic cascade of CRC is a complex multi-factor and multi-step biological process (Fig. 1), wherein a small subset of CRC cells acquire a capacity to evade from the primary CRC, in part by morphological changes such as EMT, migration through the ECM, invasion into the neighboring tissues, intravasation, survival in the circulation, extravasation and finally, colonization to distant liver forming more aggressive secondary CRLM.8,29 On one hand, this process is related to genome abnormalities in the tumor cells themselves, including activation of proto-oncogenes and inactivation of tumor suppressor genes. The occurrence and development of CRLM is especially complex and involves many molecular mechanisms, including non-coding RNAs (ncRNAs), Notch pathway, TGFβ signaling, Tyrosine kinase c-MET signaling, phosphatase of regenerating liver (PRL3), tumor-associated calcium signal transducer 2(Trop-2), L1 cell adhesion molecule (L1CAM), S100 family proteins S100A4 and S100A8 and other pathways (Table 1). It is also closely related to the tumor microenvironment (TME), which mainly involves various immune cells (macrophages, T cells, B cell etc.), cytokines, chemokines and exsomes.30,31,32 The interaction of internal and external environment jointly initiates and drives the occurrence of CRLM.
Immune cells
Immune microenvironment plays a pivotal role in CRLM.33 The liver is an important immune organ of the human body. If its immune killing ability is weakened, it can promote the survival and growth of CRC metastases. Therefore, a suppressive immunologic microenvironment plays tumor-promoting role of CRLM, which is related to tumor-associated-macrophages (TAMs) and regulatory T cells (Treg). However, TAMs maintains the immunosuppressive environment by expressing checkpoint ligand programmed death ligand 1 (PDL1), PDL2 and other inhibitory receptors34, and activates Treg cells by secreting IL-10 and TGFβ.35 TAMs also release a plethora of ECM remodeling factors (plasminogen activation system, matrix metalloproteinases (MMPs), and kallikrein-related peptidases) and a diverse array of proteolytic enzymes such as MMPs that degrades the ECM proteins. These factors, in turn, enhance migration of tumor cells.36 In addition, when targeting the CCL2/CCR2 chemokine axis, TAMs infiltration at the metastatic site is reduced and metastatic CRC (mCRC) is sensitized to tumor T cells.37
Meanwhile, Treg cells inhibit aberrant immune response against self-antigens and antitumor immune response.38 The ability to inhibit adaptive antitumor immune responses via Treg is associated with clinical outcomes of CRLM.39 The intratumoral Treg could inhibit MMPs expression and activity with IL-17 producing T cells involved which reduces the probability of postoperative metastasis of CRC.40 Inhibition of Treg activity may be a therapeutic approach to improve antitumor immunity in the future.39
The tumor-associated neutrophils (TANs) may also promote tumor growth and metastasis through a variety of mechanisms.41 Hyperlipidemia can promote neutrophil infiltration, thereby increasing the metastasis of CRC.42 In the early phase of CRC cell dissemination, neutrophils expressed CCR1 exclusively and MMP9 preferentially, which contribute to the early expansion of cancer.43 TANs produce large amounts of lysine oxidase-like 4 protein during CRLM resistance anti-angiogenic therapy.44 TANs could be recruited by loss of SMAD4 promoting chemokine CCL15 expression via the CCL15-CCR1 axis.45 In addition, neutrophil extracellular traps (NET) trigger the release of HMGB1 and promote the migration and invasion of cancer cells during stress response.41 NETs could also directly trap CRC cells in liver boosted tumorous proliferation and invasion capacity, which is because of the heightened expression of tumorous interleukin (IL)-8 which is triggered by NETs.46 Interestingly, overproduced IL-8 in turn activated neutrophils towards NETs formation, thus forming a positive loop enhanced CRC liver metastasis. Therefore, eliminating NET may reduce risks of tumor relapse after surgical stress. Recently, Xia et al.47 reported neutrophil infiltration and NETs formation were inhibited in tumor tissues with Adeno‐associated virus‐mediated DNase I liver gene transfer treatment. NET-associated carcinoembryonic Ag cell adhesion molecule 1 (CEACAM1) as an essential element for inducing CRC metastatic phenotype, which prompted CEACAM1 as a potential therapeutic target for the prevention of CRC metastasis.
Myeloid-derived suppressor cells (MDSC) are another key factor that regulates the immune response under many pathological conditions, and immune suppressive activity is an intrinsic feature of MDSC. Neutrophils or monocytes that has undergone maturation is not able to transform into potent immune suppressive cells in vitro when simply activating MDSC with hazard associated molecular patterns (DAMPs) and pathogen-associated molecular patterns or pro-inflammatory cytokines.48 CCR1(+) MDSC are recruited to the microenvironment of disseminated CRC cells by loss of SMAD4 promoting chemokine CCL15 expression via the CCL15-CCR1 axis, and produce metalloproteinases MMP9 and MMP2 to enhance metastatic colonization.43,49 S1PR1–activator of transcription 3 (STAT3) upregulation in CRC cells promotes IL-6, which could activate S1PR1–STAT3 in MDSC in the liver, leading to premetastatic niche formation before CRC cell arrival.50
There has been some progress regarding the immune status of CRLM and the effect of tumor immunological characteristics on clinical outcome in CRLM patients. The density of macrophages in CRC is also closely related to patient prognosis.51 The CRLM T cell number is an independent correlate of long-term survival following liver resection, in particular, CD8 + T‐cell infiltration in liver metastasis is associated with better prognosis.52,53,54 Moreover, high frequencies of NK and T cells in response to chemotherapy predict OS in CRLM patients.55
Cytokines
Cytokines IL-6 active IL-6 receptor (IL-6R), which induce STAT3 to bind to MIR34A inhibit miR-34A expression, thereby promoting EMT-mediated invasion and metastasis of CRC.56 Growth differentiation factor 15 in inflammatory microenvironment induces CRC metastasis by regulating EMT genes by activating c-Fos, which may be a new direction for CRLM treatment in the future.57 Similarly, cytokines IL-33 modulates the TME to potently induce the occurrence of liver metastasis.58 In addition, the levels of arachidonic acid (AA) and eicosapentaenoic acid (EPA) promoted the development of an inflammatory microenvironment and AA/EPA ratio was elevated levels in patients with metastatic CRC.59
Chemokines
Chemokines, as an important component of TME, play an important role in the entry of tumor cells into the TME.60 For instance, chemokines are critical for the tumor-stroma interaction, which promote tumor metastasis via chemokines signaling. Chemokines could also act as a bridge between the microenvironment outside the tumor and the tumor itself, and their cognate receptors are expressed by both tumor and stromal cells.61 In recent years, it has been reported that chemokine ligands and receptors (such as CXCL5, CCL3, CCL4, CXCL2, CXCL3, CXCL8, CCL3L3, CCL4L2, CCL18) have been significantly dysregulated, which implies a potential intercellular communication in the immune microenvironment of CRC. Chemokine ligand–receptor interactions between different cells further complicate the signal transduction in CRLM (Table 1).
As a member of the CXC chemokine family, CXCL5 is the ligand of CXCR2 and is not only derived from primary tumor cells but is also secreted by immune cells in the TME.62
The CXCL5/CXCR2 biological axis promoted tumor angiogenesis by activating the AKT/NF-κB/FOXD1/VEGF-A pathway.63 Moreover, Zhao et al.64 found the elevated expression of CXCL5 in CRC induced cell migration by the ERK/Elk-1/Snail signaling pathway and promoted cell invasion through the AKT/GSK3β/β-catenin/MMP7 signaling pathway. They also found that high expression of CXCL5 was a favorable factor to promote the metastasis of CRC cells to the liver in nude mice intrasplenic injection model.
The earlier study confirmed that CCR1+ immature myeloid cells (iMCs) could migrate toward CRC-derived CCL9 and accumulate at tumor invasion front in mice model.65 Later studies demonstrated CRC cells secrete CC-chemokine ligands CCL9 and CCL15, respectively, and recruit CD34(+) Gr-1(−) iMCs to the metastatic liver.66 Additionally, CRC-derived CCL2 and Myeloid-derived S100A8/A9 enhanced CD11b(+)Gr-1(+) iMCs recruitment and increased tumor burden via CCL2/CCR2.67,68 Also, CRC-derived CXCL1 increased migration of CXCR2+ microvascular endothelial cells in vitro and mediated tumor angiogenesis by binding with CXCR2 receptor on endothelial cells in xenograft CRC models.69,70 Moreover, CXCL1 is found to recruit CXCR2+ neutrophil and macrophages from the bone marrow to premetastatic liver in CRC nude mice.71 It was also reported that CCL15 secreted by CRC attracted CCR1(+) MDSCs to the premetastatic niche via the CCL15-CCR1 axis, creating conditions conducive to metastasis.49 TAMs also recruited CXCR2-positive MDSCs to promote liver metastases via producing CXCL1.72 High expression of CCL4 in CRC could induce the infiltration of TAMs and specifically a pro-tumor macrophage profile (CD163 + cells).73 CCL2/CCR2 chemokine axis could also facilitate TAMs accumulation in liver metastasis and maintain immunosuppressive TME. In addition, CXCL8 and its receptor (CXCL8-R) promote liver metastasis by promoting angiogenesis and inducing EMT in CRC cells.74 Elevated serum CXCL8 levels are associated with poor prognosis in patients with CRC.45,75
In clinical CRC specimens, CX3CR1, the chemoattractant cytokine CX3CL1 receptor, was expressed on TAMs and in a microenvironment lacking CX3CR1, the liver metastasis of CRC cells was significantly inhibited.76 However, the high expression of CX3CR1 promoted the increase of T cell invasion in tumor tissue, and thus inhibited tumor growth.77 Low-molecular-weight heparin, a common drug for venous thromboembolism, inhibited the formation of liver metastasis of CRC by disrupting the interaction of CXCR4 and CXCL12.78 Stromal cell-derived factor 1alpha activated CXCR4-expressing CRC cells, then resulted in a significant increase of cell migration.79 Moreover, hepatic stellate cells (HSCs) play important role in liver metastasis of CRC cells by the action of SDF-1/CXCR4 axis.79
The emerging roles of exosomes in CRLM
Like chemokine, exosomes also built crosstalk between various cells. Exosomes are lipid bilayer vesicles with a diameter of 30–100 nm.80 Exosomes are very widespread and are found in almost all body fluids.81 Cancer patients have more circulating exosomes than non-cancer patients.82 Exosomes can also mediate colonic epithelial-stromal interactions involved in the regulation of tumor growth and metastatic invasion.83 Exosomes play a key role in building a supportive microenvironment in the metastatic organs, namely the premetastatic niches (PMNs), which include vascular leakiness, inflammation, and immunosuppression.84 We review the role of exosomes in CRLM development, and the clinical application of exosomes (Figs. 2 and 3).
CRC-derived exosomes contain various biomolecules such as protein (PAD4, FasL, TGFβ, angiopoietin-like protein 1 (ANGPTL1), AKt, tumor necrosis factor-α (TNF-α), β-catenin), RNA (miR-246, miR-21, miR-25-3p, miR-375-3p, miR-92b-3p, miR-27a-3p, miR-17, MALAT1). These biomolecules establish PMNs in liver and induce proliferation, inflammation, EMT, invasion, migration and EMT, thereby promoting CRC metastasis.
The Complicated network of the exosomal interactions among the CRC cells and TAMs promotes metastasis. CRC induces macrophages to M2 macrophages in TME via secreting exosomal miRNA (miR-934, miR-25-3p, miR-130-3p, miR-425-5p and miR-1246). M2 macrophages also promoted CRC EMT and maintained stemness by exosomal miR-21-5p and WNTs. Moreover, CXCR5, IL-10, IL-4 and VEGF were also the medium of communication between CRC cells and TAMs.
The substances in exosomes
The substance in exosomes includes proteins, miRNAs, lncRNAs and mRNAs, which can spread in the circulatory system and play an important role in CRLM. ANGPTL1, a tumor suppressor, was decreased in CRC tissue.85 Exosomal ANGPTL1 decrease MMP9 production in Kupffer cell by suppressing the JAK2-STAT3 signaling pathway.86 Tumor cells can transfer EMT inducers, such as IL-6, Akt and TNF-α, through exosomes to induce EMT in neighboring tumor cells.87
In addition to protein, miRNAs are also a common substance in exosomes. Circulating exosomal miRNA can be used to assist in the diagnosis or prognosis evaluation of CRC patients. Circulating miRNAs are mainly derived from exosomes miRNA.88 Therefore, the analysis of circulating exosomal miRNAs can promote the early metaphase diagnosis rate of CRLM and intervention in the metastatic process. Zeng et al.89 found that exosomal miR-25-3p secreted by CRC cells was transferred to vascular endothelial cells and promoted CRC metastasis by targeting Kruppel-like factor 2 and Krüppel-like factor 4 (KLF4) to increase vascular permeability and promote angiogenesis in animal models. The clinical data further showed that miR-25-3p from circulating CRC-derived exosomal miR-25-3p can be used as a biomarker to predict metastasis. Similarly, CRC-derived exosomes packed with miR-21-5p created a liver pro-inflammatory phenotype and liver metastasis of CRC via the miR-21-Toll like receptor 7-IL-6 axis.90 Metastasis-associated miR-106b-3p from serum exosomes targeted deleted in liver cancer-1.91 Elisabetta et al.92 reported that exosomal miR-210 can perhaps be reckoned as an EMT signal promoter which maintain the local cancer-growth milieu and influence the adhesion and migration of CRC cells. Moreover, Matsumura et al.93 showed that six exosomal miRNAs (miR-19a, miR-19b, miR-4437, miR-23a, miR-320a and miR-92a) were related to the liver metastasis. Interestingly, miR-375 mimicThe tumor-derived exosome containing miR-375 mimic that could inhibit the EMT process.94 According to Fu et al.95 the high expression of miR-17-5p and miR-92a-3p were consistent with the tumorigenesis and metastasis of CRC by exploring serum exosomal miRNAs levels of normal controls and CRC patients. Interestingly, exosomal miR-1246/92b-3p/27a-3p derived by fusobacterium nucleatum-infected CRC cells facilitate uninfected cells metastasis.96 Recently, Zhang et al.97 reported the secretion of CRC exosome miR-1255b-5p was decreased under hypoxia, thereby promoting human telomerase reverse transcriptase inhibition to enhance EMT and telomerase activity.
Moreover, exosomal miRNAs are secreted by cancer-associated fibroblasts (CAFs), then they are transferred to CRC cells.98 Mechanistically, miR-92a-3p inhibits mitochondrial apoptosis by activating the Wnt/β-catenin pathway and inhibiting FBXW7 and MOAP1, thereby enhancing stemness, EMT, metastasis and 5-FU/L-OHP resistance of CRC cells.98 Reducing exosomal miR-92a-3p may contribute to the prediction and treatment of CRLM. CRC cells can also induce CAF via exosomal tumor growth factor-β (TGF-β).99,100 Moreover, exosomes miR-142-3p secreted by bone marrow-derived mesenchymal stem/stromal cells inhibited Numb expression in CRC cells, thereby increasing the population of CSCs.101
MiRNAs in exosomes isolated from CRLM are different from the profile generated from an orthotopic cecum tumor model and naïve colon tissues because higher levels of tumor suppressor miRNAs are encapsulated in the exosomes in more advanced disease.102 The level of oncogenic miR-21 was much higher in the primary colon tumor tissue and metastatic colon tumor in the liver than in their exosomes. However, the level of tumor suppressive miR-18a and miR-193a in the exosomes is enhanced.102 The serum levels of exosomal miR-200c and miR-141 were proved to predict CRC patients with poor prognosis.103 The monitoring of CRC metastasis was monitored in real time based on miR-139-3p content in the plasma of CRC patients.104 Similarly, in CRC patients with LM, the serum exosomal miR-122 can serve potentially as both a novel diagnostic and prognostic biomarker.105 A recent study provides a new concept that miR-25-3p, miR-130b-3p, miR-425-5p, miR-193a, let-7g,miR-106b-3pand miR-934 contained in exosomes could participate in progression and metastasis of CRC.91,106,107,108 Thus, all the evidences point out that exosome-mediated promotion of tumor progression, and understanding these mechanism-based circulating exosomal miRNAs will reveal new avenues for future diagnosis and treatment.
Furthermore, the exosome-derived lncRNAs could play essential roles in tumorigenesis by regulating the TME.109 Exosomal lncRNAs BCAR4 could be potential candidates to detect CRC.110 Exosomal MALAT1 regulated fucosyltransferase 4 (FUT4) expression by sponging miR-26a/26b to promote CRC progress.111 CRC cells secreted exosome cirC-133 into relatively normoxic CRC cells under hypoxic conditions. Then circ-133 adsorbed miR-133a to target GEF-H1/RhoA, as a result of reducing the distribution of E-cadherin on the membrane.112 Guo et al.96 reported fusobacterium infection may stimulate CRC cells to generate CXCL16/RhoA/IL-8 exosomes that are delivered to uninfected cells to promote prometastatic behaviors.
The exosomal interaction between CRC cells and TAMs
There is also a noticeable interaction between CRC cells and TAMs via exosomes.113 Exosomes secreted by M2 macrophages promoted CRC migration by exosomal transfer of miR-21-5p and miR-155-5p.114 Mechanistically, these miRNAs mentioned above targeted the core component of the Switch/sucrose non-fermentable complex BRG1. Moreover, M2 macrophages -derived exosomes could induce CRC stem cell activity because of containing WNT.115 CRC cell-derived exosomal miR-934 induced M2 macrophage polarization via downregulating PTEN expression and activating the PI3K/AKT signaling pathway.107 Polarized M2 macrophages secreted CXCL13, then activated a CXCL13/CXCR5/NFκB/p65/miR-934 positive feedback loop to induce PMN formation in CRC cells. Exosomal miRNAs miR-25-3p, miR-130b-3p, miR-425-5p secreted by CRC cells via activation of the CXCL12/CXCR4 axis, could be transferred to macrophages which can then target PTEN and shift the macrophage towards M2 phenotype.106 Additionally, mutant p53 CRC could reprogram macrophages into M2 macrophages by transferring exosomal miR-1246.116
The exosomal interaction between CRC cells and other types of immune cells
CRC-exosomes containing transferring TGF-β can activate TGF-β/Smad signaling and inactivate SAPK signaling which induces phenotypic alteration of T cells to Treg-like cells.117,118,119 Delivery of Fas-ligand-containing CRC cell exosomes to T cells can induce cell apoptosis.120 Citrullination generated by peptidylarginine deiminase 4 (PAD4) derived from CRC cells as a driver of liver metastases.121 In another study, it was reported that mutated KRAS is transferred to recipient cells through exosomes and induces increased IL-8 production, recruiting more neutrophils and further aggravating CRC.122
Key molecules and signaling pathway driving liver metastasis
Various signaling pathways and factors may be involved in the process of CRLM, including hepatocyte growth factor/c‐Met (HGF/c-Met) signaling pathway, phosphatase of regenerating liver (PRL3), Notch pathway, TGFβ signaling, Tyrosine kinase c-MET signaling, tumor-associated calcium signal transducer 2 (Trop-2), L1 cell adhesion molecule (L1CAM), metastasis-related gene 1 (MACC1), S100 family proteins and other pathways. There are multiple intersections between these molecular mechanisms driving CRLM.
HGF/c-Met signaling pathway
HGF/c-Met signaling pathway promotes metastasis of cancer cells by regulating a diverse downstream prometastatic effector molecules, then overactivated phosphatidylinositol-3-kinase (PI3K) and mitogen-activated protein kinase (MAPK) signaling (Fig. 4).123,124,125 Met, a receptor of hepatocyte growth factor (HGF), has a positive correlation with tumor stages of CRC liver metastasis. High expression of circulating HGF and aberrant activity of cMet were detected in patients with CRC.126 Inhibition of HGF/C-MET signaling pathway could decrease the proliferation and invasion of CRLM.127 By assessing the patient tissue samples, Yao et al.128 reported higher levels of c-Met expression (mRNA and protein) in CRLM than primary CRC by assessing the patient tissue samples. HGF/c-Met could regulate urokinase plasminogen activator (uPA) and uPAR in vitro CRC migration and invasion models.129 Ectopic TIMP1 expression provoked prometastatic microenvironment in the liver by inducing the HGF/c-Met signaling as well as uPA expression.130,131 Inhibition of ADAM-10 was in principle able to prevent shedding of c-Met, which may be one explanation for the increase of cell-associated c-Met in livers with elevated TIMP-1.130 Indeed, in livers of uPA-ablated mice upregulated TIMP-1 expression did not trigger HGF/c-Met signaling.131 Moreover, elevated levels of TIMP-1 in the TME could induce metastasis by enhancing HIF-1α-dependent HGF-signaling.132
In addition, SOX13 is induced by HGF through JAK2/STAT3 signaling and then upregulated SOX13 transactivates the expression of c-MET by directly binding to its promoters, which contributes to c-MET overexpression in CRC.133 This feedback loop induces SOX13-mediated CRC migration, invasion, and metastasis. Similar to SOX13, Forkhead box protein C2 (FOXC2) was directly associated with c-MET promoter to increase the transcriptional activity of MET.134 Blocking KPNB1 (a novel gene) expression showed a significant inhibitory role in metastasis both in vitro and in vivo through interacting with MET proto-oncogene. COX-2/PGE(2) pathway as an important mediator of HGF/Met signaling is closely associated with the survival, proliferation and invasion of CRC cells.135 HGF/Met signaling is an important regulator of the COX-2/PGE 2 pathway in CRC cells, stimulating PGE 2 synthesis via COX-2 upregulation and inhibiting PGE 2 degradation via Ras-MAPK/ERK while PI3K/AKT signaling mediate HGF-driven 15-PGDH downregulation.136 In fact, IL-6/IL-6R, HGF/c-Met, STAT3, VEGF cytokinetic pathway is a major mechanism of the pro-oncogenic effect induced by hepatic RFA.137
The association between C-Met and CRC was even stronger due to c-MET being identified as a transcriptional target of colon cancer MACC1.138 MACC1 promotes proliferation, invasion, and HGF-induced scattering of CRC cells in cell culture and tumor growth and metastasis in mouse models. In advanced metastatic CRC patients, MACC1 and c-Met were both upregulated.138 MACC1, a new detectable biomarker in cancer, is also an independent prognostic factor for the recurrence after liver resection of CRC metastasis.139
HGF/c-Met signaling promoted metastasis of cancer cells by regulating a diverse downstream prometastatic effector molecules, via Ras-MAPK/ERK, PI3K/AKT signaling, JAK2/STAT3 signaling. MACC1, KPNB1 and FOXC2 could transactivate the expression of c-MET.
CRLM and PRL3
PRL3 has received widespread attention as a potential cause for metastasis (Fig. 5).140 Since PRL3 transcripts are overexpressed in CRC metastases found in the liver, and PRL3 expression cannot be detected in non-metastatic primary tumors and normal colorectal epithelium. PRL3-induced enhancement of EMT dependent on EGFR activation and PRL3 promoted cell invasion and upregulated MMPS by activating AKT in vitro and in vivo.141,142,143,144 The levels of PRL-3 mRNA expression can be used as biomarkers for increased risk of liver metastasis.145 The PRL-3 expression did not represent a direct causative mechanism of liver metastasis, but modulated multiple signaling pathways, including PI3K/AKT and MAPK/ERK in various cancer cells. It was reported liver metastasis by PRL-3 is mediated through lymph node metastasis and elevated tumor markers (CEA and CA19-9) in the serum.146 Overexpression of PRL-3 can also promote the proliferation and invasiveness of CRC cells, mainly by activating STAT3 to increase the expression of miR-17, miR-19a and miR-21.147 There is evidence that ubiquitin-specific protease 4 drives CRC invasion and metastasis via binding with and deubiquitinating PRL-3 to stabilize PRL-3.148 Moreover, PRL-3 improved IL-8 secretion in CRC cells and mediated enhancement of glycolysis, which in turn contributed to the promotion of cancer metastasis.149 PRL-3 also promoted CRC invasion and metastasis by upregulating chemokine ligand 26 (CCL26) to induce cancer metastasis.150 Recently, it was shown that PRL-3 promoted cell metastasis via activating MAPK pathways in TAMs to initiate EMT, and the NF-κB pathway activated by PRL-3 contributed to angiogenesis in CRC cells.151
PRL3 promoted metastasis of cancer cells by regulating a diverse downstream prometastatic effector molecule, via NF-κB pathway, AKT, STAT3, EGFR, IL-8 and CCL26.
CRLM and Notch pathway
Furthermore, there are other signaling pathways that are related to CRLM. The Notch pathway plays a positive role in CRLM and inhibition of Notch signaling interferes with CRLM.122 The human Notch system includes four Notch receptors (Notch1-4) and five Notch ligands (Jagged1-2, DLL1, DLL3, DLL4).152 Activation of the Notch pathway is related to the poor prognosis of CRC. Notch1 gene copy number amplification may indicate a decrease in patient survival.153 The positive expression of Notch3 protein is an unfavorable prognostic factor of disease-free survival (DFS) and overall survival (OS).154 However, the functions of notch2 and notch4 are opposite to them, their overexpression can block the proliferation, invasion or migration of cancer cells.155,156 Meanwhile, multiple Notch signaling pathway related mechanisms play a significant role in metastatic CRC(Fig. 6)
The enhanced aggressiveness of colorectal tumors caused by Notch signal activation may be related to EMT. Retroviruses are used to construct colorectal tumor cells that can stably express Notch-1 intracellular domain (NICD-1) in the cytoplasm, and the Notch ligand Jagged-1 and CD44, Slug, Smad-3 and other EMT-related proteins are found to be upregulated in these CRC cells.157 Further treatment of the cells with γ-secretase inhibitor DAPT can effectively inhibit this process. Interestingly, NICD-1 produced by retroviruses is not affected by DAPT, which suggests that this may be caused by a more complicated mechanism-after the activated Notch1 receptor releases NICD, the expression of the ligand Jagged1 is upregulated. Animal experiments have shown that when Notch/TGF-β2 signaling is inhibited, it is accompanied by a decrease in neutrophils in the target organs of metastasis, as well as the gradual accumulation of CD3+, CD4+ and CD8+ T cells, resulting in a significant reduction in metastasis.158 In addition, in the EMT-related literature of other types of cancer, it has also been reported that hey-1, an important downstream target gene of the Notch signaling pathway, can regulate the TGF-β/Smad signaling pathway, and the TGF-β-dependent signaling pathway is closely related to the induction of EMT phenotypes.159 This indicates that the crosstalk of these two signaling pathways may play a more essential role in the process of colorectal tumor metastasis.
Song et al.160 confirmed that autocrine motor factor receptor (AMFR) and notch1 are the direct target genes of miR-139-5p in colorectal tumors, upregulation of miR-139-5p promotes the expression of AMFR and NOTCH1 to enhance the migration and metastasis of CRC. In addition, knockdown of Notch ligand Jagged1 can lead to a decrease in Notch signal transduction activity of CRC cells, thereby reducing the migration of CRC cells. Moreover, in Jagged1 silenced nude mouse xenograft tumors, tumor metastasis markers MMP-2 and MM-9 are also significantly downregulated.161 Ligand DLL4 can upregulate MUSASHI-1 (MSI-1) through a Notch3-mediated mechanism, while MSI-1 inhibits the translation of Notch signaling negative regulator NUMB, and then reduces the degradation of NICD in tumor cells, and promotes the activation of Notch signaling pathway.162 In addition, Amino-terminal Enhancer of Split (AES) can inhibit the metastasis of colorectal tumors by lowering Notch signal.163 In the past, there was little knowledge about the regulation of AES expression. The latest research found that it may be affected by the human Casein kinase 1δ/ε (CK1δ/ε), meanwhile CK1δ/ε is closely related to Wnt signal and Hedgehog signal in the occurrence and development of CRC.164
Ectopic expression of tRNA-derived fragments (tRF)/miR-1280 reduced cell proliferation and metastasis by directly inhibiting the Notch signaling via targeting JAG2, leading to the decreased activity of Notch pathway components and Notch signaling.165 Moreover, the intensity of Notch1 expression was related to depth of invasion, tumor node metastasis (TNM) staging and lymph node metastasis of CRC.166 Inhibition of tumor-derived Tumor-Derived Laminin α5 (LAMA5) activated the Notch pathway in tumor endothelial cells, thereby reducing branching angiogenesis.167
MSI-1, MUSASHI-1; CK1δ/ε, Casein kinase 1δ/ε; NICD, Notch intracellular domain; EMT, epithelial-to-mesenchymal transition.
CRLM and TGFβ signaling
The TGF-β superfamily signaling contains more than 30 members, among which the more important ones are TGF-β, Activins, Nodal, Bone Morphogenetic Proteins, etc.168 In mice with progressive liver metastatic disease, blockade of TGFβ signaling rendered tumors susceptible to anti-programmed cell death 1 (PD1) therapy.169 TGF-β1 could also downregulate the E-cadherin expression and increase the Vimentin expression, inducing EMT to promote the invasion and migration of CRC.170 Moreover, CRC-derived CXCR4 activated HSCs to release SDF-1, resulting in TGF-β1 secretion in CRC cells to promote liver metastasis of CRC.171 Therefore, TGF-β1 signaling blockage may be an effective clinical strategy for CRLM.
TGF-β plays a dual role in the tumorigenesis process as illustrated in Fig. 7. In the early stage of tumorigenesis, TGF-β plays a role in inhibiting tumors. However, in the later stages, TGF-β can promote EMT and is closely related to tumor invasion and metastasis.172 In CRC, CMS4 type has a higher degree of TGF-β activation.173 Studies have shown that TGF-β1 can promote the occurrence of EMT by activating a variety of transcriptional regulators including SNAI1/2, Twist and ZEB1/2, etc.174 Increased expression of TGF-β1 can increase the metastasis of CRC cells, leading to the secretion of IL-11 by CAFs to trigger the STAT3 signaling pathway.175
Equally important, SMAD4 is also an essential molecule in the TGF-β signaling pathway. It has been reported that up to 62% of CRC patients with liver metastases have SMDA4 downregulation176, and the loss of SMAD4 has been shown to cause abnormal activation of STAT3 in pancreatic cancer cells, which then makes the expression of E-cadherin decrease, N-cadherin and vimentin increase, leading to the occurrence of EMT.177 By constructing gene-virus CD55-Smad4 and overexpressing SMAD4, Xiao et al. confirmed that the metastasis and tumor cell stemness of CRC cells can be effectively inhibited by activating the Wnt/β-catenin signaling pathway.178 In addition, there are a significant amount of studies confirming that some ncRNAs, affect the invasion and metastasis of CRC by regulating the TGF-β/Smad axis. For instance, LINC00941 promotes CRC metastasis via activation of the TGF-β/SMAD2/3 axis179, and lncRNA ANRIL promotes CRC metastasis through activating the let-7a/TGF- β 1/Smad Axis.180
CXCR4 secreted by CRC cells promoted SDF-1 secretion by HSCs. SDF-1 in turn acted on CRC and promoted its secretion of TGF-β. TGF-β can not only promote the secretion of IL-11 by CAFs to activate the STAT3 signaling pathway, but also further regulated gene transcription through SMAD in CRC cells and promoted the occurrence of EMT.
Other molecular targets and pathways involved in CRLM
L1CAM is a marker and mediator of metastasis initiating cells, which are required for orthotopic carcinoma propagation, liver metastatic colonization and chemoresistance.181,182 L1CAM is induced after loss of epithelial integrity, it then promotes tumor growth and metastasis.183 The mechanism that epithelial cells enter into an L1CAM high phenotypic state is epithelial dissociation causes transcriptional downregulation of REST, in turn reducing REST occupancy of an L1CAM intronic enhancer and promoting L1CAM expression. Research and development of L1CAM inhibitory molecules-as a treatment of diffuse residual disease and metastatic disease is of great significance.183 The scattered Lgr5 low and L1CAM high cells in the primary tumor can trigger the distant metastasis of CRC, but will dynamically emerge from the Lgr5 high primary tumor when the epithelial integrity is lost during the self-destructive tumor infiltration process, which is a distant metastatic organ necessary for the survival and re-growth of diffuse cancer cells. Future studies may elucidate the close relationship between wound healing and metastasis.
Akt2 is one of the three subtypes in the Akt family, as it promotes cell movement/invasion.184,185 Akt2 is highly expressed in stage IV CRCs and liver metastases, meanwhile, loss of PTEN and overexpression of Akt2 synergistically promote metastasis.186 The inducible loss of Akt2 in CRC cells strongly upregulates metastasis Suppressor 1 (MTSS1) at the messenger RNA and protein levels.187 MTSS1 is a new gene regulated by Akt2, where the inhibition of MTSS1 is a key step for Akt2 to promote metastasis in CRC cells. Moreover, Elevated eukaryotic translation initiation factor 4A (EIF4A) in CRC patients was associated with poorer survival rate, poor response to oxaliplatin and more distant metastasis.188 The EIF4A inhibitor silvestrol and oxaliplatin have a synergistic effect, and the combined use of the two may represent a new treatment strategy for patients with CRLM. The Trop-2 expression is another necessary for tumorigenesis and invasiveness of CRC cells.189 Interestingly, ZFP57, an embryonic stem cell-specific transcription factor has been shown to promote liver metastasis of CRC.190 Tang et al.191 found phosphoprotein enriched in astrocytes-15 kDa (PEA15) was highly expressed in CRLM patients compared to non-metastatic. PEA15 promote CRC proliferation and the abilities of invasion and migration though activating the ERK/MAPK signaling pathway. PEA15 and ZFP57 may be a potential therapy biomarker for CRLM.
There are various epidermal growth factor-like domain protein 6 (MEGF6) in CRC, which induce EMT via transforming growth factor β (TGFβ)/SMAD signaling pathway to promote the transfer of CRC.192 Through the analysis of expression profile microarray data, it is concluded that apolipoprotein E (ApoE) expression in normal mucosal tissues, primary CRC and CRLM increased sequentially.193 The overexpression of ApoE is related to the progression of CRC, especially for stage II and simultaneous liver metastases, resulting in a poor prognosis for CRC patients. Moreover, lgG Fc binding protein RNA and protein were significantly downregulated in metastatic lesions, which were associated with the prognosis of CRLM.194 In addition, two members of the S100 gene family, S100A6 and S100A4, are thought to be involved in the invasion and metastasis of cancer. When cancer cells form a glandular structure again in the center of metastatic nodules, the expression level of S100A6 decreases.195,196 Knockdown of S100A4 restricts metastasis formation in a xenografted mouse model of CRC.197 Moreover, S100A8, another member of the S100 family is correlated with TIMP-1-induced PMN in liver.198
A new member of the F-box protein family, FBX8, containing F-box and Sec 7 domain, can ubiquitinate and degrade HIF-1α such as CDK4 and C-Myc, downregulating their ability for promotion of angiogenesis, cycle progression and cell proliferation, respectively, thus regulating the CRC liver metastasis dormancy.199 Moreover, Thrombospondin-1 (THBS1) depletion inhibited migration and invasion of CRC cells through attenuating EMT.200 Upregulated THBS1 may be significantly correlated with CRLM, which requires further study.
Hypermethylated B4GALT1 was detected in the mCRC case group and had adverse prognostic effects on CRC.201 Glyco-genes B4GALT1 may act as an additional novel biomarker for CRLM. Moreover, Ferritin Light Chain (FTL) as an oncogene leads to CRC cell resistance against 5-FU treatment and promoted CRC metastasis by competing with lncRNA Linc00467 for miR-133b binding site.202 The expression of defensin β 4 A (DEFB4A) was significantly upregulated and experimentally proved that DEFB4A gene knockdown proved that DEF4BA promoted cell migration.203 Sialyltransferase ST6GAL1 can increase the stability of ICAM-1 through sialylation, thereby inhibiting the transfer characteristics of CRC.204
Under hypoxic conditions, loss of the tumor suppressor p53 (encoded by TP53) provides cancer cells with a selective advantage, hypoxia which causes resistance to therapy and promotes metastasis in CRC.205,206 Recently, Li et al. reported the hypoxia inducible factor 1 alpha subunit (HIF1A) directly repressed the miRNA-34a in p53-defective CRC cells under hypoxia.207 Conversely, p53 increases expression of miRNA-34a in CRC cells without hypoxia. Inhibition of Protein phosphatase 1 regulatory inhibitor subunit 11 (PPP1R11) by miRNA-34a prevented activation of STAT3 and inhibited the hypoxia-induced EMT and metastasis. Similarly, upregulated PPP1R11 was associated with TP53 mutations and metastasis to the liver. Targeting this pathway may represent a therapeutic opportunity for managing metastatic disease. The NF-κB/p65 signaling pathway has a critical role in the mediation of GNA13 in CRC which was indicated by a decrease in GNA13-induced migration, invasion, and in addition, CXCL chemokine level increases after the inhibition of NF-κB/p65 signaling pathway with an inhibitor.70 The novel quinazoline derivative MJ-56 interfered with the NF-κB signaling via impairing PI3K/AKT activation and subsequently reduced the NF-κB-mediated transcription of MMPs.208
Chu et al.209 reported that mutant KRAS transcriptionally activates IGF-IR gene expression through Y-box-binding protein (YB)-1 upregulation via a novel MEK-Sp1-DNMT1-miR-137 pathway in CRC cells. Moreover, suppression of the expression of YB-1 and IGF-IR via inhibition of MEK hampers KRAS-driven CRLM in animal model studies. KRAS-driven MEK-Sp1-DNMT1-miR-137-YB-1-IGF-IR signaling pathway, which might provide a mechanistic rationale for the use of a MEK inhibitor as an adjuvant, in combination with standard of care, to prevent the recurrence of CRLM in KRAS mutant CRC patients after receiving liver resection, however, further investigation is needed. Analysis of the TCGA database confirmed the upregulation of retinoblastoma binding protein 4 (RBBP4) in CRC tissues, and the overexpression of RBBP4 is related to nerve infiltration and poor chemotherapy effects.210
NcRNAs in CRLM
NcRNAs include long non-coding RNA (lncRNAs), microRNAs (miRNAs) and circular RNAs (circRNAs), which are emerging as the master regulators of cancer. The dysregulated expression NcRNAs have coding-independent functions in the progression of CRLM. In this section, we discuss some lncRNA and their role in CRLM (Tables 2, 3), circRNAs in CRLM (Table 4), miRNAs in the regulation of cascade of CRLM (Table 5).
LncRNAs in CRLM
Recently, Zhang et al.211 demonstrated that lncRNA LALC recruited DNA methyltransferases (DNMTs) to the LZTS1 promoter by coupling with EZH2 and then modified the expression of LZTS1 through DNMTs-mediated DNA methylation in CRLM. Methyltransferase like 3 behaves as the ‘writer’ of m6A promoted m6A-methylation of oncogenic lncRNA XIST to inhibit CRC proliferation and invasion.212 LncRNA LINC01578 activity was enhanced when nuclear factor kappa B (NF-κB) and Yin Yang 1 (YY1) were directly bound to the LINC01578 promoter.213 Meanwhile, upregulated LINC01578 interacted with and recruited EZH2 to NFKBIB promoter and further repressed NFKBIB expression, thereby activating NF-κB signaling. LINC01578 and NF-κB/YY1 formed a positive feedback loop, which promoted CRC metastasis. LncRNA HOTAIR suppressed HNF4α via recruiting SNAIL to promote the migration, invasion and EMT of CRC.214 Intriguingly, a novel lncRNA, ENSG00000274093.1 binds to histone deacetylase 2 (HDAC2) and may act as a modular scaffold for the HDAC1/HDAC2 and EZH2 complexes, thereby altering EMT in CRC.215 LncRNA TPT1-AS1 induces angiogenesis and metastasis in CRC via theTPT1-AS1/NF90/VEGFA axis.216 Additional research have shown that the expression level of LncRNA clmat3 and SNHG15 in CRC with liver metastasis was significantly higher than in those CRC without liver metastasis.217,218 Conversely, some lncRNAs play a role of tumor inhibition, for example, low expressed lncRNA SATB2-AS1 inhibited cell metastasis and regulated the immune response of CRC by cis-activating SATB2.219 Moreover, lncRNA MIR22HG, as a tumor suppressor in CRC, competitively interacted with SMAD2 and modulated the activity of TGFβ pathway, thereby inhibited cell survival and tumor metastasis in vitro and in vivo.220
LncRNA UICLM acted as a ceRNA for miR-215 to regulate ZEB2 expression, then induced CRC liver metastasis, which may offer a novel prognostic marker and therapeutic target for this disease.221 LncRNA GAPLINC, HOTAIR, RP11-757G1.5, LINC00460, MIR4435-2HG, PlncRNA-1, DANCR and TP73-AS1 were proven to be positively associated with the proliferation and liver metastasis of CRC through the GAPLINC/miR-34a/c-MET axis, HOTAIR/miR-214/ST6GAL1 axis, RP11-757G1.5/miR-139-5p/YAP1 axis, LINC00460/miR-613/sphingosine kinase 1 (SphK1) axis and MIR4435-2HG/miR-206/YAP1 axis, PlncRNA-1/miR-204/Wnt/β-catenin regulatory network, DANCR/miR-577/HSP27 signaling axis and TP73-AS1/miR-194/TGFαsignaling axis, respectively.222,223,224,225,226,227,228,229 Furthermore, lncRNA ZEB1-AS1 acted as ceRNA to upregulate p21-activated kinases 2 (PAK2) by sponging miR-455-3p, thus facilitating colon adenocarcinoma cell growth and metastasis.230 Zhou et al.231 reported that SRY-box transcription factor 9 (SOX9) activated the transcription of lncRNA FARSA-AS1. FARSA-AS1 upregulated SOX9 and FARSA via binding to miR-18b-5p directly. Overall, SOX9-FARSA-AS1-SOX9/FARSA loop participated in cell growth, stemness, and metastasis in CRC. In addition, lncRNA HSD17B11-1:1 acted as a sponge for miR-338-3p to upregulate the expression of MACC1 to promote CRC cell proliferation, mobility and invasion in vitro and in vivo.232 B3GALT5‐AS1, is an antisense lncRNA, located in Chr21q22.2, which was first reported in Wang et al. study.233 They found B3GALT5‐AS1 was decreased in CRC contrast to normal colonic epithelium. B3GALT5-AS1 directly bound to the promoter of miR-203, repressed miR-203 expression, upregulated miR-203 targets ZEB2 and SNAI2, and induced EMT, suggesting that downregulated B3GALT5‐AS1 is a biomarker of CRC metastasis and, more importantly, activating B3GALT5-AS1/miR-203/EMT axis may be a potential therapeutic strategy for CRLM.233 Recently, B3GALT5‐AS1 has been reported to be related to TNM stage and histological differentiation.234 There are also other lncRNA that regulated CRLM through ceRNA mode, for example MALAT1, SNHG14, SNHG7, H19, and LINC00485. Taken together, these lncRNA could be a therapeutic target and promising biomarkers for prognosis prediction of CRLM.
CircRNAs in CRLM
NSUN2 is a circRNA recently identified upregulated in CRLM and N6-methyladenosine modification of circRNA among which circRNA circPPP1R12A activated the Hippo-YAP signaling pathway to promote the growth and metastasis of CRC235. NSUN2 was more likely to enter the cytoplasm, then in the cytoplasm increased the stability of HMGA2 mRNA to induce CRC metastasis progression.236 Equally noteworthy, circRNA circ_0124554 inhibited the ubiquitination of AKT to promote the early metastasis particularly for the lymph node-negative CRC patients with synchronous liver metastasis.237
CircRNAs serve as a sponge to protect multiple oncogenes from being attacked by miRNAs and among these, global exaltation of circ_0115744 circRNAs circAPLP2, hsa_circ_000984, hsa_circ_0071589 and hsa_circRNA_102209 acted as ceRNA to enhance tumor metastasis via circ_0115744/miR-144/EZH2 axis, circAPLP2/miR-101-3p/Notch1 axis, hsa_circ_000984/miR-106b/CDK6 axis, hsa_circ_0071589/miR-600/EZH2 axis and hsa_circRNA_102209/miR-761/RIN1 axis, respectively.238,239,240 Similarly, circRNA NSD2 promoted metastasis of CRC by sponging miR-199b-5p, which could increase the expression of DDR1 and activate JAG1 signaling.241 More interestingly, one circRNA contains various binding sites for different miRNAs reveals the complex role of circRNA in cancer malignancy. CircRNA CCDC66 functions as miRNA sponge to reduce the destruction of MYC mRNA by miRNA-33b and miR-93.242 In addition, Zhi et al.243 also identified a novel and conserved circRNA hsa_circ_102049 as a promoter of CRC metastasis. Mechanistically, hsa_circ_102049 acted as a sponge of the tumor suppressor miR‐761 and miR‐192‐3p and modified the subcellular localization of the RNA binding protein DGCR8, then regulated the levels of the FRAS1, which synergistically enhanced the adhesion, migration, and invasion abilities of CRC cells. Interestingly, hsa_circ_0001178 sponging miR-382, miR-587 and miR-616 to upregulate ZEB1, which in turn increase hsa_circ_0001178 expression via physically binding to hsa_circ_0001178 promoter region, thereby based on this positive feedback ceRNA axis, consequently facilitated the invasion and metastasis of CRC.244
circRNAs also play an inhibition role of CRLM, for example, circRNA Hsa_circ_0009361 acted as an inhibitor for miR-582 and suppressed the CRC cell metastasis.245 Similarly, circRNA-FNDC3B negatively and directly regulated miR‐937‐5p to promote the expression level of the tumor‐suppressor TIMP3, thereby blocking the metastasis, invasion and angiogenesis of CRC.246 In addition, circITGA7 inhibited the Ras signaling pathway and promoted the transcription of ITGA7, thereby suppressed the proliferation and metastasis of CRC cells.247
In the latest studies, circRNA_0001178, circRNA_0000826 and circRNA hsa_circ_0000826 induced by the hypoxia in CRC were dramatically upregulated in CRLM tissues, which can be a potential biomarker of CRC liver metastasis.248,249 Validation studies of specific molecular mechanisms are needed to identify optimal miRNAs and marker panels that can be used in clinical care.
MiRNAs in CRLM
Recent studies have shown that miRNAs were involved in the context of EMT in CRC. It has been reported that miR-200 family (miR-200a, miR-200b, miR-200c, miR-141, and miR-429) are recognized as regulators of the epithelial phenotype through repression of ZEB1 and ZEB2 mRNA translation250,251,252 The miR-200c levels in primary CRC without liver metastasis is lower than the metastasis with primary tumor tissues, which highlight a crucial role for miR-200c in CRC metastasis.251 The overexpression of miR-429 could play an oncogenic role in the cellular processes of CRC by targeting SOX2.253
Additionally, miR-181a and miR-30b were highly expressed in CRLM by promoting EMT through inhibiting Wnt inhibitory factor-1 (WIF-1) and inhibiting the SIX1 gene, respectively.254,255 Moreover, miR-30a is an important regulator of transmembrane-4-L-six-family protein (TM4SF1), VEGF, and E-cadherin for CRC cell motility and EMT.256 Manganese superoxide dismutase (MnSOD) was required for downregulation of epithelial markers and upregulation of mesenchymal markers in CRC cells, indicating that it promoted the EMT, which was reduced by overexpression of miR-212.257 Methylated miR‐34c‐5p significantly suppressed the metastasis of CRC cells via directly modulating the SATB2.258 Recently, the cytokine IL-6 was found to activate the oncogenic STAT3 transcription factor, which directly repressed the MIR34A gene, while miR-34a directly regulated IL-6R. As a result, the IL-6R/STAT3/miR-34a loop promoted CRC invasion and metastasis.56 MiR-186-5p affected metastasis and EMT process of CRC cell by inhibition of ZEB1, while miR-17-5p regulated EMT by targeting vimentin.259,260
Furthermore miR-34a/SNAIL loop and miR-200/ZEB1/2 loops, miR-15a/16-1/AP4 feedback loop is also found in primary CRC. The tumor-suppressive miR-15a and miR-16-1, which targeted AP4 3′-UTR, and inhibited CRC cell migration and invasion.261,262 Overexpression of miRNA let-7 promoted EMT via targeting HMGA2263, while Lin28 could inhibit let-7 in conjunction with OCT4, SOX2, and KLF4.264 MiR-10a suppressed CRC metastasis by regulating the EMT via targeting matrix metalloproteinase 14 and actin gamma 1 (ACTG1).265 Unlike miR-10a, upregulated miR-10b in metastatic CRC tissues and cell lines inhibited E-cadherin expression and enhanced cyclin D1, which were partly abrogated after targeting KLF4.266
The expression of miR-21 was higher in tumor tissue than in adjacent normal tissue of 156 CRC patients by TaqMan MicroRNA assays.267 It is reported that transglutaminase 2 (TG2) expression was observed in CRC primary tumors but lost in liver metastases and TG2 inhibited by miR-19 could affect the invasive ability of CRC cells.268 Moreover, Zhang et al.147 demonstrated that the overexpression of PRL-3 in CRC cells induced the expression of miR-21, miR-17 and miR-19a by activating signal transducer and STAT3. A positive correlation was observed between PRL-3 and these miRNAs in matched primary colon cancer tissues and metastatic lesions.147 MiR-21 could significantly reduce Pdcd4-protein amounts and increased invasion.269 Additionally, Feiersinger et al.270 have identified that the expression of miR -21 was significantly lower in liver metastases as compared to the primary CRC. The overexpression of miR-200 and miR-141 inhibited apoptosis and induced migration.271 These indicated that miR-21 might be involved in the initiation and miR-200 and miR-141 exacerbated liver metastasis of CRC.
Interestingly, highly expressed miRNA-20a-5p and miR-298 were positively correlated with CRLM by suppressing drosophila mothers against decapentaplegic protein4 (smad4) expression and targeting PTEN, respectively.272,273 Inhibition of miR-15b significantly decreased colony formation ability, invasion, and migration of HCT116 cells in vitro and liver metastasis of HCT116 tumors in vivo via increasing metastasis suppressor-1 (MTSS1) and Klotho protein expression.274,275 Upregulation of miR-885-5p had strong tumor-promoting effects through by targeting cytoplasmic polyadenylation element binding protein 2 (cpeb2), von Willebrand factor and insulin-like growth factor binding protein 5, and its potential role in promoting cell migration, invasion and liver metastasis.276,277 The expression of miR-224 increased consistently with tumor burden and the steady state of microsatellites, and can enhance CRC metastasis in vitro and in vivo.278
For long, it was known that the patients with CRC have demonstrated a significant evaluated expression of serum miR-497.279 In the mouse model, Qiu et al.280 found that a combination of miR-497 and bufalin had a synergistic effect on the inhibition of CRC metastasis. They also found that miR-497 targeted the expression of vascular endothelial growth factor-A (VEGF-A).281 Notably, miR-497 exerted its oncogenic function by targeting fos-related antigen-1 (Fra-1).282 Recently, AGAP2-AS1 regulated fibroblast growth factor receptor 1 (FGFR1) expression by sponging miR-497 in the migration and invasion of CRC cells.283
However, liver metastasis cells of CRC also downregulate the expression of miR-133a, miR-17-5p, miR-99b-5p, miR-214, miR-26, miR-30e-5p and miR-328-3p that suppress cell migration and invasion. MiR-133a, as a tumor suppressor, inhibited cell proliferation, invasion, and migration by targeting oncogenic EIF4A1.284 MiR-99b-5p was differently expressed in primary CRC and liver metastasis and functioned as a tumor-suppressive miRNA to affect cell migration by targeting mTOR in metastatic CRC.285 In an orthotopic mouse model of CRC, Ding et al.286 found that APOBEC3G enhanced CRC cell migration and invasion via inhibition of miR-29-mediated suppression of MMP2. Downregulation of miR-214 promoted proliferation, migration, and invasion in CRC cell lines via increasing level of FGFR1, which can lead to the occurrence of liver metastasis.287,288 Li et al.289 reported that tumor-suppressive miR-26a and miR-26b inhibited the target gene FUT4 expression, resulting in migratory behavior of CRC. Laudato et al.290 found that miR-30e-5p was a novel effector of P53-induced suppression of migration, invasion, and metastasis by directly targeting both integrin alpha‐6 (ITGA6) and integrin beta‐1 (ITGB1). In addition, miR-30b-5p functioned as a metastasis suppressor by targeting Rap1b, a Ras family small GTPase that regulates cell adhesion and mobility.291
A recent study has shown that low miR-196b-5p expression is significantly associated with metastases and poor outcomes, further proving that miR-196b-5p inhibition led to significantly increased CRC cell migration/invasion and metastases formation in mice via the interaction with HOXB7 and GALNT5.292 A contemporary study by Luo et al. has demonstrated that miR-432-5p functions as a tumor suppressor to inhibit cell migration and invasion by negatively regulating CXCL5 expression.293 Interestingly, miR-328-3p may inhibit proliferation and metastasis of CRC cells via suppressing Girdin expression and associated PI3K/Akt signaling pathway294 MiR-365a-3p to inhibit the metastasis of CRC cells by negatively regulating ADAM10 and inactivating the JAK/STAT signaling pathway.295 Moreover, miR-487b directly targeted LRP6, a receptor for WNT/β-catenin signaling to inhibit liver metastasis.296 MiR-143-3p, miR-132 and miR-146a significantly abolished the development of liver metastases by directly targeting ITGA6/ASAP3, anoctamin 1 (ANO1) and c-Met, respectively.297,298,299 In addition to control the intravasation, miRNAs also targeted genes; for example, miR-21 regulated Pdcd4 which induced intravasation and metastasis.269 A positive significant correlation between expression of miR-126 and epidermal growth factor–like domain 7 (EGFL7) were seen in liver metastases, which supported miR-126 may act as a regulator in angiogenesis and intravasation process.300 Further experiments are needed to explore more detailed forms of connection between miR-126 and EGFL7. Low expression of miR-26b was significantly associated with the invasiveness and metastasis of CRC cells. Hansen et al.301 reported that miR-31-mediated repression of E-selectin impaired the metastatic potential of CRC cells.
One study showed that miR-103/107 potentiated the colonization of CRC cells at a metastatic site by targeting the known metastasis suppressors death-associated protein kinase (DAPK) and KLF4 in CRC cells.302 Similarly, in mice models of CRC, the overexpression of miR-103/107 enhanced local invasion and liver metastasis effects. Another report revealed that miR-483 and miR-551a inhibited liver colonization and metastasis.303 Metastatic foci in new organs promote angiogenesis due to increased demand for oxygen and nutrients. Therefore, angiogenesis has become a necessary condition for the survival of metastatic foci.
Cancer stem cells (CSCs) in the progression to CRLM
CSCs as seed cells for tumorigenesis can initiate and sustain the growth of tumors. CSCs can be identified by specific markers, including CD133, LGR5,CD44, ALDH1, CD24, CD166, CD29, CD26, and CD51.304,305 CRC-CSCs can increase the ability of distant metastasis and colonization.306,307 It is thus possible that targeting CSCs may have widespread clinical implications.308 Among the epithelial cells that form the surface of the intestine, crypt-based columnar cells (Lgr5+ cells) expressing the receptor protein Lgr5 function as stem cells during the maintenance of intestinal homeostasis, and they are also the starting cells for CRC.309 The intestinal epithelium has significant self-renewal ability, and the rapidly proliferating Lgr5+ cells are usually responsible for the daily production of all types of intestinal epithelial cells.309 Lgr5− cells can drive wound healing by first transdifferentiating into a stem-like Lgr5+ state.310 The selective elimination of LGR5+ cells leads to temporary tumor regression, and other cells exhibit compensatory proliferation, then the reappearance of LGR5+CSCs drives tumor regeneration.311 It has been demonstrated that selective Lgr5+ cell ablation restricts primary CRC growth, but does not result in tumor regression, because proliferative Lgr5− cells could continuously attempt to replenish the Lgr5+ CSCs pool, leading to rapid re-initiation of tumor growth upon treatment cessation.312 Moreover, Fumagalli et al. provided direct evidence most disseminated CRC cells in circulation were Lgr5- and formed liver metastases in which Lgr5+ CSCs appeared.313 Cell state plasticity is very important to promote CRC metastasis. The signaling pathways responsible for the reproduction of Lgr5+ cells may yield exciting new strategies for treating CRLM. Wang et al.314 reported prostaglandin E2 induced CSCs and enhanced liver metastasis by activating NF-κB via EP4-MAPK and EP4-PI3K-Akt pathways.
DNA methylation is considered to be a potential epigenetic mechanism to maintain CSCs, and the loss of DNMT can reduce the occurrence of tumors by limiting the CSC pool.315 Li et al.316 found that the DNMT inhibitor 5-Aza-2'-deoxycytidine (5-AzaDC) significantly reduced the abundance of colorectal CSC in vitro and inhibited the growth of liver metastatic tumors in vivo. They also found that 5-AzaDC inhibits the expression of active β-catenin and downregulates the Wnt signaling pathway. A new study points out the mechanism exerted by methylation on endothelin system genes expression can be compromised in CRLM.317 Notably, CSCs are critical for the formation and maintenance of liver metastasis. Together, our data highlights distinct CSC dependencies for primary versus metastatic tumor growth and suggest that targeting CSCs may be one direction of treatment for CRLM.
Role of circulating tumor cells (CTCs) in CRLM
Patient-derived CTCs has been shown to bear all the functional attributes of CSCs.318 The markers expressed by CTCs are similar to the cancer niche, which are conducive to liver metastasis.319,320 CD133+CD44 + CD54 + cellular subpopulation of CTCs has a prognostic value in CRLM, especially in the survival of CRLM who did not receive surgical treatment for metastasis.321 Patient-derived CTCs lines are tumorigenic in subcutaneous xenografts and are also able to colonize the liver after intrasplenic injection.318 Drug test by in vitro culture of CTCs may facilitate access to personalized medicine. TAMs regulated JAK2/STAT3 signaling pathway by secreting IL6, thereby inhibiting miR-506-3p expression and promoting FoxQ1 expression. CTC cells then produced CCL2 to recruit more TAMs. TAMs and CTC both interacted to promote the occurrence of metastasis.322 These findings suggest targeting strategies against CTC clusters may be effective in the treatment of liver metastases.
Metabolic factors in CRLM
Tumors are also a metabolic disease, and metabolic changes are closely related to every process of tumor metastasis. The process of transfer requires a large amount of energy supply, and some enzymes and molecules that affect energy metabolism also act as gas stations in the process of transfer. In the extracellular space, creatine kinase brain (CKB) used ATP-catalyzed phosphorylation of the metabolite creatine to form phosphocreatine, which can produce large amounts of ATP into CRC cells as an energy reserve to maintain the energy requirements of CRC cells during anoxia during metastasis.323 CKB promoted the development of liver metastasis, and targeted inhibition of its activity was also the direction of future treatment of CRLM. In addition to phosphocreatine generating energy for cancer cells, the fatty acid oxidation (FAO) pathway is also an important energy source within cancer cells. In detached CRC cells, under the action of carnitine palmitoyl transferase 1, FAO was greatly activated, thus increasing the ability of cells to metastasize.324 CRC cells implanted in the liver promoted the metabolism of fructose by upregulating enzyme aldolase B (ALDOB), which provided energy for cell growth in liver metastases.325 This particular metabolic pathway change was found only in liver metastases and not in other sites of metastasis or primary tumors. All in all, the metabolic pathways of tumors adapt to the environment during metastasis, and there are still many problems to be studied.
Diagnosis
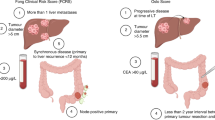
CRLM should be identified in patients with a confirmed diagnosis of CRC. Among them, ~10–15% of patients have synchronous liver metastases—the liver metastasis is diagnosed at the same time with CRC.10 The main content of our diagnostic section is centered on synchronous liver metastases, and the assessing approaches of metachronous liver metastases are similar to the former and executed during the follow-up period.
Biomarker testing should be performed routinely for diagnosis of the synchronous liver metastasis. Serum cancer embryo antigen (CEA) could be the routine choice for screening, and carbohydrate antigen 19-9 (CA19-9) might be a supplementary indicator without an evaluated CEA value.326 Detection of CEA and CA19-9 should also be executed at intervals after resection. RAS is also a biomarker predicting the efficacy of anti-epidermal growth factor receptor (EGFR) therapy, which determines the choice of treatment strategy. Thus, RAS testing should be executed in all patients with CRC, and include NRAS exons 2, 3, 4 and KRAS exons 2, 3, 4 at least.327 BRAF testing also could help to select the therapy option and predict the prognosis. In addition to serum tumor marker examination and pathological staging evaluation, imaging examinations such as liver ultrasound and abdominal contrast-enhanced CT should be routinely performed to screen and diagnose liver metastases.328
Magnetic resonance imaging (MRI) with liver specific contrast should be used prior to surgery when the liver metastases are resectable. Moreover, MMR/microsatellite instability (MSI) testing, UGT1A1 detection of UGT1A1 protein, human epidermal growth factor receptor 2 detection are recommended to provide the basis for clinical decisions on post line treatment in advanced patients. The PET-CT examination is not used as a regular recommendation and may be applied as appropriate when the condition is required. Needle biopsies of liver metastases are rarely required.329
After radical CRC, patients should follow up regularly to understand whether the occurrence of metachronous liver metastases. During CRC surgery, routine exploration of the liver must be performed to further exclude the possibility of liver metastases. Suspicious liver nodules found during the operation can also be biopsied.329 Tumor markers such as serum CEA, CA19-9 and, liver ultrasound and chest/abdominal/pelvic enhancement CT scan should be routinely examined to conduct screenings and diagnose liver metastases. Patients with high suspicion in ultrasound or CT imaging should not be diagnosed with liver MRI, and recommended consistency of imaging methods during follow-up. PET-CT scan is not normally recommended. Electronic colonoscopy should be performed within 1 year after surgery.330
Treatment
Surgery
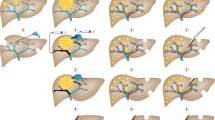
Surgery has become the standard of care for patients with CRLM. When CRLM may be limited to a few liver metastatic foci, surgical resection is the first choice. Patients with liver metastases that cannot be resected initially should also receive surgical treatment when they are transformed into resectable lesions after treatment.331 For these patients who are suitable for complete surgical resection, they are often accompanied by lymph node infiltration and dissemination of occult micro metastases. Therefore, it is necessary to cooperate with adjuvant radiotherapy and chemotherapy and targeted therapy.
Radiation therapy
In patients with CRLM, radiation therapy of liver metastasis remains controversial, due to the tolerated dose of whole liver radiation being much lower than the lethal dose required by tumor cells. The use of image guidance technology can make radiotherapy more precise. For patients with liver metastases from CRC with normal liver function, conventional radiotherapy techniques can be used to treat liver metastases.332,333
Ablation therapy
In cases where liver metastases cannot be surgically removed, appropriate ablation treatments, such as radiofrequency ablation, microwave ablation, and cryotherapy, should be selected on the basis of systemic chemotherapy based on their location, treatment goals, treatment-related complications, and the patient’s own conditions to strengthen control of local lesions.334 However, care should be taken to avoid extrahepatic heat damage and incomplete ablation when performing ablation treatment.331
Systemic therapy
Patients with CRLM usually receive chemotherapy after surgical resection. It recommends FOLFOX (5‐fluorouracil [5‐FU] + oxaliplatin + leucovorin [LV]), FOLFIRI (5‐FU + irinotecan+ LV), XELOX (capecitabine + oxaliplatin), infusional 5‐FU/LV or capecitabine, or FOLFOXIRI (5‐FU + oxaliplatin + irinotecan + LV) for patients with mCRC who are suitable for intensive therapy according to the current National Comprehensive Cancer Network guidelines.335 Research data in recent decades has shown that patients with metastatic CRC receive systematic chemotherapy, and their OS duration is extended to nearly 20 months.335,336 Currently, patients with initially unresectable CRLM has received hepatic arterial infusion chemotherapy in combination with systemic chemotherapy, which is not widely used.337
Anti-angiogenesis therapy
Tumor increases energy supply through angiogenesis during metastasis, and anti-angiogenesis therapy has become an important therapeutic strategy for CRC. It is of great significance to explore the angiogenesis mechanism of CRLM. The targeted agents for anti-angiogenesis therapy under clinical trials in CRLM are summarized in Table 6. Bevacizumab and Cetuximab are molecularly targeted drugs that have been developed specifically for endothelial growth factor receptor (EGFR). Clinical trials have proven that anti-EGFR monoclonal antibodies panitumumab and cetuximab can effectively inhibit metastatic CRC.338 Saltz et al.339 evaluated the efficacy of adding bevacizumab to XELOX or FOLFOX-4 in 1401 CRLM patients. The median duration of PFS was 9.4 months in the bevacizumab group and 8.0 months in the placebo group (P = 0.0023). Moreover, cetuximab added to FOLFOX-4 compared with FOLFOX-4 alone, cetuximab was associated with a clinically significant increase in overall response (61% vs. 37%; P compared with FOLFOX-4 alone, especially In KRAS wild-type tumors, it can reach a level of 0.011), which reduces the risk of disease progression (hazard ratio = 0.57; P = 0.0163).340 Patients with RAS wt mCRC do not respond to anti-EGFR antibodies, probably due to ineffective inhibition of oncogenic RAS signaling.341 A recent randomized study conducted by Qin et al.342 confirmed cetuximab in combination with FOLFOX as an effective standard-of-care first-line treatment regimen for patients with RAS wt mCRC. However, the NORDIC-VII multicenter phase III trial suggested that cetuximab did not add significant benefit to the fluorouracil, leucovorin, and oxaliplatin (Nordic FLOX) regimen in first-line treatment of mCRC.343
It was confirmed by clinical trials that dual chemotherapy (fluoropyrimidine plus oxaliplatin or irinotecan) with targeted drugs that add anti-EGFR antibodies can further increase the response rate to about 60%.339,344,345 Bevacizumab plus XELOX (CapeOX) chemotherapy has no negative impact on intrahepatic immune cells in resectable CRLM patients.346 Although the survival rate is improved by FOLFOX treatment, chemotherapy still has many disadvantages, such as the destruction of the human immune system, loss of appetite, weight loss, hair loss and so on after chemotherapy. In a recent study, nano codelivery of oxaliplatin and folinic acid (Nano-Folox/5-Fu) with 5-fluorouracil significantly promoted the blood circulation and tumor accumulation of drugs in the orthotopic CRC mouse model.347 Nano-Folox/5-Fu with 5-fluorouracil can not only provide anti-cancer cell toxicity, but also induce immunogenic cell death. In addition, the combination of anti-PDL1 monoclonal antibody with Nano-Folox/5-FU significantly reduced liver metastasis in mice. These results suggest that the combination strategy based on Nano Folox has some therapeutic significance in CRLM. Moreover, gene set enrichment analysis discovered that the mTOR pathway was activated in patients undergoing oxaliplatin based therapy, which suggested it could be used in combination with cytotoxic chemotherapy and targeted drugs, and may have good therapeutic effects.348 For patients with initially resectable CRLM, chemotherapy can improve progression-free survival (PFS) but cannot improve OS349, and further addition of EGFR-targeted antibodies is not beneficial.350 However, in patients with unresectable CRLM, the use of bevacizumab may further increase the proportion of patients eligible for surgical resection.351 For patients whose CRLM is initially unsatisfactory or unresectable, targeted anti-angiogenesis drugs plus chemotherapy can also produce a higher remission rate and improve resectable.352
TSU68[(Z)-5-[(1,2-dihydro-2-oxo-3H-indol-3-ylidene)methyl]-2,4-dimethyl-1H-pyrrole-3-propanoic acid; SU6668] is a potent anti-angiogenic agent. In some preclinical models, TSU68 has previously been shown to have a powerful effect in preventing liver metastasis of CRC.353,354 Inhibiting CXCL1 expression in the premetastatic liver was the detailed mechanisms by which TSU68 suppresses tumor metastases.71 Anti-angiogenic agents may modulate the PMN in target organs.
Immunotherapy
In addition to anti-angiogenesis therapy, immunotherapy becomes an attractive therapeutic option as more progress has been made in the exploration of immune checkpoint in several cancer types, mainly including DNA mismatch repair defects(dMMR)/high microsatellite instability (MSI-H) CRC and high tumor infiltrating lymphocyte (TIL) tumors. By contrast, immune checkpoint monotherapy has not demonstrated significant clinical success in patients with strong mismatch repair capacity (pMMR)/low microsatellite instability (MSI-L) CRC. Immune checkpoint drugs target the inhibitory receptors present on T cells, such as PD1, LAG3 and cytotoxic T-lymphocyte associated antigen 4 (CTLA-4).355 These cells mainly exist at the tumor-stroma interface, while pMMR/MSI-L CRC shows a conventional morphology without obvious infiltrating lymphocyte TIL.356
PD1, a novel member of the immunoglobulin gene superfamily, has two functional ligands: PDL1 and programmed cell death ligand 2(PDL2).357,358 PD1/PDL1 signaling pathway can inhibit the activation of T effector cell while inhibiting the production of interferon gamma and the release of interleukin (IL)-2 and other inflammatory cytokines, thereby enabling cancer cells to escape from the host’s antitumor immunosurveillance.359,360 Masugi et al.361 reported that tumor PD1 expression was inversely associated with MSI-H in CRC. However, Wyss et al.362 reported stromal PDL1 might function as a prognostic marker in CRC patients independent of microsatellite-stable (MSS) or with MSI-L. Masugi et al.361 found that stromal PDL1 expression was only present in 5% CRC patients, but the studies by Wyss et al.362 and Taube et al.363 displayed a stromal PDL1 expression rate of 60.9% and 50%, respectively. However, epithelial PDL1 staining is less common in CRC than in other solid tumors, such as melanoma.364 While there is an inequality of PDL1 expression between primary tumors and metastases in melanoma patients,365 PDL1 expression in primary colon tumors largely corresponds to matched liver metastases.362 Another study revealed the gene networks of EMT, angiogenesis, immune-suppression and T cell exhaustion are the key events closely associated with CRC metastasis and intrinsic anti-PD1 resistance.366 If anti-PDL1 treatment for metastatic colon cancer is needed, it may not be necessary to analyze PDL1 in liver metastases. The PDL1 and CRLM research is highly debated, therefore PD1/PDL1 signaling pathway may be closely related to CRLM and further exploration is needed.
A small number of patients with MSI-H CRC were sensitive to immune checkpoint blockade with antibodies to PD1/PDL1.367,368 The incidence of dMMR in CRLM is also low.369 Recently, a study has found that tumor mutational burden (TMB) is a predictor of response to immune checkpoint inhibitors in the MSI-H population. Patients with high TMB may respond particularly well to immune checkpoint inhibitors, and these patients can be further selected to receive anti-PD1 monotherapy rather than combination therapy (such as nivolumab) as first-line treatment.370 Nevertheless, in recent years, there have been new advances in immunotherapy for CRLM. Next, we summarized the latest progress in the clinical development of immune checkpoint blockade therapy for patients with CRLM.
CTLA-4 is predominantly expressed on T cells and inhibits the activation and response of T cells.371 In a MSS highly aggressive orthotopic mouse model of CRC, dual inhibition of CTLA-4 and PDL1 resulted in the arrest of tumor growth and entirely blocked liver metastasis. Nevertheless, inhibition of CTLA-4 or PDL1 alone only moderately reduced metastatic diffusion of CRC cells.372 Furthermore, pMMR CRLM differs immunologically from primary CRC in terms of immune infiltration.373 Zhou et al.374 focus on TIL regulated by the inhibitory receptor-ligand pathway regulates MMR-proficient CRLM. They found that blocking LAG3 enhances tumor-infiltrating T-cell responses of MMR-proficient CRLM, and may thus be a new target of immunotherapy for CRLM.
Unlike primary CRC, tumor-infiltrating regulatory T cells (Ti-Treg) in liver metastases effectively inhibit tumor-specific T cell responses by expressing high levels of Treg-related molecule glucocorticoid-induced tumor necrosis factor receptor (GITR) and CTLA-4.375,376 It has demonstrated that treating CRLM by inhibiting the inhibitory receptor CTLA4 with antibodies and activating the stimulant receptor GITR with natural ligand of GITR can reduce Ti-Treg-mediated inhibition, thereby restoring effector T cell proliferation and cytokine formation.377 More importantly, compared with any single treatment, the low-dose combination therapy of the two molecules showed a stronger ability to restore T cell function. Based on gene ontology and Kyoto Encyclopedia of Genes and Genomes pathway analyses, Liu et al.378 identified some immune genes CCL20, CCL24 and CD70 that were associated with CRLM, but needs further experimental validation.
Furthermore, the relationship of gut microbiome and immune system is nowadays rapidly changing and expanding. Host factors, including age, obesity and gut microbiome greatly affect the effectiveness of immunotherapy.379,380 Meanwhile, gut microbiome is closely linked to the development and progression of CRC. Lipopolysaccharide (LPS), an important product of intestinal Gram-negative microbiota, was also found to participate in the whole process of CRLM. Systemic inflammation caused by elevated LPS blood levels in the intestinal cavity and portal vein in CRC patients could increase the liver recruitment of cancer cells.381 LPS enhanced CRLM by stimulating Toll-like receptor 4 signaling and increasing β1 integrin-mediated cell adhesion.382 LPS was observed to promote the migration capacity of CRC cells by activating the SDF-1α/CXCR4 axis and EMT.381 These results revealed trapping LPS may prevent CRLM. Recently, Song et al.194 reported nanotechnology-based trapping LPS in the orthotopic CRC tumor could promote T-cell infiltration into tumors and promote enhanced immunotherapy.
Antartina, an antitumor agent isolated from Deschampsia antarctica Desv, potently inhibited tumor growth and liver metastases in an immunocompetent colorectal carcinoma mice model.383 Antartina induced a potent specific cytotoxic T-cell response against CRC and a long-lasting antitumor immunity. Antartina can induce antitumor immunity against CRLM. Based on the results of one phase III multicenter trial, Keytruda is superior to chemotherapy in the first-line treatment of MSI-H or dMMR CRC, resulting in longer PFS and fewer treatment-related adverse events.384
Other potential therapies
Current chemotherapy for CRLM is still not desirable owing to off-target effect. Therefore, it is necessary to develop new methods to improve target effect or even replace the existing CRLM chemotherapy. Zhao et al.385 found α5β1 integrin receptor expression on metastatic cells was higher than that of the original cells from orthotopic tumors. They synthesized RPM-conjugated, α5β1-targeted micelles (RPM-CSOSA), which could enhance the cellular internalization and distribution in metastatic lesions by binding to α5β1 integrin. Doxorubicin (DOX) was a potent cytotoxic drug and curcumin (CUR) exerts its anti-cancer effects as chemosensitizer.386 Zhao et al.385 also found that the therapy of RPM-CSOSA/DOX and RPM-CSOSA/CUR significantly inhibited the progression of liver metastasis in vivo and in vitro.
Endothelin, a coreceptor of transforming growth factor-β, is preferentially expressed in solid tumor angiogenic endothelial cells.387 Serum endoglin is a useful marker for monitoring early signs of CRC metastasis.388 As an endoglin neutralizing antibody, TRC105 binds to human endoglin with high affinity and is associated with antibody-dependent cell-mediated cytotoxicity.389 Targeting endoglin with TRC105 strongly inhibits metastatic spread of breast cancer in vivo.390 Similarly, Targeting endoglin using TRC105 decreased metastatic spread of CRC cells to the mouse liver.391 In a phase I first-in-human study, TRC105 showed clinical efficacy on preexisting metastases in 2 patients.392 Collectively, these results demonstrated that in addition to endothelial cells, targeting endoglin to CAFs may be a potent method to prevent metastases formation, and emphasizes the potential of TRC105 for the treatment of metastatic tumors, not just a classic anti-vascular generate drugs.
The cytoskeleton of the cell and ECM determine the stiffness of the tissue.393,394 Highly activated metastasis-associated fibroblasts cause the ECM to stiffen, which in turn increase tissue stiffness. Metastasis stiffness influences the effect of anti-angiogenic therapy on intratumoral blood vessel reduction. Shen et al.395 reported that anti-hypertensive drugs targeting the renin-angiotensin system (anti-RAS) in combination with bevacizumab could significantly improve anti-angiogenic efficacy in CRLM. The mechanism is that anti-RAS inhibit fibroblast contraction and ECM deposition, thereby reducing the hardening of liver metastases and enhancing the anti-angiogenic effect of bevacizumab. This research highlights a new mechanism of action for anti-RAS drugs in cancer that could lead to new therapeutics.
The Cyclin-dependent kinase 8 (CDK8) is an early clinical stage drug that targets the overexpression in colon cancer.396 From Liang et al.397, inhibition of CDK8 almost has no effect on the growth of CRC cells and orthotopic transplanted tumors in subcutaneous, splenic, or cecum, but significantly inhibited the liver metastases of mouse and human colon cancer cells. CDK8 mediated CRLM due to downregulating of MMP inhibitor TIMP3 via TGFβ/SMAD-driven expression of a TIMP3-targeting microRNA, miR-181b, along with promotion of MMP3 in murine or MMP9 in human colon cancer cells via Wnt/β-catenin-driven transcription.397 This study illustrated that CDK8-targeting drugs can be used in the treatment of CRLM in the future.
We also would like to point out that the new development in cancer nanotechnology and photodynamic therapy (PDT) may provide potential solutions for CRC diagnosis and treatment.398 PDT has been exploited as a promising cancer treatment modality for many years. PDT can destroy cancer cells by generating reactive oxygen species (ROS) when photosensitizers are irradiated with light of suitable wavelength. PDT has the benefit of dual targeting by the drugs themselves and light, so its side-effects are much lower compared with Chemotherapy or radiation therapy. However, in traditional PDT, the light from the applied high-frequency source is scattered and cannot penetrate deep tissue, which limits its clinical application.399 Several possible solutions have been proposed to solve this problem, such as inventing new photosensitizers that use near infrared light and X-rays, and developing up-conversion nanoparticles.400,401,402 Among them, X-ray induced PDT has attracted much attention because of its unlimited tissue penetration ability.398,403,404,405,406 Most recently, Chen et al. reported Cu-Cy, Cu3Cl(SR)2 (R = CH2CH2NH2),407,408 a new type of sensitizer, has strong luminescence407,409 and can produce ROS under UV irradiation,407,410 X-rays,411,412,413 microwave radiation,414,415,416,417,418 and ultrasound.419 All these observations reported so far suggest that the Cu-Cy NPs are a new type of sensitizers with potential applications for infection inactivation420 and antitumor therapies.421 The most recent studies prove that this new Cu-Cy nanoparticle possess anti-tumoral effects and potential molecular mechanisms on B16 melanoma through X-PDT and X-ray induced antitumor immunity,422 and most importantly is that X-PDT is exceptionally effective with clinical X-rays used for irradiation in clinic settings.423 We believe that these new technologies could provide good solutions for CRC treatment.
Perspective
CRC is still a common cancer and liver metastasis is the main cause of death in CRC patients. The treatment of advanced CRLM remains a major challenge. Previous studies have determined the main steps of the metastasis process. The role of TME, the dysregulation of NcRNAs, and the activation of various signaling pathways are closely related to liver metastasis. However, due to the lack of adequate experimental models to detect this complex process fully and continuously, the molecular mechanisms involved in the formation of CRLM have still much room for exploration. Detailed molecular mechanisms that mediate CRC metastasis to liver contribute to early detection and prevention. Future work should involve clarifying CRLM, based on the molecular mechanisms and clinical characteristics, which guide clinical precision treatment. At the same time, combined treatment and multidisciplinary cooperation should be established. Intervention at the early stage of liver metastasis, such as the stage of metastasis and colonization, will be more beneficial to improve patient survival. In addition, clinical and experimental data strongly suggest that postoperative tumor recurrence remains a possibility, therefore adjuvant targeted therapy is necessary for patients with postoperative resection. Defining high-risk factors for CRLM, accurate selection of high-risk individuals and minimizing the controllable risk factors are essential for the prevention of CRLM and further reducing mortality of CRC. It is also possible that the new development in cancer nanotechnology like nanoparticle self-lighting PDT may be a new hope for CRC treatment.
References
Torre, L. A. et al. Global cancer incidence and mortality rates and trends–an update. Cancer Epidemiol. Biomark. Prev. 25, 16–27 (2016).
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 71, 209–249 (2021).
Parkin, D. M., Pisani, P. & Ferlay, J. Global cancer statistics. CA Cancer J. Clin. 49, 33–64 (1999). 1.
Parkin, D. M. et al. Global cancer statistics, 2002. CA Cancer J. Clin. 55, 74–108 (2005).
Jemal, A. et al. Global cancer statistics. CA Cancer J. Clin. 61, 69–90 (2011).
Torre, L. A. et al. Global cancer statistics, 2012. CA Cancer J. Clin. 65, 87–108 (2015).
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Yu, X. et al. Emerging Role of Immunotherapy for Colorectal Cancer with Liver Metastasis. Onco Targets Ther. 13, 11645–11658 (2020).
Tauriello, D. V. et al. Determinants of metastatic competency in colorectal cancer. Mol. Oncol. 11, 97–119 (2017).
de Ridder, J. et al. Incidence and origin of histologically confirmed liver metastases: an explorative case-study of 23,154 patients. Oncotarget 7, 55368–55376 (2016).
Hugen, N. et al. Metastatic pattern in colorectal cancer is strongly influenced by histological subtype. Ann. Oncol. 25, 651–657 (2014).
Valderrama-Treviño, A. I. et al. Hepatic Metastasis from Colorectal Cancer. Euroasian J. Hepatogastroenterol. 7, 166–175 (2017).
Akgül, Ö. et al. Role of surgery in colorectal cancer liver metastases. World J. Gastroenterol. 20, 6113–6122 (2014).
Al, B. M. & Kim, N. K. Current status and future perspectives on treatment of liver metastasis in colorectal cancer (Review). Oncol. Rep. 37, 2553–2564 (2017).
Takahashi, H. & Berber, E. Role of thermal ablation in the management of colorectal liver metastasis. Hepatobiliary Surg. Nutr. 9, 49–58 (2020).
Pan, Z. et al. Is there a survival benefit from adjuvant chemotherapy for patients with liver oligometastases from colorectal cancer after curative resection? Cancer Commun. 38, 29 (2018).
Norén, A., Eriksson, H. G. & Olsson, L. I. Selection for surgery and survival of synchronous colorectal liver metastases; a nationwide study. Eur. J. Cancer 53, 105–114 (2016).
Price, T. J. et al. Does the primary site of colorectal cancer impact outcomes for patients with metastatic disease? Cancer 121, 830–835 (2015).
Engstrand, J. et al. Colorectal cancer liver metastases—a population-based study on incidence, management and survival. BMC Cancer 18, 78 (2018).
Low, E. E. et al. Risk factors for early-onset colorectal cancer. Gastroenterology 159, 492–501.e7 (2020).
Siegel, R., Naishadham, D. & Jemal, A. Cancer statistics, 2012. CA Cancer J. Clin. 62, 10–29 (2012).
Riddiough, G. E. et al. Searching for the link; mechanisms underlying liver regeneration and recurrence of colorectal liver metastasis post partial hepatectomy. J. Gastroenterol. Hepatol. 34, 1276–1286 (2019).
Siegel, R. L. et al. Colorectal cancer statistics, 2020. CA Cancer J. Clin. 70, 145–164 (2020).
Margonis, G. A. et al. Association of BRAF Mutations With Survival and Recurrence in Surgically Treated Patients With Metastatic Colorectal Liver Cancer. JAMA Surg. 153, e180996 (2018).
Shi, R. et al. Prediction of KRAS, NRAS and BRAF status in colorectal cancer patients with liver metastasis using a deep artificial neural network based on radiomics and semantic features. Am. J. Cancer Res. 10, 4513–4526 (2020).
Tsilimigras, D. I. et al. Clinical significance and prognostic relevance of KRAS, BRAF, PI3K and TP53 genetic mutation analysis for resectable and unresectable colorectal liver metastases: a systematic review of the current evidence. Surg. Oncol. 27, 280–288 (2018).
Pitroda, S. P. et al. Integrated molecular subtyping defines a curable oligometastatic state in colorectal liver metastasis. Nat. Commun. 9, 1793 (2018).
Paget, S. The distribution of secondary growths in cancer of the breast. 1889. Cancer Metastasis Rev. 8, 98–101 (1989).
Jafri, M. A., Al-Qahtani, M. H. & Shay, J. W. Role of miRNAs in human cancer metastasis: implications for therapeutic intervention. Semin Cancer Biol. 44, 117–131 (2017).
Hui, L. & Chen, Y. Tumor microenvironment: Sanctuary of the devil. Cancer Lett. 368, 7–13 (2015).
Vitale, I. et al. Macrophages and metabolism in the tumor microenvironment. Cell Metab. 30, 36–50 (2019).
Yang, L. & Lin, P. C. Mechanisms that drive inflammatory tumor microenvironment, tumor heterogeneity, and metastatic progression. Semin Cancer Biol. 47, 185–195 (2017).
Mantovani, A. et al. Cancer-related inflammation. Nature 454, 436–444 (2008).
Mantovani, A. et al. Tumour-associated macrophages as treatment targets in oncology. Nat. Rev. Clin. Oncol. 14, 399–416 (2017).
Mantovani, A. et al. Macrophage polarization: tumor-associated macrophages as a paradigm for polarized M2 mononuclear phagocytes. Trends Immunol. 23, 549–555 (2002).
Murray, P. J. et al. Macrophage activation and polarization: nomenclature and experimental guidelines. Immunity 41, 14–20 (2014).
Grossman, J. G. et al. Recruitment of CCR2(+) tumor associated macrophage to sites of liver metastasis confers a poor prognosis in human colorectal cancer. Oncoimmunology 7, e1470729 (2018).
Tanaka, A. & Sakaguchi, S. Regulatory T cells in cancer immunotherapy. Cell Res. 27, 109–118 (2017).
Brudvik, K. W. et al. Regulatory T-cell-mediated inhibition of antitumor immune responses is associated with clinical outcome in patients with liver metastasis from colorectal cancer. Cancer Immunol. Immunother. 61, 1045–1053 (2012).
Wang, Q. et al. Intratumoral regulatory T cells are associated with suppression of colorectal carcinoma metastasis after resection through overcoming IL-17 producing T cells. Cell Immunol. 287, 100–105 (2014).
Tohme, S. et al. Neutrophil Extracellular Traps Promote the Development and Progression of Liver Metastases after Surgical Stress. Cancer Res. 76, 1367–1380 (2016).
Shen, Y. et al. A comprehensive look at the role of hyperlipidemia in promoting colorectal cancer liver metastasis. J. Cancer 9, 2981–2986 (2018).
Hirai, H. et al. CCR1-mediated accumulation of myeloid cells in the liver microenvironment promoting mouse colon cancer metastasis. Clin. Exp. Metastasis 31, 977–989 (2014).
Palmieri, V. et al. Neutrophils expressing lysyl oxidase-like 4 protein are present in colorectal cancer liver metastases resistant to anti-angiogenic therapy. J. Pathol. 251, 213–223 (2020).
Ogawa, R. et al. Loss of SMAD4 promotes colorectal cancer progression by recruiting tumor-associated neutrophils via the CXCL1/8-CXCR2 Axis. Clin. Cancer Res. 25, 2887–2899 (2019).
Yang, L. et al. IL-8 mediates a positive loop connecting increased neutrophil extracellular traps (NETs) and colorectal cancer liver metastasis. J. Cancer 11, 4384–4396 (2020).
Xia, Y. et al. AAV-mediated gene transfer of DNase I in the liver of mice with colorectal cancer reduces liver metastasis and restores local innate and adaptive immune response. Mol. Oncol. 14, 2920–2935 (2020).
Gabrilovich, D. I. Myeloid-derived suppressor cells. Cancer Immunol. Res. 5, 3–8 (2017).
Inamoto, S. et al. Loss of SMAD4 promotes colorectal cancer progression by accumulation of myeloid-derived suppressor cells through the CCL15-CCR1 chemokine axis. Clin. Cancer Res 22, 492–501 (2016).
Lin, Q. et al. The mechanism of the premetastatic niche facilitating colorectal cancer liver metastasis generated from myeloid-derived suppressor cells induced by the S1PR1-STAT3 signaling pathway. Cell Death Dis. 10, 693 (2019).
Väyrynen, J. P. et al. The prognostic role of macrophage polarization in the colorectal cancer microenvironment. Cancer Immunol. Res 9, 8–19 (2021).
Katz, S. C. et al. T cell infiltrate predicts long-term survival following resection of colorectal cancer liver metastases. Ann. Surg. Oncol. 16, 2524–2530 (2009).
Mlecnik, B. et al. Comprehensive Intrametastatic Immune Quantification and Major Impact of Immunoscore on Survival. J. Natl. Cancer Inst. 110, 97–108 (2018).
Tumeh, P. C. et al. Liver metastasis and treatment outcome with anti-PD-1 monoclonal antibody in patients with melanoma and NSCLC. Cancer Immunol. Res. 5, 417–424 (2017).
Donadon, M. et al. Increased infiltration of natural killer and T cells in colorectal liver metastases improves patient overall survival. J. Gastrointest. Surg. 21, 1226–1236 (2017).
Rokavec, M. et al. IL-6R/STAT3/miR-34a feedback loop promotes EMT-mediated colorectal cancer invasion and metastasis. J. Clin. Investig. 124, 1853–1867 (2014).
Ding, Y. et al. c-Fos separation from Lamin A/C by GDF15 promotes colon cancer invasion and metastasis in inflammatory microenvironment. J. Cell Physiol. 235, 4407–4421 (2020).
Zhang, Y. et al. IL-33 promotes growth and liver metastasis of colorectal cancer in mice by remodeling the tumor microenvironment and inducing angiogenesis. Mol. Carcinog. 56, 272–287 (2017).
Tutino, V. et al. Elevated AA/EPA ratio represents an inflammatory biomarker in tumor tissue of metastatic colorectal cancer patients. Int. J. Mol. Sci. 20, 2050 (2019).
Balkwill, F. R. The chemokine system and cancer. J. Pathol. 226, 148–157 (2012).
Mollica, P. V. et al. Chemokines and chemokine receptors: new targets for cancer immunotherapy. Front. Immunol. 10, 379 (2019).
Zhang, W. et al. CXCL5/CXCR2 axis in tumor microenvironment as potential diagnostic biomarker and therapeutic target. Cancer Commun. 40, 69–80 (2020).
Chen, C. et al. CXCL5 induces tumor angiogenesis via enhancing the expression of FOXD1 mediated by the AKT/NF-κB pathway in colorectal cancer. Cell Death Dis. 10, 178 (2019).
Zhao, J. et al. Tumor-derived CXCL5 promotes human colorectal cancer metastasis through activation of the ERK/Elk-1/Snail and AKT/GSK3β/β-catenin pathways. Mol. Cancer 16, 70 (2017).
Kitamura, T. et al. SMAD4-deficient intestinal tumors recruit CCR1+ myeloid cells that promote invasion. Nat. Genet 39, 467–475 (2007).
Kitamura, T. et al. Inactivation of chemokine (C-C motif) receptor 1 (CCR1) suppresses colon cancer liver metastasis by blocking accumulation of immature myeloid cells in a mouse model. Proc. Natl Acad. Sci. USA 107, 13063–13068 (2010).
Zhao, L. et al. Recruitment of a myeloid cell subset (CD11b/Gr1 mid) via CCL2/CCR2 promotes the development of colorectal cancer liver metastasis. Hepatology 57, 829–839 (2013).
Ichikawa, M. et al. S100A8/A9 activate key genes and pathways in colon tumor progression. Mol. Cancer Res. 9, 133–148 (2011).
Wang, D. et al. CXCL1 induced by prostaglandin E2 promotes angiogenesis in colorectal cancer. J. Exp. Med. 203, 941–951 (2006).
Zhang, Z. et al. GNA13 promotes tumor growth and angiogenesis by upregulating CXC chemokines via the NF-κB signaling pathway in colorectal cancer cells. Cancer Med. 7, 5611–5620 (2018).
Yamamoto, M. et al. TSU68 prevents liver metastasis of colon cancer xenografts by modulating the premetastatic niche. Cancer Res. 68, 9754–9762 (2008).
Wang, D. et al. CXCL1 is critical for premetastatic niche formation and metastasis in colorectal cancer. Cancer Res. 77, 3655–3665 (2017).
De la Fuente, L. M. et al. The relationship between chemokines CCL2, CCL3, and CCL4 with the tumor microenvironment and tumor-associated macrophage markers in colorectal cancer. Tumour Biol. 40, 1010428318810059 (2018).
Itatani, Y. et al. The role of chemokines in promoting colorectal cancer invasion/metastasis. Int. J. Mol. Sci. 17, 643 (2016).
Li, E. et al. CXCL8 associated dendritic cell activation marker expression and recruitment as indicators of favorable outcomes in colorectal cancer. Front Immunol. 12, 667177 (2021).
Zheng, J. et al. Chemokine receptor CX3CR1 contributes to macrophage survival in tumor metastasis. Mol. Cancer 12, 141 (2013).
Siddiqui, I. et al. Enhanced recruitment of genetically modified CX3CR1-positive human T cells into Fractalkine/CX3CL1 expressing tumors: importance of the chemokine gradient. J. Immunother. Cancer 4, 21 (2016).
Ma, L. et al. Modulating the interaction of CXCR4 and CXCL12 by low-molecular-weight heparin inhibits hepatic metastasis of colon cancer. Investig. N. Drugs 30, 508–517 (2012).
Schimanski, C. C. et al. Effect of chemokine receptors CXCR4 and CCR7 on the metastatic behavior of human colorectal cancer. Clin. Cancer Res 11, 1743–1750 (2005).
Wortzel, I. et al. Exosome-mediated metastasis: communication from a distance. Dev. Cell 49, 347–360 (2019).
Mathieu, M. et al. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 21, 9–17 (2019).
Kalluri, R., V. S. LeBleu. The biology, function, and biomedical applications of exosomes. Science. 367, 6777 (2020).
Valcz, G. et al. Exosomes in colorectal carcinoma formation: ALIX under the magnifying glass. Mod. Pathol. 29, 928–938 (2016).
Peinado, H. et al. Pre-metastatic niches: organ-specific homes for metastases. Nat. Rev. Cancer 17, 302–317 (2017).
Dhanabal, M. et al. Angioarrestin: an antiangiogenic protein with tumor-inhibiting properties. Cancer Res. 62, 3834–3841 (2002).
Jiang, K. et al. Exosomal ANGPTL1 attenuates colorectal cancer liver metastasis by regulating Kupffer cell secretion pattern and impeding MMP9 induced vascular leakiness. J. Exp. Clin. Cancer Res. 40, 21 (2021).
Mashouri, L. et al. Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol. Cancer 18, 75 (2019).
Kalluri, R. The biology and function of exosomes in cancer. J. Clin. Investig. 126, 1208–1215 (2016).
Zeng, Z. et al. Cancer-derived exosomal miR-25-3p promotes pre-metastatic niche formation by inducing vascular permeability and angiogenesis. Nat. Commun. 9, 5395 (2018).
Shao, Y. et al. Colorectal cancer-derived small extracellular vesicles establish an inflammatory premetastatic niche in liver metastasis. Carcinogenesis 39, 1368–1379 (2018).
Liu, H. et al. Colorectal cancer-derived exosomal miR-106b-3p promotes metastasis by down-regulating DLC-1 expression. Clin. Sci. 134, 419–434 (2020).
Bigagli, E. et al. Exosomes secreted from human colon cancer cells influence the adhesion of neighboring metastatic cells: Role of microRNA-210. Cancer Biol. Ther. 17, 1062–1069 (2016).
Matsumura, T. et al. Exosomal microRNA in serum is a novel biomarker of recurrence in human colorectal cancer. Br. J. Cancer 113, 275–281 (2015).
Rezaei, R. et al. Exosome-mediated delivery of functionally active miRNA-375-3p mimic regulate epithelial mesenchymal transition (EMT) of colon cancer cells. Life Sci. 269, 119035 (2021).
Fu, F. et al. Circulating Exosomal miR-17-5p and miR-92a-3p Predict Pathologic Stage and Grade of Colorectal Cancer. Transl. Oncol. 11, 221–232 (2018).
Guo, S. et al. Exosomes derived from Fusobacterium nucleatum-infected colorectal cancer cells facilitate tumour metastasis by selectively carrying miR-1246/92b-3p/27a-3p and CXCL16. Gut 71, e1–3 (2020).
Zhang, X. et al. Exosomal miR-1255b-5p targets human telomerase reverse transcriptase in colorectal cancer cells to suppress epithelial-to-mesenchymal transition. Mol. Oncol. 14, 2589–2608 (2020).
Hu, J. L. et al. CAFs secreted exosomes promote metastasis and chemotherapy resistance by enhancing cell stemness and epithelial-mesenchymal transition in colorectal cancer. Mol. Cancer 18, 91 (2019).
Wu, J. et al. The malignant role of exosomes in the communication among colorectal cancer cell, macrophage and microbiome. Carcinogenesis 40, 601–610 (2019).
Rai, A. et al. Exosomes derived from human primary and metastatic colorectal cancer cells contribute to functional heterogeneity of activated fibroblasts by reprogramming their proteome. Proteomics 19, e1800148 (2019).
Li, H. & Li, F. Exosomes from BM-MSCs increase the population of CSCs via transfer of miR-142-3p. Br. J. Cancer 119, 744–755 (2018).
Teng, Y. et al. MVP-mediated exosomal sorting of miR-193a promotes colon cancer progression. Nat. Commun. 8, 14448 (2017).
Santasusagna, S. et al. Prognostic impact of miR-200 family members in plasma and exosomes from tumor-draining versus peripheral veins of colon cancer patients. Oncology 95, 309–318 (2018).
Liu, W. et al. Plasma exosomal miRNA-139-3p is a novel biomarker of colorectal cancer. J. Cancer 11, 4899–4906 (2020).
Sun, L. et al. Serum exosomal miR-122 as a potential diagnostic and prognostic biomarker of colorectal cancer with liver metastasis. J. Cancer 11, 630–637 (2020).
Wang, D. et al. Exosome-encapsulated miRNAs contribute to CXCL12/CXCR4-induced liver metastasis of colorectal cancer by enhancing M2 polarization of macrophages. Cancer Lett. 474, 36–52 (2020).
Zhao, S. et al. Tumor-derived exosomal miR-934 induces macrophage M2 polarization to promote liver metastasis of colorectal cancer. J. Hematol. Oncol. 13, 156 (2020).
Cho, W. C. et al. Exosomal miR-193a and let-7g accelerate cancer progression on primary colorectal cancer and paired peritoneal metastatic cancer. Transl. Oncol. 14, 101000 (2021).
Pan, L. et al. Exosomes-mediated transfer of long noncoding RNA ZFAS1 promotes gastric cancer progression. J. Cancer Res. Clin. Oncol. 143, 991–1004 (2017).
Dong, L. et al. Circulating long RNAs in serum extracellular vesicles: their characterization and potential application as biomarkers for diagnosis of colorectal cancer. Cancer Epidemiol. Biomark. Prev. 25, 1158–1166 (2016).
Xu, J. et al. Exosomal MALAT1 sponges miR-26a/26b to promote the invasion and metastasis of colorectal cancer via FUT4 enhanced fucosylation and PI3K/Akt pathway. J. Exp. Clin. Cancer Res. 39, 54 (2020).
Yang, H. et al. Hypoxia induced exosomal circRNA promotes metastasis of Colorectal Cancer via targeting GEF-H1/RhoA axis. Theranostics 10, 8211–8226 (2020).
Yin, Y. et al. Colorectal Cancer-Derived Small Extracellular Vesicles Promote Tumor Immune Evasion by Upregulating PD-L1 Expression in Tumor-Associated Macrophages. Adv. Sci. 17, e2102620 (2022).
Lan, J. et al. M2 macrophage-derived exosomes promote cell migration and invasion in colon cancer. Cancer Res. 79, 146–158 (2019).
Saha, S. et al. Macrophage-derived extracellular vesicle-packaged WNTs rescue intestinal stem cells and enhance survival after radiation injury. Nat. Commun. 7, 13096 (2016).
Cooks, T. et al. Mutant p53 cancers reprogram macrophages to tumor supporting macrophages via exosomal miR-1246. Nat. Commun. 9, 771 (2018).
Clayton, A. et al. Human tumor-derived exosomes selectively impair lymphocyte responses to interleukin-2. Cancer Res. 67, 7458–7466 (2007).
Sun, B. et al. Colorectal cancer exosomes induce lymphatic network remodeling in lymph nodes. Int J. Cancer 145, 1648–1659 (2019).
Yamada, N. et al. Colorectal cancer cell-derived extracellular vesicles induce phenotypic alteration of T cells into tumor-growth supporting cells with transforming growth factor-β1-mediated suppression. Oncotarget 7, 27033–27043 (2016).
Huber, V. et al. Human colorectal cancer cells induce T-cell death through release of proapoptotic microvesicles: role in immune escape. Gastroenterology 128, 1796–1804 (2005).
Yuzhalin, A. E. et al. Colorectal cancer liver metastatic growth depends on PAD4-driven citrullination of the extracellular matrix. Nat. Commun. 9, 4783 (2018).
Shang, A. et al. Exosomal KRAS mutation promotes the formation of tumor-associated neutrophil extracellular traps and causes deterioration of colorectal cancer by inducing IL-8 expression. Cell Commun. Signal 18, 52 (2020).
Tacchini, L. et al. Hepatocyte growth factor signaling regulates transactivation of genes belonging to the plasminogen activation system via hypoxia inducible factor-1. Exp. Cell Res. 290, 391–401 (2003).
Gherardi, E. et al. Targeting MET in cancer: rationale and progress. Nat. Rev. Cancer 12, 89–103 (2012).
Demkova, L. & Kucerova, L. Role of the HGF/c-MET tyrosine kinase inhibitors in metastasic melanoma. Mol. Cancer 17, 26 (2018).
Trusolino, L. & Comoglio, P. M. Scatter-factor and semaphorin receptors: cell signalling for invasive growth. Nat. Rev. Cancer 2, 289–300 (2002).
Organ, S. L. et al. Quantitative phospho-proteomic profiling of hepatocyte growth factor (HGF)-MET signaling in colorectal cancer. J. Proteome Res. 10, 3200–3211 (2011).
Yao, J. F. et al. Role of HGF/c-Met in the treatment of colorectal cancer with liver metastasis. J. Biochem Mol. Toxicol. 33, e22316 (2019).
Bauer, T. W. et al. Insulinlike growth factor-I-mediated migration and invasion of human colon carcinoma cells requires activation of c-Met and urokinase plasminogen activator receptor. Ann. Surg. 241, 748–756 (2005).
Kopitz, C. et al. Tissue inhibitor of metalloproteinases-1 promotes liver metastasis by induction of hepatocyte growth factor signaling. Cancer Res. 67, 8615–8623 (2007).
Schrötzlmair, F. et al. Tissue inhibitor of metalloproteinases-1-induced scattered liver metastasis is mediated by host-derived urokinase-type plasminogen activator. J. Cell Mol. Med. 14, 2760–2770 (2010).
Schelter, F. et al. Tissue inhibitor of metalloproteinases-1-induced scattered liver metastasis is mediated by hypoxia-inducible factor-1α. Clin. Exp. Metastasis 28, 91–99 (2011).
Du, F. et al. SOX13 promotes colorectal cancer metastasis by transactivating SNAI2 and c-MET. Oncogene 39, 3522–3540 (2020).
Cui, Y. M. et al. FOXC2 promotes colorectal cancer metastasis by directly targeting MET. Oncogene 34, 4379–4390 (2015).
Pai, R. et al. Prostaglandins promote colon cancer cell invasion; signaling by cross-talk between two distinct growth factor receptors. FASEB J. 17, 1640–1647 (2003).
Moore, A. E. et al. HGF/Met signalling promotes PGE(2) biogenesis via regulation of COX-2 and 15-PGDH expression in colorectal cancer cells. Carcinogenesis 30, 1796–1804 (2009).
Liao, H. et al. Thermal Ablation Induces Transitory Metastatic Growth by Means of the STAT3/c-Met Molecular Pathway in an Intrahepatic Colorectal Cancer Mouse Model. Radiology 294, 464–472 (2020).
Stein, U. et al. MACC1, a newly identified key regulator of HGF-MET signaling, predicts colon cancer metastasis. Nat. Med. 15, 59–67 (2009).
Isella, C. et al. MACC1 mRNA levels predict cancer recurrence after resection of colorectal cancer liver metastases. Ann. Surg. 257, 1089–1095 (2013).
Leiphrakpam, P. D. et al. Correlation of PRL3 expression with colorectal cancer progression. J. Surg. Oncol. 123, 42–51 (2021).
Saha, S. et al. A phosphatase associated with metastasis of colorectal cancer. Science 294, 1343–1346 (2001).
Lee, S. K. et al. Phosphatase of regenerating liver-3 promotes migration and invasion by upregulating matrix metalloproteinases-7 in human colorectal cancer cells. Int. J. Cancer 131, E190–E203 (2012).
Al-Aidaroos, A. Q. et al. Metastasis-associated PRL-3 induces EGFR activation and addiction in cancer cells. J. Clin. Investig. 123, 3459–3471 (2013).
Guo, K. et al. PRL-3 initiates tumor angiogenesis by recruiting endothelial cells in vitro and in vivo. Cancer Res. 66, 9625–9635 (2006).
Kim, N. W. et al. Correlation between liver metastases and the level of PRL-3 mRNA expression in patients with primary colorectal cancer. J. Korean Soc. Coloproctol. 27, 231–236 (2011).
Hatate, K. et al. Liver metastasis of colorectal cancer by protein-tyrosine phosphatase type 4A, 3 (PRL-3) is mediated through lymph node metastasis and elevated serum tumor markers such as CEA and CA19-9. Oncol. Rep. 20, 737–743 (2008).
Zhang, J. et al. miR-21, miR-17 and miR-19a induced by phosphatase of regenerating liver-3 promote the proliferation and metastasis of colon cancer. Br. J. Cancer 107, 352–359 (2012).
Xing, C. et al. Ubiquitin-specific protease 4-mediated deubiquitination and stabilization of PRL-3 is required for potentiating colorectal oncogenesis. Cancer Res. 76, 83–95 (2016).
Xu, H. et al. PRL-3 improves colorectal cancer cell proliferation and invasion through IL-8 mediated glycolysis metabolism. Int. J. Oncol. 51, 1271–1279 (2017).
Lan, Q. et al. CCL26 participates in the PRL-3-induced promotion of colorectal cancer invasion by stimulating tumor-associated macrophage infiltration. Mol. Cancer Ther. 17, 276–289 (2018).
Zhang, T. et al. Interaction with tumor‑associated macrophages promotes PRL‑3‑induced invasion of colorectal cancer cells via MAPK pathway‑induced EMT and NF‑κB signaling‑induced angiogenesis. Oncol. Rep. 41, 2790–2802 (2019).
Casey, L. M. et al. Jag2-Notch1 signaling regulates oral epithelial differentiation and palate development. Dev. Dyn. 235, 1830–1844 (2006).
Arcaroli, J. J. et al. A NOTCH1 gene copy number gain is a prognostic indicator of worse survival and a predictive biomarker to a Notch1 targeting antibody in colorectal cancer. Int. J. Cancer 138, 195–205 (2016).
Rallis, G. et al. Association of Notch and Hedgehog Pathway Activation With Prognosis in Early-stage Colorectal Cancer. Anticancer Res. 39, 2129–2138 (2019).
Wang, W. J. et al. Increased LEF1 expression and decreased Notch2 expression are strong predictors of poor outcomes in colorectal cancer patients. Dis. Markers 35, 395–405 (2013).
Zhang, Z. et al. NOTCH4 regulates colorectal cancer proliferation, invasiveness, and determines clinical outcome of patients. J. Cell Physiol. 233, 6975–6985 (2018).
Fender, A. W. et al. Notch-1 promotes stemness and epithelial to mesenchymal transition in colorectal cancer. J. Cell Biochem. 116, 2517–2527 (2015).
Jackstadt, R. et al. Epithelial NOTCH signaling rewires the tumor microenvironment of colorectal cancer to drive poor-prognosis subtypes and metastasis. Cancer Cell 36, 319–336.e7 (2019).
Zavadil, J. et al. Integration of TGF-beta/Smad and Jagged1/Notch signalling in epithelial-to-mesenchymal transition. EMBO J. 23, 1155–1165 (2004).
Song, M. et al. MiR-139-5p inhibits migration and invasion of colorectal cancer by downregulating AMFR and NOTCH1. Protein Cell 5, 851–861 (2014).
Dai, Y. et al. Silencing of Jagged1 inhibits cell growth and invasion in colorectal cancer. Cell Death Dis. 5, e1170 (2014).
Pastò, A. et al. NOTCH3 signaling regulates MUSASHI-1 expression in metastatic colorectal cancer cells. Cancer Res. 74, 2106–2118 (2014).
Sonoshita, M. et al. Suppression of colon cancer metastasis by Aes through inhibition of Notch signaling. Cancer Cell 19, 125–137 (2011).
Wang, Z. et al. The CK1δ/ε-AES axis regulates tumorigenesis and metastasis in colorectal cancer. Theranostics 11, 4421–4435 (2021).
Huang, B. et al. tRF/miR-1280 suppresses stem cell-like cells and metastasis in colorectal cancer. Cancer Res. 77, 3194–3206 (2017).
Zheng, C. G. et al. Immunohistochemical expression of Notch1, Jagged1, NF-κB and MMP-9 in colorectal cancer patients and the relationship to clinicopathological parameters. Cancer Biomark. 15, 889–897 (2015).
Gordon-Weeks, A. et al. Tumour-derived laminin α5 (LAMA5) promotes colorectal liver metastasis growth, branching angiogenesis and notch pathway inhibition. Cancers. 11, 630 (2019).
Weiss, A. & Attisano, L. The TGFbeta superfamily signaling pathway. Wiley Interdiscip. Rev. Dev. Biol. 2, 47–63 (2013).
Tauriello, D. et al. TGFβ drives immune evasion in genetically reconstituted colon cancer metastasis. Nature 554, 538–543 (2018).
Ji, Q. et al. Resveratrol suppresses epithelial-to-mesenchymal transition in colorectal cancer through TGF-β1/Smads signaling pathway mediated Snail/E-cadherin expression. BMC Cancer 15, 97 (2015).
Tan, H. X. et al. CXCR4/TGF-β1 mediated hepatic stellate cells differentiation into carcinoma-associated fibroblasts and promoted liver metastasis of colon cancer. Cancer Biol. Ther. 21, 258–268 (2020).
Roberts, A. B. & Wakefield, L. M. The two faces of transforming growth factor beta in carcinogenesis. Proc. Natl Acad. Sci. USA 100, 8621–8623 (2003).
van den Bulk, J. et al. Neoantigen-specific immunity in low mutation burden colorectal cancers of the consensus molecular subtype 4. Genome Med. 11, 87 (2019).
Massagué, J. TGFβ signalling in context. Nat. Rev. Mol. Cell Biol. 13, 616–630 (2012).
Calon, A. et al. Dependency of colorectal cancer on a TGF-β-driven program in stromal cells for metastasis initiation. Cancer Cell 22, 571–584 (2012).
Losi, L., Luppi, G. & Benhattar, J. Assessment of K-ras, Smad4 and p53 gene alterations in colorectal metastases and their role in the metastatic process. Oncol. Rep. 12, 1221–1225 (2004).
Zhao, S. et al. Inhibition of STAT3 Tyr705 phosphorylation by Smad4 suppresses transforming growth factor beta-mediated invasion and metastasis in pancreatic cancer cells. Cancer Res 68, 4221–4228 (2008).
Xiao, B. et al. Oncolytic Adenovirus CD55-Smad4 Suppresses Cell Proliferation, Metastasis, and Tumor Stemness in Colorectal Cancer by Regulating Wnt/β-Catenin Signaling Pathway. Biomedicines. 8 (2020).
Wu, N. et al. LINC00941 promotes CRC metastasis through preventing SMAD4 protein degradation and activating the TGF-β/SMAD2/3 signaling pathway. Cell Death Differ. 28, 219–232 (2021).
Zhang, L. et al. Qingjie Fuzheng Granule Inhibited the Migration and Invasion of Colorectal Cancer Cells by Regulating the lncRNA ANRIL/let-7a/TGF-β1/Smad Axis. Evid. Based Complement Altern. Med 2020, 5264651 (2020).
Altevogt, P., Doberstein, K. & Fogel, M. L1CAM in human cancer. Int J. Cancer 138, 1565–1576 (2016).
Min, J. K. et al. L1 cell adhesion molecule is a novel therapeutic target in intrahepatic cholangiocarcinoma. Clin. Cancer Res. 16, 3571–3580 (2010).
Ganesh, K. et al. L1CAM defines the regenerative origin of metastasis-initiating cells in colorectal cancer. Nat. Cancer 1, 28–45 (2020).
Ericson, K. et al. Genetic inactivation of AKT1, AKT2, and PDPK1 in human colorectal cancer cells clarifies their roles in tumor growth regulation. Proc. Natl Acad. Sci. USA 107, 2598–2603 (2010).
Irie, H. Y. et al. Distinct roles of Akt1 and Akt2 in regulating cell migration and epithelial-mesenchymal transition. J. Cell Biol. 171, 1023–1034 (2005).
Rychahou, P. G. et al. Akt2 overexpression plays a critical role in the establishment of colorectal cancer metastasis. Proc. Natl Acad. Sci. USA 105, 20315–20320 (2008).
Agarwal, E. et al. Role of Akt2 in regulation of metastasis suppressor 1 expression and colorectal cancer metastasis. Oncogene 36, 3104–3118 (2017).
Chen, Z. H. et al. Eukaryotic initiation factor 4A2 promotes experimental metastasis and oxaliplatin resistance in colorectal cancer. J. Exp. Clin. Cancer Res. 38, 196 (2019).
Wang, J. et al. Identification of Trop-2 as an oncogene and an attractive therapeutic target in colon cancers. Mol. Cancer Ther. 7, 280–285 (2008).
Shoji, Y. et al. The Embryonic Stem Cell-Specific Transcription Factor ZFP57 Promotes Liver Metastasis of Colorectal Cancer. J. Surg. Res. 237, 22–29 (2019).
Tang, B. et al. PEA15 promotes liver metastasis of colorectal cancer by upregulating the ERK/MAPK signaling pathway. Oncol. Rep. 41, 43–56 (2019).
Hu, H. et al. MEGF6 Promotes the Epithelial-to-Mesenchymal Transition via the TGFβ/SMAD Signaling Pathway in Colorectal Cancer Metastasis. Cell Physiol. Biochem. 46, 1895–1906 (2018).
Zhao, Z. et al. Apolipoprotein E Overexpression Is Associated With Tumor Progression and Poor Survival in Colorectal Cancer. Front. Genet 9, 650 (2018).
Song, W. et al. Trapping of Lipopolysaccharide to Promote Immunotherapy against Colorectal Cancer and Attenuate Liver Metastasis. Adv. Mater. 30, e1805007 (2018).
Komatsu, K. et al. Increased expression of S100A6 at the invading fronts of the primary lesion and liver metastasis in patients with colorectal adenocarcinoma. Br. J. Cancer 83, 769–774 (2000).
Komatsu, K. et al. Expression of S100A6 and S100A4 in matched samples of human colorectal mucosa, primary colorectal adenocarcinomas and liver metastases. Oncology 63, 192–200 (2002).
Dahlmann, M. et al. Systemic shRNA mediated knock down of S100A4 in colorectal cancer xenografted mice reduces metastasis formation. Oncotarget 3, 783–797 (2012).
Seubert, B. et al. Tissue inhibitor of metalloproteinases (TIMP)-1 creates a premetastatic niche in the liver through SDF-1/CXCR4-dependent neutrophil recruitment in mice. Hepatology 61, 238–248 (2015).
Zhu, X. et al. FBX8 promotes metastatic dormancy of colorectal cancer in liver. Cell Death Dis. 11, 622 (2020).
Liu, X. et al. THBS1 facilitates colorectal liver metastasis through enhancing epithelial-mesenchymal transition. Clin. Transl. Oncol. 22, 1730–1740 (2020).
Picardo, F. et al. Diagnostic and Prognostic Value of B4GALT1 Hypermethylation and Its Clinical Significance as a Novel Circulating Cell-Free DNA Biomarker in Colorectal Cancer. Cancers 11, 1598 (2019).
Li, Z. et al. Ferritin Light Chain (FTL) competes with long noncoding RNA Linc00467 for miR-133b binding site to regulate chemoresistance and metastasis of colorectal cancer. Carcinogenesis 41, 467–477 (2020).
Wu, Q. et al. DEFB4A is a potential prognostic biomarker for colorectal cancer. Oncol. Lett. 20, 114 (2020).
Zhou, L. et al. The β-galactoside α2,6-sialyltranferase 1 (ST6GAL1) inhibits the colorectal cancer metastasis by stabilizing intercellular adhesion molecule-1 via sialylation. Cancer Manag Res. 11, 6185–6199 (2019).
Yang, Y. et al. Prognostic value of a hypoxia-related microRNA signature in patients with colorectal cancer. Aging12, 35–52 (2020).
Řehulka, J. et al. Peloruside A-Induced Cell Death in Hypoxia Is p53 Dependent in HCT116 Colorectal Cancer Cells. J. Nat. Prod. 81, 634–640 (2018).
Li, H. et al. Antagonistic Effects of p53 and HIF1A on microRNA-34a Regulation of PPP1R11 and STAT3 and Hypoxia-induced Epithelial to Mesenchymal Transition in Colorectal Cancer Cells. Gastroenterology 153, 505–520 (2017).
Chen, H. J. et al. Dual inhibition of EGFR and c-Met kinase activation by MJ-56 reduces metastasis of HT29 human colorectal cancer cells. Int J. Oncol. 43, 141–150 (2013).
Chu, P. C. et al. Mutant KRAS promotes liver metastasis of colorectal cancer, in part, by upregulating the MEK-Sp1-DNMT1-miR-137-YB-1-IGF-IR signaling pathway. Oncogene 37, 3440–3455 (2018).
Li, Y. D. et al. Retinoblastoma binding protein 4 up-regulation is correlated with hepatic metastasis and poor prognosis in colon cancer patients. Hepatobiliary Pancreat. Dis. Int 18, 446–451 (2019).
Zhang, C. et al. Long non-coding RNA Lnc-LALC facilitates colorectal cancer liver metastasis via epigenetically silencing LZTS1. Cell Death Dis. 12, 224 (2021).
Yang, X. et al. METTL14 suppresses proliferation and metastasis of colorectal cancer by down-regulating oncogenic long non-coding RNA XIST. Mol. Cancer 19, 46 (2020).
Liu, J. et al. Long noncoding RNA LINC01578 drives colon cancer metastasis through a positive feedback loop with the NF-κB/YY1 axis. Mol. Oncol. 14, 3211–3233 (2020).
Jin, L. et al. LncRNA HOTAIR recruits SNAIL to inhibit the transcription of HNF4α and promote the viability, migration, invasion and EMT of colorectal cancer. Transl. Oncol. 14, 101036 (2021).
Qi, Z. P. et al. HDAC2 promotes the EMT of colorectal cancer cells and via the modular scaffold function of ENSG00000274093.1. J. Cell Mol. Med. 25, 1190–1197 (2021).
Zhang, Y. et al. Long non-coding RNA TPT1-AS1 promotes angiogenesis and metastasis of colorectal cancer through TPT1-AS1/NF90/VEGFA signaling pathway. Aging 12, 6191–6205 (2020).
Ye, L. C. et al. Aberrant expression of long noncoding RNAs in colorectal cancer with liver metastasis. Tumour Biol. 36, 8747–8754 (2015).
Huang, L. et al. Aberrant expression of long noncoding RNA SNHG15 correlates with liver metastasis and poor survival in colorectal cancer. J. Cell Physiol. 234, 7032–7039 (2019).
Xu, M. et al. LncRNA SATB2-AS1 inhibits tumor metastasis and affects the tumor immune cell microenvironment in colorectal cancer by regulating SATB2. Mol. Cancer 18, 135 (2019).
Xu, J. et al. MIR22HG acts as a tumor suppressor via TGFβ/SMAD signaling and facilitates immunotherapy in colorectal cancer. Mol. Cancer 19, 51 (2020).
Chen, D. L. et al. Long non-coding RNA UICLM promotes colorectal cancer liver metastasis by acting as a ceRNA for microRNA-215 to regulate ZEB2 expression. Theranostics 7, 4836–4849 (2017).
Zhu, X. et al. Long noncoding RNA RP11-757G1.5 sponges miR-139-5p and upregulates YAP1 thereby promoting the proliferation and liver, spleen metastasis of colorectal cancer. J. Exp. Clin. Cancer Res. 39, 207 (2020).
Wang, L. et al. Long Noncoding RNA LINC00460 Facilitates Colorectal Cancer Progression by Negatively Regulating miR-613. Onco Targets Ther. 13, 7555–7569 (2020).
Dong, X. et al. Long Non-coding RNA MIR4435-2HG Promotes Colorectal Cancer Proliferation and Metastasis Through miR-206/YAP1 Axis. Front. Oncol. 10, 160 (2020).
Jia, G. Q. et al. Long non-coding RNA PlncRNA-1 promotes cell proliferation and hepatic metastasis in colorectal cancer. J. Cell Biochem. 119, 7091–7104 (2018).
Wang, Y. et al. Long noncoding RNA DANCR promotes colorectal cancer proliferation and metastasis via miR-577 sponging. Exp. Mol. Med. 50, 1–17 (2018).
Cai, Y. et al. Long non-coding RNA TP73-AS1 sponges miR-194 to promote colorectal cancer cell proliferation, migration and invasion via up-regulating TGFα. Cancer Biomark. 23, 145–156 (2018).
Liu, B. et al. The HOTAIR/miR-214/ST6GAL1 crosstalk modulates colorectal cancer procession through mediating sialylated c-Met via JAK2/STAT3 cascade. J. Exp. Clin. Cancer Res. 38, 455 (2019).
Luo, Y. et al. Long Noncoding RNA GAPLINC Promotes Cells Migration and Invasion in Colorectal Cancer Cell by Regulating miR-34a/c-MET Signal Pathway. Dig. Dis. Sci. 63, 890–899 (2018).
Ni, X. et al. Long non-coding RNA ZEB1-AS1 promotes colon adenocarcinoma malignant progression via miR-455-3p/PAK2 axis. Cell Prolif. 53, e12723 (2020).
Zhou, T. et al. SOX9-activated FARSA-AS1 predetermines cell growth, stemness, and metastasis in colorectal cancer through upregulating FARSA and SOX9. Cell Death Dis. 11, 1071 (2020).
Zhang, W. et al. Lnc-HSD17B11-1:1 Functions as a Competing Endogenous RNA to Promote Colorectal Cancer Progression by Sponging miR-338-3p to Upregulate MACC1. Front. Genet 11, 628 (2020).
Wang, L. et al. Long noncoding RNA B3GALT5-AS1 suppresses colon cancer liver metastasis via repressing microRNA-203. Aging 10, 3662–3682 (2018).
Ding, Y. et al. Serum level of long noncoding RNA B3GALT5-AS1 as a diagnostic biomarker of colorectal cancer. Future Oncol. 16, 827–835 (2020).
Zheng, X. et al. A novel protein encoded by a circular RNA circPPP1R12A promotes tumor pathogenesis and metastasis of colon cancer via Hippo-YAP signaling. Mol. Cancer 18, 47 (2019).
Chen, R. X. et al. N(6)-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat. Commun. 10, 4695 (2019).
Tang, J. et al. CircRNA circ_0124554 blocked the ubiquitination of AKT promoting the skip lymphovascular invasion on hepatic metastasis in colorectal cancer. Cell Death Dis. 12, 270 (2021).
Xu, X. W. et al. Circular RNA hsa_circ_000984 promotes colon cancer growth and metastasis by sponging miR-106b. Oncotarget 8, 91674–91683 (2017).
Yong, W. et al. Hsa_circ_0071589 promotes carcinogenesis via the miR-600/EZH2 axis in colorectal cancer. Biomed. Pharmacother. 102, 1188–1194 (2018).
Wu, H. B. et al. CircAPLP2 regulates the proliferation and metastasis of colorectal cancer by targeting miR-101-3p to activate the Notch signalling pathway. Am. J. Transl. Res. 12, 2554–2569 (2020).
Chen, L. Y. et al. NSD2 circular RNA promotes metastasis of colorectal cancer by targeting miR-199b-5p-mediated DDR1 and JAG1 signalling. J. Pathol. 248, 103–115 (2019).
Hsiao, K. Y. et al. Noncoding Effects of Circular RNA CCDC66 Promote Colon Cancer Growth and Metastasis. Cancer Res. 77, 2339–2350 (2017).
Zhi, Q. et al. Circular RNA profiling identifies circ102049 as a key regulator of colorectal liver metastasis. Mol. Oncol. 15, 623–641 (2021).
Ren, C. et al. Circular RNA hsa_circ_0001178 facilitates the invasion and metastasis of colorectal cancer through upregulating ZEB1 via sponging multiple miRNAs. Biol. Chem. 401, 487–496 (2020).
Geng, Y. et al. Hsa_circ_0009361 acts as the sponge of miR-582 to suppress colorectal cancer progression by regulating APC2 expression. Clin. Sci. 133, 1197–1213 (2019).
Zeng, W. et al. CircFNDC3B sequestrates miR-937-5p to derepress TIMP3 and inhibit colorectal cancer progression. Mol. Oncol. 14, 2960–2984 (2020).
Li, X. et al. Circular RNA circITGA7 inhibits colorectal cancer growth and metastasis by modulating the Ras pathway and upregulating transcription of its host gene ITGA7. J. Pathol. 246, 166–179 (2018).
Xu, H. et al. RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers. Mol. Cancer 18, 8 (2019).
Shi, L. et al. Hypoxia-induced hsa_circ_0000826 is linked to liver metastasis of colorectal cancer. J. Clin. Lab. Anal. 34, e23405 (2020).
Paterson, E. L. et al. Down-regulation of the miRNA-200 family at the invasive front of colorectal cancers with degraded basement membrane indicates EMT is involved in cancer progression. Neoplasia 15, 180–191 (2013).
Hur, K. et al. MicroRNA-200c modulates epithelial-to-mesenchymal transition (EMT) in human colorectal cancer metastasis. Gut 62, 1315–1326 (2013).
Park, S. M. et al. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. 22, 894–907 (2008).
Li, J. et al. MiR-429 is an independent prognostic factor in colorectal cancer and exerts its anti-apoptotic function by targeting SOX2. Cancer Lett. 329, 84–90 (2013).
Ji, D. et al. MicroRNA-181a promotes tumor growth and liver metastasis in colorectal cancer by targeting the tumor suppressor WIF-1. Mol. Cancer 13, 86 (2014).
Zhao, H. et al. miR-30b regulates migration and invasion of human colorectal cancer via SIX1. Biochem J. 460, 117–125 (2014).
Park, Y. R. et al. MicroRNA-30a-5p (miR-30a) regulates cell motility and EMT by directly targeting oncogenic TM4SF1 in colorectal cancer. J. Cancer Res. Clin. Oncol. 143, 1915–1927 (2017).
Meng, X. et al. Genetic and epigenetic down-regulation of microRNA-212 promotes colorectal tumor metastasis via dysregulation of MnSOD. Gastroenterology 145, 426–36.e1-6 (2013).
Gu, J. et al. SATB2 targeted by methylated miR-34c-5p suppresses proliferation and metastasis attenuating the epithelial-mesenchymal transition in colorectal cancer. Cell Prolif. 51, e12455 (2018).
Li, J. et al. MiR-186-5p upregulation inhibits proliferation, metastasis and epithelial-to-mesenchymal transition of colorectal cancer cell by targeting ZEB1. Arch. Biochem. Biophys. 640, 53–60 (2018).
Kim, T. W. et al. MicroRNA-17-5p regulates EMT by targeting vimentin in colorectal cancer. Br. J. Cancer 123, 1123–1130 (2020).
Jackstadt, R. et al. AP4 is a mediator of epithelial-mesenchymal transition and metastasis in colorectal cancer. J. Exp. Med. 210, 1331–1350 (2013).
Shi, L. et al. p53-induced miR-15a/16-1 and AP4 form a double-negative feedback loop to regulate epithelial-mesenchymal transition and metastasis in colorectal cancer. Cancer Res. 74, 532–542 (2014).
Shell, S. et al. Let-7 expression defines two differentiation stages of cancer. Proc. Natl Acad. Sci. USA 104, 11400–11405 (2007).
King, C. E. et al. LIN28B promotes colon cancer progression and metastasis. Cancer Res. 71, 4260–4268 (2011).
Liu, Y. et al. miR-10a suppresses colorectal cancer metastasis by modulating the epithelial-to-mesenchymal transition and anoikis. Cell Death Dis. 8, e2739 (2017).
Xie, Y. et al. MicroRNA-10b controls the metastasis and proliferation of colorectal cancer cells by regulating Krüppel-like factor 4. Artif. Cells Nanomed. Biotechnol. 47, 1722–1729 (2019).
Shibuya, H. et al. Clinicopathological and prognostic value of microRNA-21 and microRNA-155 in colorectal cancer. Oncology 79, 313–320 (2010).
Cellura, D. et al. miR-19-Mediated Inhibition of Transglutaminase-2 Leads to Enhanced Invasion and Metastasis in Colorectal Cancer. Mol. Cancer Res. 13, 1095–1105 (2015).
Asangani, I. A. et al. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 27, 2128–2136 (2008).
Feiersinger, F. et al. MiRNA-21 Expression Decreases from Primary Tumors to Liver Metastases in Colorectal Carcinoma. PLoS One 11, e0148580 (2016).
Ding, M. et al. Correlation analysis between liver metastasis and serum levels of miR‑200 and miR‑141 in patients with colorectal cancer. Mol. Med. Rep. 16, 7791–7795 (2017).
Cheng, D. et al. MicroRNA-20a-5p promotes colorectal cancer invasion and metastasis by downregulating Smad4. Oncotarget 7, 45199–45213 (2016).
Arabsorkhi, Z. et al. miR-298 plays a pivotal role in colon cancer invasiveness by targeting PTEN. J. Cell Physiol. 235, 4335–4350 (2020).
Li, J. et al. Inhibition of miR-15b decreases cell migration and metastasis in colorectal cancer. Tumour Biol. 37, 8765–8773 (2016).
Wang, Y. et al. Decrease of miR-622 expression suppresses migration and invasion by targeting regulation of DYRK2 in colorectal cancer cells. Onco Targets Ther. 10, 1091–1100 (2017).
Zhang, T. et al. Identifying the key genes and microRNAs in colorectal cancer liver metastasis by bioinformatics analysis and in vitro experiments. Oncol. Rep. 41, 279–291 (2019).
Lam, C. S. et al. Identification of microRNA 885-5p as a novel regulator of tumor metastasis by targeting CPEB2 in colorectal cancer. Oncotarget 8, 26858–26870 (2017).
Ling, H. et al. The clinical and biological significance of MIR-224 expression in colorectal cancer metastasis. Gut 65, 977–989 (2016).
Zou, G., Wang, R. & Wang, M. Clinical response and prognostic significance of serum miR-497 expression in colorectal cancer. Cancer Biomark. 25, 11–18 (2019).
Qiu, Y. Y. et al. MicroRNA-497 and bufalin act synergistically to inhibit colorectal cancer metastasis. Tumour Biol. 35, 2599–2606 (2014).
Qiu, Y. et al. microRNA-497 inhibits invasion and metastasis of colorectal cancer cells by targeting vascular endothelial growth factor-A. Cell Prolif. 49, 69–78 (2016).
Zhang, N., Shen, Q. & Zhang, P. miR-497 suppresses epithelial-mesenchymal transition and metastasis in colorectal cancer cells by targeting fos-related antigen-1. Onco Targets Ther. 9, 6597–6604 (2016).
Hong, S. et al. LncRNA AGAP2-AS1 augments cell viability and mobility, and confers gemcitabine resistance by inhibiting miR-497 in colorectal cancer. Aging 12, 5183–5194 (2020).
Li, W. et al. miR-133a acts as a tumor suppressor in colorectal cancer by targeting eIF4A1. Tumour Biol. 39, 1010428317698389 (2017).
Li, W. et al. miRNA-99b-5p suppresses liver metastasis of colorectal cancer by down-regulating mTOR. Oncotarget 6, 24448–24462 (2015).
Ding, Q. et al. APOBEC3G promotes liver metastasis in an orthotopic mouse model of colorectal cancer and predicts human hepatic metastasis. J. Clin. Investig. 121, 4526–4536 (2011).
Chen, D. L. et al. Identification of microRNA-214 as a negative regulator of colorectal cancer liver metastasis by way of regulation of fibroblast growth factor receptor 1 expression. Hepatology 60, 598–609 (2014).
Cristóbal, I. et al. Downregulation of miR-214 is specific of liver metastasis in colorectal cancer and could play a role determining the metastatic niche. Int J. Colorectal Dis. 29, 885 (2014).
Li, Y. et al. Tumor-suppressive miR-26a and miR-26b inhibit cell aggressiveness by regulating FUT4 in colorectal cancer. Cell Death Dis. 8, e2892 (2017).
Laudato, S. et al. P53-induced miR-30e-5p inhibits colorectal cancer invasion and metastasis by targeting ITGA6 and ITGB1. Int J. Cancer 141, 1879–1890 (2017).
Fan, M. et al. MicroRNA-30b-5p functions as a metastasis suppressor in colorectal cancer by targeting Rap1b. Cancer Lett. 477, 144–156 (2020).
Stiegelbauer, V. et al. miR-196b-5p Regulates Colorectal Cancer Cell Migration and Metastases through Interaction with HOXB7 and GALNT5. Clin. Cancer Res. 23, 5255–5266 (2017).
Luo, M. et al. MicroRNA-432-5p inhibits cell migration and invasion by targeting CXCL5 in colorectal cancer. Exp. Ther. Med. 21, 301 (2021).
Pan, S. et al. MiR-328-3p inhibits cell proliferation and metastasis in colorectal cancer by targeting Girdin and inhibiting the PI3K/Akt signaling pathway. Exp. Cell Res. 390, 111939 (2020).
Hong, Y. G. et al. miR-365a-3p regulates ADAM10-JAK-STAT signaling to suppress the growth and metastasis of colorectal cancer cells. J. Cancer 11, 3634–3644 (2020).
Hata, T. et al. Identification of microRNA-487b as a negative regulator of liver metastasis by regulation of KRAS in colorectal cancer. Int J. Oncol. 50, 487–496 (2017).
Guo, L. et al. MicroRNA-143-3p inhibits colorectal cancer metastases by targeting ITGA6 and ASAP3. Cancer Sci. 110, 805–816 (2019).
Mokutani, Y. et al. Down-Regulation of microRNA-132 is Associated with Poor Prognosis of Colorectal Cancer. Ann. Surg. Oncol. 23, 599–608 (2016).
Bleau, A. M. et al. miR-146a targets c-met and abolishes colorectal cancer liver metastasis. Cancer Lett. 414, 257–267 (2018).
Hansen, T. F. et al. Intra-tumoural vessel area estimated by expression of epidermal growth factor-like domain 7 and microRNA-126 in primary tumours and metastases of patients with colorectal cancer: a descriptive study. J. Transl. Med. 13, 10 (2015).
Zhong, L. et al. p38 and JNK pathways control E-selectin-dependent extravasation of colon cancer cells by modulating miR-31 transcription. Oncotarget 8, 1678–1687 (2017).
Chen, H. Y. et al. miR-103/107 promote metastasis of colorectal cancer by targeting the metastasis suppressors DAPK and KLF4. Cancer Res. 72, 3631–3641 (2012).
Larsen, C. J. [Metastatic colonization of the liver by colorectal tumor cells is abolished by two micro-RNAs (miR-483-5p et miR-551a)]. Bull. Cancer 102, 713–715 (2015).
Schepers, A. G. et al. Lineage tracing reveals Lgr5+ stem cell activity in mouse intestinal adenomas. Science 337, 730–735 (2012).
Hadjimichael, C. et al. Common stemness regulators of embryonic and cancer stem cells. World J. Stem Cells 7, 1150–1184 (2015).
Zeuner, A. et al. Colorectal cancer stem cells: from the crypt to the clinic. Cell Stem Cell 15, 692–705 (2014).
Das, P. K., Islam, F. & Lam, A. K. The Roles of Cancer Stem Cells and Therapy Resistance in Colorectal Carcinoma. Cells 9, 1392 (2020).
Vermeulen, L. et al. The developing cancer stem-cell model: clinical challenges and opportunities. Lancet Oncol. 13, e83–e89 (2012).
Murata, K. et al. Ascl2-Dependent Cell Dedifferentiation Drives Regeneration of Ablated Intestinal Stem Cells. Cell Stem Cell 26, 377–390.e6 (2020).
de Sousa, E. M. F. & de Sauvage, F. J. Cellular Plasticity in Intestinal Homeostasis and Disease. Cell Stem Cell 24, 54–64 (2019).
Shimokawa, M. et al. Visualization and targeting of LGR5(+) human colon cancer stem cells. Nature 545, 187–192 (2017).
de Sousa, E. M. F. et al. A distinct role for Lgr5(+) stem cells in primary and metastatic colon cancer. Nature 543, 676–680 (2017).
Fumagalli, A. et al. Plasticity of Lgr5-Negative Cancer Cells Drives Metastasis in Colorectal Cancer. Cell Stem Cell 26, 569–578.e7 (2020).
Wang, D. et al. Prostaglandin E2 Promotes Colorectal Cancer Stem Cell Expansion and Metastasis in Mice. Gastroenterology 149, 1884–1895.e4 (2015).
Morgan, A. E., Davies, T. J. & Mc, A. M. The role of DNA methylation in ageing and cancer. Proc. Nutr. Soc. 77, 412–422 (2018).
Li, S. et al. Inhibition of DNMT suppresses the stemness of colorectal cancer cells through down-regulating Wnt signaling pathway. Cell Signal 47, 79–87 (2018).
Mahdi, M. R. et al. Modulation of the Endothelin System in Colorectal Cancer Liver Metastasis: Influence of Epigenetic Mechanisms? Front. Pharm. 11, 180 (2020).
Grillet, F. et al. Circulating tumour cells from patients with colorectal cancer have cancer stem cell hallmarks in ex vivo culture. Gut 66, 1802–1810 (2017).
Tellez-Gabriel, M., Heymann, M. F. & Heymann, D. Circulating Tumor Cells as a Tool for Assessing Tumor Heterogeneity. Theranostics 9, 4580–4594 (2019).
Castro-Giner, F. & Aceto, N. Tracking cancer progression: from circulating tumor cells to metastasis. Genome Med. 12, 31 (2020).
Fang, C. et al. Prognostic value of CD133(+) CD54(+) CD44(+) circulating tumor cells in colorectal cancer with liver metastasis. Cancer Med. 6, 2850–2857 (2017).
Wei, C. et al. Crosstalk between cancer cells and tumor associated macrophages is required for mesenchymal circulating tumor cell-mediated colorectal cancer metastasis. Mol. Cancer 18, 64 (2019).
Loo, J. M. et al. Extracellular metabolic energetics can promote cancer progression. Cell 160, 393–406 (2015).
Wang, Y. N. et al. CPT1A-mediated fatty acid oxidation promotes colorectal cancer cell metastasis by inhibiting anoikis. Oncogene 37, 6025–6040 (2018).
Bu, P. et al. Aldolase B-Mediated Fructose Metabolism Drives Metabolic Reprogramming of Colon Cancer Liver Metastasis. Cell Metab. 27, 1249–1262.e4 (2018).
Fong, Y. et al. Clinical score for predicting recurrence after hepatic resection for metastatic colorectal cancer: analysis of 1001 consecutive cases. Ann. Surg. 230, 309–318 (1999).
Therkildsen, C. et al. The predictive value of KRAS, NRAS, BRAF, PIK3CA and PTEN for anti-EGFR treatment in metastatic colorectal cancer: A systematic review and meta-analysis. Acta Oncol. 53, 852–864 (2014).
Kuipers, E. J. et al. Colorectal cancer. Nat. Rev. Dis. Prim. 1, 15065 (2015).
Hagness, M. et al. Liver transplantation for nonresectable liver metastases from colorectal cancer. Ann. Surg. 257, 800–806 (2013).
Adam, R. et al. Managing synchronous liver metastases from colorectal cancer: a multidisciplinary international consensus. Cancer Treat. Rev. 41, 729–741 (2015).
Hammill, C. W. et al. Outcome after laparoscopic radiofrequency ablation of technically resectable colorectal liver metastases. Ann. Surg. Oncol. 18, 1947–1954 (2011).
Goodman, K. A. et al. Dose-escalation study of single-fraction stereotactic body radiotherapy for liver malignancies. Int J. Radiat. Oncol. Biol. Phys. 78, 486–493 (2010).
Schefter, T. E. et al. A phase I trial of stereotactic body radiation therapy (SBRT) for liver metastases. Int J. Radiat. Oncol. Biol. Phys. 62, 1371–1378 (2005).
Abdalla, E. K. Commentary: Radiofrequency ablation for colorectal liver metastases: do not blame the biology when it is the technology. Am. J. Surg. 197, 737–739 (2009).
Cremolini, C. et al. Upfront FOLFOXIRI plus bevacizumab and reintroduction after progression versus mFOLFOX6 plus bevacizumab followed by FOLFIRI plus bevacizumab in the treatment of patients with metastatic colorectal cancer (TRIBE2): a multicentre, open-label, phase 3, randomised, controlled trial. Lancet Oncol. 21, 497–507 (2020).
Falcone, A. et al. Phase III trial of infusional fluorouracil, leucovorin, oxaliplatin, and irinotecan (FOLFOXIRI) compared with infusional fluorouracil, leucovorin, and irinotecan (FOLFIRI) as first-line treatment for metastatic colorectal cancer: the Gruppo Oncologico Nord Ovest. J. Clin. Oncol. 25, 1670–1676 (2007).
Datta, J. et al. Role of Hepatic Artery Infusion Chemotherapy in Treatment of Initially Unresectable Colorectal Liver Metastases: A Review. JAMA Surg. 154, 768–776 (2019).
Price, T. J. et al. Panitumumab versus cetuximab in patients with chemotherapy-refractory wild-type KRAS exon 2 metastatic colorectal cancer (ASPECCT): a randomised, multicentre, open-label, non-inferiority phase 3 study. Lancet Oncol. 15, 569–579 (2014).
Saltz, L. B. et al. Bevacizumab in combination with oxaliplatin-based chemotherapy as first-line therapy in metastatic colorectal cancer: a randomized phase III study. J. Clin. Oncol. 26, 2013–2019 (2008).
Bokemeyer, C. et al. Fluorouracil, leucovorin, and oxaliplatin with and without cetuximab in the first-line treatment of metastatic colorectal cancer. J. Clin. Oncol. 27, 663–671 (2009).
Pohl, M. & Schmiegel, W. Therapeutic Strategies in Diseases of the Digestive Tract - 2015 and Beyond Targeted Therapies in Colon Cancer Today and Tomorrow. Dig. Dis. 34, 574–579 (2016).
Qin, S. et al. Efficacy and Tolerability of First-Line Cetuximab Plus Leucovorin, Fluorouracil, and Oxaliplatin (FOLFOX-4) Versus FOLFOX-4 in Patients With RAS Wild-Type Metastatic Colorectal Cancer: The Open-Label, Randomized, Phase III TAILOR Trial. J. Clin. Oncol. 36, 3031–3039 (2018).
Tveit, K. M. et al. Phase III trial of cetuximab with continuous or intermittent fluorouracil, leucovorin, and oxaliplatin (Nordic FLOX) versus FLOX alone in first-line treatment of metastatic colorectal cancer: the NORDIC-VII study. J. Clin. Oncol. 30, 1755–1762 (2012).
Venook, A. P. et al. Effect of First-Line Chemotherapy Combined With Cetuximab or Bevacizumab on Overall Survival in Patients With KRAS Wild-Type Advanced or Metastatic Colorectal Cancer: A Randomized Clinical Trial. JAMA 317, 2392–2401 (2017).
Van Cutsem, E. et al. Cetuximab plus irinotecan, fluorouracil, and leucovorin as first-line treatment for metastatic colorectal cancer: updated analysis of overall survival according to tumor KRAS and BRAF mutation status. J. Clin. Oncol. 29, 2011–2019 (2011).
Hirata, F. et al. Effect of bevacizumab plus XELOX (CapeOX) chemotherapy on liver natural killer cell activity in colorectal cancer with resectable liver metastasis. Ann. Gastroenterol. Surg. 2, 383–393 (2018).
Guo, J. et al. Nano Codelivery of Oxaliplatin and Folinic Acid Achieves Synergistic Chemo-Immunotherapy with 5-Fluorouracil for Colorectal Cancer and Liver Metastasis. ACS Nano. 14, 5075–5089 (2020).
Lu, M. et al. Activation of the mTOR Pathway by Oxaliplatin in the Treatment of Colorectal Cancer Liver Metastasis. PLoS One 12, e0169439 (2017).
Nordlinger, B. et al. Perioperative FOLFOX4 chemotherapy and surgery versus surgery alone for resectable liver metastases from colorectal cancer (EORTC 40983): long-term results of a randomised, controlled, phase 3 trial. Lancet Oncol. 14, 1208–1215 (2013).
Bridgewater, J. A. et al. Systemic chemotherapy with or without cetuximab in patients with resectable colorectal liver metastasis (New EPOC): long-term results of a multicentre, randomised, controlled, phase 3 trial. Lancet Oncol. 21, 398–411 (2020).
Okines, A. et al. Surgery with curative-intent in patients treated with first-line chemotherapy plus bevacizumab for metastatic colorectal cancer First BEAT and the randomised phase-III NO16966 trial. Br. J. Cancer 101, 1033–1038 (2009).
Folprecht, G. et al. Tumour response and secondary resectability of colorectal liver metastases following neoadjuvant chemotherapy with cetuximab: the CELIM randomised phase 2 trial. Lancet Oncol. 11, 38–47 (2010).
Shaheen, R. M. et al. Tyrosine kinase inhibition of multiple angiogenic growth factor receptors improves survival in mice bearing colon cancer liver metastases by inhibition of endothelial cell survival mechanisms. Cancer Res. 61, 1464–1468 (2001).
Yorozuya, K. et al. TSU-68 (SU6668) inhibits local tumor growth and liver metastasis of human colon cancer xenografts via anti-angiogenesis. Oncol. Rep. 14, 677–682 (2005).
Postow, M. A., Sidlow, R. & Hellmann, M. D. Immune-Related Adverse Events Associated with Immune Checkpoint Blockade. N. Engl. J. Med. 378, 158–168 (2018).
Ganesh, K. et al. Immunotherapy in colorectal cancer: rationale, challenges and potential. Nat. Rev. Gastroenterol. Hepatol. 16, 361–375 (2019).
Ishida, Y. et al. Induced expression of PD-1, a novel member of the immunoglobulin gene superfamily, upon programmed cell death. EMBO J. 11, 3887–3895 (1992).
Zamani, M. R. et al. PD-1/PD-L and autoimmunity: A growing relationship. Cell Immunol. 310, 27–41 (2016).
Sharpe, A. H. & Pauken, K. E. The diverse functions of the PD1 inhibitory pathway. Nat. Rev. Immunol. 18, 153–167 (2018).
Lei, Q. et al. Resistance Mechanisms of Anti-PD1/PDL1 Therapy in Solid Tumors. Front Cell Dev. Biol. 8, 672 (2020).
Masugi, Y. et al. Tumour CD274 (PD-L1) expression and T cells in colorectal cancer. Gut 66, 1463–1473 (2017).
Wyss, J. et al. Stromal PD-1/PD-L1 Expression Predicts Outcome in Colon Cancer Patients. Clin. Colorectal Cancer 18, e20–e38 (2019).
Taube, J. M. et al. Association of PD-1, PD-1 ligands, and other features of the tumor immune microenvironment with response to anti-PD-1 therapy. Clin. Cancer Res. 20, 5064–5074 (2014).
Llosa, N. J. et al. The vigorous immune microenvironment of microsatellite instable colon cancer is balanced by multiple counter-inhibitory checkpoints. Cancer Discov. 5, 43–51 (2015).
Madore, J. et al. PD-L1 expression in melanoma shows marked heterogeneity within and between patients: implications for anti-PD-1/PD-L1 clinical trials. Pigment Cell Melanoma Res. 28, 245–253 (2015).
Liu, J. et al. Molecular dissection of CRC primary tumors and their matched liver metastases reveals critical role of immune microenvironment, EMT and angiogenesis in cancer metastasis. Sci. Rep. 10, 10725 (2020).
Le, D. T. et al. PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Alvarado-Bachmann, R. et al. The incidence of mismatch repair gene defects in colorectal liver metastases. Mol. Med. Rep. 10, 1003–1006 (2014).
Schrock, A. B. et al. Tumor mutational burden is predictive of response to immune checkpoint inhibitors in MSI-high metastatic colorectal cancer. Ann. Oncol. 30, 1096–1103 (2019).
Rowshanravan, B., Halliday, N. & Sansom, D. M. CTLA-4: a moving target in immunotherapy. Blood 131, 58–67 (2018).
Fiegle, E. et al. Dual CTLA-4 and PD-L1 Blockade Inhibits Tumor Growth and Liver Metastasis in a Highly Aggressive Orthotopic Mouse Model of Colon Cancer. Neoplasia 21, 932–944 (2019).
Halama, N. et al. Hepatic metastases of colorectal cancer are rather homogeneous but differ from primary lesions in terms of immune cell infiltration. Oncoimmunology 2, e24116 (2013).
Zhou, G. et al. Blockade of LAG3 enhances responses of tumor-infiltrating T cells in mismatch repair-proficient liver metastases of colorectal cancer. Oncoimmunology 7, e1448332 (2018).
Pedroza-Gonzalez, A. et al. Activated tumor-infiltrating CD4+ regulatory T cells restrain antitumor immunity in patients with primary or metastatic liver cancer. Hepatology 57, 183–194 (2013).
Unitt, E. et al. Compromised lymphocytes infiltrate hepatocellular carcinoma: the role of T-regulatory cells. Hepatology 41, 722–730 (2005).
Pedroza-Gonzalez, A. et al. GITR engagement in combination with CTLA-4 blockade completely abrogates immunosuppression mediated by human liver tumor-derived regulatory T cells ex vivo. Oncoimmunology 4, e1051297 (2015).
Liu, W. et al. Discovery of core gene families associated with liver metastasis in colorectal cancer and regulatory roles in tumor cell immune infiltration. Transl. Oncol. 14, 101011 (2021).
Iida, N. et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342, 967–970 (2013).
Viaud, S. et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science 342, 971–976 (2013).
Liu, W. T. et al. LPS-induced CXCR4-dependent migratory properties and a mesenchymal-like phenotype of colorectal cancer cells. Cell Adh. Migr. 11, 13–23 (2017).
Hsu, R. Y. et al. LPS-induced TLR4 signaling in human colorectal cancer cells increases beta1 integrin-mediated cell adhesion and liver metastasis. Cancer Res. 71, 1989–1998 (2011).
Malvicini, M. et al. A Tricin Derivative from Deschampsia antarctica Desv. Inhibits Colorectal Carcinoma Growth and Liver Metastasis through the Induction of a Specific Immune Response. Mol. Cancer Ther. 17, 966–976 (2018).
André, T. et al. Pembrolizumab in Microsatellite-Instability-High Advanced Colorectal Cancer. N. Engl. J. Med. 383, 2207–2218 (2020).
Shao, S. et al. Targeting High Expressed α(5) β(1) Integrin in Liver Metastatic Lesions To Resist Metastasis of Colorectal Cancer by RPM Peptide-Modified Chitosan-Stearic Micelles. Mol. Pharm. 15, 1653–1663 (2018).
Willenbacher, E. et al. Curcumin: New Insights into an Ancient Ingredient against Cancer. Int. J. Mol. Sci. 20, 1808 (2019).
Kasprzak, A. & Adamek, A. Role of Endoglin (CD105) in the Progression of Hepatocellular Carcinoma and Anti-Angiogenic Therapy. Int. J. Mol. Sci. 19, 3887 (2018).
Takahashi, N. et al. Association of serum endoglin with metastasis in patients with colorectal, breast, and other solid tumors, and suppressive effect of chemotherapy on the serum endoglin. Clin. Cancer Res. 7, 524–532 (2001).
Mehta, C. R., Liu, L. & Theuer, C. An adaptive population enrichment phase III trial of TRC105 and pazopanib versus pazopanib alone in patients with advanced angiosarcoma (TAPPAS trial). Ann. Oncol. 30, 103–108 (2019).
Paauwe, M. et al. Endoglin targeting inhibits tumor angiogenesis and metastatic spread in breast cancer. Oncogene 35, 4069–4079 (2016).
Paauwe, M. et al. Endoglin Expression on Cancer-Associated Fibroblasts Regulates Invasion and Stimulates Colorectal Cancer Metastasis. Clin. Cancer Res. 24, 6331–6344 (2018).
Rosen, L. S. et al. A phase I first-in-human study of TRC105 (Anti-Endoglin Antibody) in patients with advanced cancer. Clin. Cancer Res. 18, 4820–4829 (2012).
Grady, M. E., Composto, R. J. & Eckmann, D. M. Cell elasticity with altered cytoskeletal architectures across multiple cell types. J. Mech. Behav. Biomed. Mater. 61, 197–207 (2016).
Humphrey, J. D., Dufresne, E. R. & Schwartz, M. A. Mechanotransduction and extracellular matrix homeostasis. Nat. Rev. Mol. Cell Biol. 15, 802–812 (2014).
Shen, Y. et al. Reduction of Liver Metastasis Stiffness Improves Response to Bevacizumab in Metastatic Colorectal Cancer. Cancer Cell 37, 800–817.e7 (2020).
Firestein, R. et al. CDK8 is a colorectal cancer oncogene that regulates beta-catenin activity. Nature 455, 547–551 (2008).
Liang, J. et al. CDK8 Selectively Promotes the Growth of Colon Cancer Metastases in the Liver by Regulating Gene Expression of TIMP3 and Matrix Metalloproteinases. Cancer Res. 78, 6594–6606 (2018).
Chen, W. Nanoparticle Self-Lighting Photodynamic Therapy For Cancer Treatment. J. Biomed. Nanotechnol. 4, 369–276 (2008).
Fan, W., Huang, P. & Chen, X. Overcoming the Achilles' heel of photodynamic therapy. Chem. Soc. Rev. 45, 6488–6519 (2016).
Ma, L., Zou, X. & Chen, W. A new X-ray activated nanoparticle photosensitizer for cancer treatment. J. Biomed. Nanotechnol. 10, 1501–1508 (2014).
Guo, H., Qian, H., Idris, N. M. & Zhang, Y. Singlet oxygen-induced apoptosis of cancer cells using upconversion fluorescent nanoparticles as a carrier of photosensitizer. Nanomedicine 6, 486–495 (2010).
Wang, C., Tao, H., Cheng, L. & Liu, Z. Near-infrared light induced in vivo photodynamic therapy of cancer based on upconversion nanoparticles. Biomaterials 32, 6145–6154 (2011).
Chen, W. & Zhang, J. Using Nanoparticles to Enable Simultaneous Radiation and Photodynamic Therapies for Cancer Treatment. J. Nanosci. Nanotechnol. 6, 1159–1166 (2006).
Liu, F., Chen, W., Wang, S. P. & Joly, A. G. Investigation of Water-Soluble X-ray Luminescence Nanoparticles For Photodynamic Activation. Appl. Phys. Lett. 92, 43901 (2008).
Zou, X. et al. X-ray Induced Nanoparticles Based Photodynamic Therapy of Cancer. Future. Nanomedicine 9, 2339–2351 (2014).
Zhang, W. et al. Ultra-high FRET efficiency NaGdF4: Tb(3+)-Rose Bengal biocompatible nanocomposite for X-ray excited photodynamic therapy application. Biomaterials 184, 31–40 (2018).
Ma, L. et al. A new Cu–cysteamine complex: structure and optical properties. J. Mater. Chem. C. 2, 4239–4246 (2014).
Wang, Y. et al. A new type of cuprous-cysteamine sensitizers: Synthesis, optical properties and potential applications. Matetials Today Phys. 19, 100435 (2021).
Guo, H. et al. Ions Self-Activated White Emitting Phosphor under Single Excitation. Adv. Funct. Mater. 25, 6833–6838 (2015).
Huang, X. et al. Investigation of copper-cysteamine nanoparticles as a new photosensitizer for anti-hepatocellular carcinoma. Cancer Biol. Therap. 20, 812–825 (2019).
Ma, L., Zou, X., Chen, W. & New, A. X-ray Induced Nanoparticle Photosensitizers For Cancer Treatment. J. Biomed. Nanotechnol. 10, 1501–1508 (2014).
Liu, Z. et al. The Investigation of Copper Cysteamine Nanoparticles as a new type of radiosenstizers for Colorectal Carcinoma. Sci. Rep. 7, 9290 (2017).
Shrestha, S. et al. X-ray Induced Photodynamic Therapy with pH-Low Insertion Peptide Targeted Copper-Cysteamine Nanoparticles in Mice. PNAS 116, 16823–16828 (2019).
Yao, M. et al. Modality for Cancer Treatment—Nanoparticle Mediated Microwave Induced Photodynamic Therapy. J. Biomed. Nanotechnol. 12, 1835–1851 (2016).
Chu, X. et al. Exploration of Graphitic‑C3N4 Quantum Dots for Microwave-Induced Photodynamic Therapy. ACS Biomater. Sci. Eng. 3, 1836–1844 (2017).
Chu, X. et al. Exploration of TiO2 Nanoparticle Mediated Microdynamic Therapy on Cancer Treatment. Nanomedicine. NBM 18, 272–281 (2019).
Pandey N. K. et al. Aggregation-induced emission luminogens for highly effective microwave dynamic therapy. Bioact. Mater. 7, 112–125 (2021).
Pandey, N. K. et al. A facile method for the synthesis of copper-cysteamine nanoparticles and study of ROS production for cancer treatment. J. Mater. Chem. B 7, 6630–6642 (2019).
Wang, P. et al. Nanosonosensitization by Using Copper–Cysteamine Nanoparticles Augmented Sonodynamic Cancer Treatment. Part Part Syst. Charact. 35, 1700378 (2018). (1700371-1700310).
Huang, L. et al. Exploration of Copper-Cysteamine nanoparticles as a new type of agents for Antimicrobial photodynamic inactivation. J. Biomed. Nanotechnol. 15, 2142–2148 (2019).
Shi, L. et al. The effectiveness and safety of copper-cysteamine nanoparticle mediated X-PDT for cutaneous squamous cell carcinoma and melanoma. Nanomedicine 14, 2027–2043 (2019).
Zhang, Q. et al. Use of copper-cysteamine nanoparticles to simultaneously enable radiotherapy, oxidative therapy and immunotherapy for melanoma treatment. Signal Transduct. Target. Ther. 5, 58 (2020).
Chen, X. et al. Study of copper-cysteamine based X-ray induced photodynamic therapy and its effects on cancer cell proliferation and migration in a clinical mimic setting. Bioact. Mater. 7, 504–514 (2021).
Wang, J. H. et al. The Long Noncoding RNA LOC441461 (STX17-AS1) Modulates Colorectal Cancer Cell Growth and Motility. Cancers. 12, 3171 (2020).
Liang, W. C. et al. The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget 6, 22513–22525 (2015).
Shan, Y. et al. LncRNA SNHG7 sponges miR-216b to promote proliferation and liver metastasis of colorectal cancer through upregulating GALNT1. Cell Death Dis. 9, 722 (2018).
Di, W. et al. Long noncoding RNA SNHG14 facilitates colorectal cancer metastasis through targeting EZH2-regulated EPHA7. Cell Death Dis. 10, 514 (2019).
Zhuang, M. et al. MALAT1 sponges miR-106b-5p to promote the invasion and metastasis of colorectal cancer via SLAIN2 enhanced microtubules mobility. EBioMedicine 41, 286–298 (2019).
Li, C. et al. Long intergenic non-coding RNA LINC00485 exerts tumor-suppressive activity by regulating miR-581/EDEM1 axis in colorectal cancer. Aging 13, 3866–3885 (2021).
Li, C. & Zhou, H. Circular RNA hsa_circRNA_102209 promotes the growth and metastasis of colorectal cancer through miR-761-mediated Ras and Rab interactor 1 signaling. Cancer Med. 9, 6710–6725 (2020).
Ma, X., Lv, L. & Xing, C. Circ_ 0115744 acts as miR-144 sponge to promote and predict the metastasis of colorectal cancer. Aging 13, 5892–5905 (2021).
Wang, R. et al. MicroRNA-195 suppresses angiogenesis and metastasis of hepatocellular carcinoma by inhibiting the expression of VEGF, VAV2, and CDC42. Hepatology 58, 642–653 (2013).
Wang, L. G. & Gu, J. Serum microRNA-29a is a promising novel marker for early detection of colorectal liver metastasis. Cancer Epidemiol. 36, e61–e67 (2012).
Acknowledgements
We are grateful for the support by the Natural Science Foundation of China (81773293, 81873640, 81970569, 82000756), the Key Project of Science and Technology Program of Hunan Provincial Science and Technology Department (2015GK3117, 2017WK2063). We also would like to acknowledge the supports from Guangxi Jialouyuan Medical Inc., Solgro, and the distinguished award from UT Arlington.
Author information
Authors and Affiliations
Contributions
H.Z. and Z.L.: searched the information, materials and updates and wrote the drafts. Y.W., X.W., E.H.A., L.Y. and X.R.: helped for editing, checking and formatting; W.C., L.X., Y.R. and Y.W.: proposed the concept of design, organized, conceived, led the writing and supervising. H.Z. edited and formatted the figures, E.H.A. checked the English and W.C. oversaw and reproved the entire work. All authors have read and approved the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, H., Liu, Z., Wang, Y. et al. Colorectal liver metastasis: molecular mechanism and interventional therapy. Sig Transduct Target Ther 7, 70 (2022). https://doi.org/10.1038/s41392-022-00922-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-022-00922-2
This article is cited by
-
METTL3 recruiting M2-type immunosuppressed macrophages by targeting m6A-SNAIL-CXCL2 axis to promote colorectal cancer pulmonary metastasis
Journal of Experimental & Clinical Cancer Research (2024)
-
Tumor-derived exosomal ADAM17 promotes pre-metastatic niche formation by enhancing vascular permeability in colorectal cancer
Journal of Experimental & Clinical Cancer Research (2024)
-
Non-alcoholic fatty liver disease promotes liver metastasis of colorectal cancer via fatty acid synthase dependent EGFR palmitoylation
Cell Death Discovery (2024)
-
Periostin–TGF-β feedforward loop contributes to tumour-stroma crosstalk in liver metastatic outgrowth of colorectal cancer
British Journal of Cancer (2024)
-
A chemical proteomics approach for global mapping of functional lysines on cell surface of living cell
Nature Communications (2024)