Abstract
MicroRNAs (miRNAs), a class of endogenous single-stranded short noncoding RNAs, have emerged as vital epigenetic regulators of both pathological and physiological processes in animals. They direct fundamental cellular pathways and processes by fine-tuning the expression of multiple genes at the posttranscriptional level. Growing evidence suggests that miRNAs are implicated in the onset and development of rheumatoid arthritis (RA). RA is a chronic inflammatory disease that mainly affects synovial joints. This common autoimmune disorder is characterized by a complex and multifaceted pathogenesis, and its morbidity, disability and mortality rates remain consistently high. More in-depth insights into the underlying mechanisms of RA are required to address unmet clinical needs and optimize treatment. Herein, we comprehensively review the deregulated miRNAs and impaired cellular functions in RA to shed light on several aspects of RA pathogenesis, with a focus on excessive inflammation, synovial hyperplasia and progressive joint damage. This review also provides promising targets for innovative therapies of RA. In addition, we discuss the regulatory roles and clinical potential of extracellular miRNAs in RA, highlighting their prospective applications as diagnostic and predictive biomarkers.
Similar content being viewed by others
Introduction
Rheumatoid arthritis (RA) is a chronic, inflammatory, autoimmune disorder characterized by progressive and irreversible joint damage as a consequence of sustained synovitis. Patients with RA typically experience pain, swelling and stiffness in multiple joints bilaterally. Extra-articular organs and systems may also be involved as RA proceeds. Cardiovascular disease is a significant and potentially lethal comorbidity, and the risk of its occurrence is intimately related to RA disease activity.1 RA is a global health issue, and the prevalence of RA is ~240 per 100 000 individuals. Furthermore, RA prevalence is increasing with the rapidly expanding aging population throughout the world.2 The etiology and pathogenesis of RA, involving genetic predisposition, environmental triggers and epigenetic modifications, are highly complex and remain to be elucidated.3,4 Synovial hyperplasia, persistent inflammation, articular cartilage degradation and bone erosion are generally considered the cardinal hallmarks of RA pathology.5,6
In the past two decades, considerable insights into RA pathophysiology have fostered the development of novel therapeutics in the forms of both biologic and small molecule inhibitors.7 Despite this progress, a notable percentage of RA patients still suffer from progressive disability, loss of function, decreased quality of life and diminished life expectancy due to inadequate treatment. Thus, a deeper understanding of the cellular and molecular mechanisms underlying synovial hyperplasia, chronic inflammation and tissue destruction during RA is required to achieve improvement in clinical outcomes.
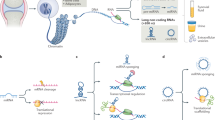
MicroRNAs (miRNAs), a class of short noncoding RNA molecules, are posttranscriptional regulators of gene expression. The first known miRNA, lin-4, was recognized in Caenorhabditis elegans in 1993.8 For the past few years, the field of miRNA biology has expanded considerably. MiRNAs have been shown to play essential roles in the developmental timing of stage-specific cell lineages and control multiple biological processes related to development, differentiation, growth, and metabolism.9 The widely observed deregulation of miRNAs in diverse diseases, including cancer,10 cardiovascular disorders and inflammatory diseases,11,12 has rendered miRNAs attractive candidate targets. In addition, the role of miRNAs in the development of RA has recently attracted attention, as a growing body of evidence has revealed the aberrant expression of numerous miRNAs in clinical samples and experimental models of RA. Studying deregulated miRNAs in RA will provide novel mechanistic and therapeutic insights into the progression of the disease (Fig. 1).
Schematic diagram of miRNAs in normal development, homeostasis and RA progression. a In normal synovial joints, many cell types serve as major sources of short noncoding RNAs, including miRNAs, which are transcribed from DNA. Precursor miRNAs are cleaved and processed into miRNAs, which further function within parental cells and neighboring functional cells and can be secreted into biological fluids such as blood and joint fluid as free molecules or extracellular vesicles. b In normal conditions in synovial joints, miRNAs are mainly produced by a variety of cells involved in the composition of bone remodeling, osteoimmunology and synovial systems. MiRNAs procedurally regulate well-organized bone remodeling, appropriate levels of immune responses and the secretion of normal synovial fluid. Conversely, persistent synovial hyperplasia, progressive inflammation, and subsequent destruction of affected joints are prominent characteristics of RA. The development, differentiation and homeostasis of diverse cell populations in the inflamed synovial compartment are dysregulated as a result of miRNA deregulation. Interactions between these abnormal cell types with disrupted function ultimately contribute to RA pathogenesis. miRNA microRNA, APC antigen-presenting cell, RANKL receptor activator of NF-κB ligand, FLS fibroblast-like synoviocyte, MMP matrix metalloproteinase
Cellular landscape in RA pathology
RA is a systemic disease characterized by a complex pathogenesis involving interactions between various cell types located in synovial compartments and peripheral blood, rather than resulting from a single pathogenic factor.5,6 These cell populations, comprising fibroblast-like synoviocytes (FLSs), innate and adaptive immune cells, and bone-related cells, change in number, status, and behavior in response to the dynamic microenvironment, which perturbs cytokine secretion, intracellular signaling networks and homeostasis and consequently leads to corresponding pathology. Multiple dysfunctional cell types and the crosstalk between these pathogenic cells collectively contribute to the onset, progression and perpetuation of RA.
Fibroblast-like synoviocytes (FLSs)
FLSs are highly specialized mesenchymal-derived cells that are a prominent component of the synovial lining layer of diarthrodial joints. Under normal physiological conditions, FLSs express components of the extracellular matrix (ECM) and synovial fluid to lubricate and nourish cartilage surfaces, thereby maintaining the homeostasis of joints. However, accumulating studies have identified FLSs as key players in many pathogenic events in the RA synovium. In pathological conditions, such as RA, FLSs increase rapidly in number and are redistributed in the synovium and joints, exhibiting heterogeneity across different locations within the synovium and across different joints.13,14 FLSs in RA display unique aggressive behavior that arises from their reduced rate of apoptosis; deregulated proliferation, migration, and invasion; and improved ability to secrete inflammatory mediators and matrix metalloproteinases (MMPs) into the synovial fluid.13,15,16 Researchers have detected constitutive overexpression of the tumor suppressor p53 and mutations in the gene encoding p53 in FLSs from RA patients. Interestingly, these mutations are similar or identical to those found in certain tumors but are not found in skin fibroblasts from the same patients or in FLSs from osteoarthritis (OA) patients.17
Immune cells
Immune dysfunction is undoubtedly a key factor in RA pathogenesis. A variety of immune cells are implicated in and mediate autoimmune inflammation in RA, among which CD4+ T lymphocytes (T cells) and monocyte-macrophage lineage cells are acknowledged as the two most significant cellular components.18
Adaptive immune cells
The significant number of infiltrating T cells, primarily CD4+ T cells, in the inflammatory synovial milieu and circulation of patients with RA implies that T cells are one of the major players in RA pathology. CD4+ T cells include helper T cells (Th cells), regulatory T cells (Tregs) and follicular helper T cells (Tfh cells). The specific CD4+ T-cell subpopulations and their secreted cytokines play distinct and essential roles in the pathogenesis of RA and experimentally induced arthritis.19 Pathogenic Th17 cells secreting proinflammatory cytokines, such as interleukin (IL)-17A, IL-17F, and IL-22, are regarded as the predominant positive regulators of the immune response in RA.5 Attention has also been increasingly focused on the imbalance of Th17 cells/Tregs. Forkhead box P3 (Foxp3)-positive Tregs, a subset with low proliferative capacity, can secrete cytokines such as IL-4, IL-10 and TGF-β to exert immunosuppressive effects, thereby managing immune homeostasis and inducing immune tolerance. Treg differentiation was reported to be significantly inhibited in patients with RA.20 However, others have noted high levels of proliferating Tregs in inflamed joints of RA patients, suggesting that expansion of Tregs is induced by autoimmune inflammatory environments.21 Despite the controversy concerning Treg frequencies, it is clear that Tregs become less functional or even pathogenic in the setting of RA. Factors controlling Th17 cell and Treg plasticity may be attractive targets for RA immunotherapy.
The role of B lymphocytes (B cells) in RA pathogenesis goes beyond autoantibody production, antigen presentation and cytokine release. Toll-like receptors (TLRs) drive the differentiation of self-reactive B cells into plasma cells that synthesize autoantibodies.22 The best known RA-associated autoantibodies are rheumatoid factor (RF) and anti-citrullinated protein antibody (ACPA). In particular, ACPA has the most pronounced predictive value for the onset of RA in symptomatic, at-risk patients.23
Innate immune cells
Innate immunity is the first line of defense against the invasion of pathogenic microorganisms. Monocytes and macrophages, as the most important components of the innate immune system, are also key effectors of synovitis in RA. Synovial tissue-resident macrophages in the steady state reside in healthy synovium and perform homeostatic functions but attain an activated phenotype under inflammatory conditions.24 Despite having controversial cellular origins,25 these tissue-resident macrophages are deemed to self-renew locally throughout adulthood without continuous repopulation from circulating monocytes.26 In the context of RA, in addition to activated tissue-resident macrophages, massive bone marrow-derived monocytes accumulate in the blood and consistently migrate into the inflamed joints, where they can differentiate into macrophages to replenish the synovial macrophage pool and thus induce inflammatory reactions.27,28 However, infiltrating macrophages are difficult to distinguish from tissue-resident macrophages in diseased human tissues due to the absence of specific markers. Notably, macrophages display remarkable plasticity and heterogeneity, with dynamic phenotypes and distinct functions that are modulated by the surrounding microenvironmental signals.29 For simplicity, researchers have constructed a polarization model that classifies macrophage phenotypes on a continuum in which classically activated or inflammatory macrophages (M1) and alternatively activated or reparative macrophages (M2) represent the two opposite extremes.30 While disruption of the M1/M2 ratio, which is usually inclined toward M1 polarization, is considered to be one of the pathogenic factors during RA,31 the molecular mechanisms that govern M1/M2 polarization remain incompletely defined.
In addition to the monocyte-macrophage lineage, several other innate effector cells, such as dendritic cells (DCs) and neutrophils, are also critical players in the RA process. Indeed, the large amounts of neutrophils, which mainly reside in synovial fluid, contribute to sustained synovitis in RA via the synthesis of proteases, prostaglandins and reactive oxygen intermediates.32 DCs might be involved in the promotion of tolerance by mediating the generation and maintenance of Tregs as well as T-cell unresponsiveness.33 Conversely, DCs exhibit antigen presentation capacity and can thus foster the priming and/or effector differentiation of self-reactive T cells. The majority of antigen-presenting cells (APCs) in the inflamed RA synovium are fully differentiated DCs expressing high levels of class I and II major histocompatibility complex (MHC) and several T-cell costimulatory molecules.34 In summary, the role of DCs in the pathogenesis of RA is complex and dichotomous, and DCs may induce either tolerance or autoimmunity based on the integrated signals received in the microenvironment of synovial joints.
Bone-related cells
Osteoclasts
RA is an independent risk factor for generalized osteopenia and osteoporosis. Osteoclasts, long considered the only effector cells in vivo that participate in bone remodeling by resorbing bone matrix, have emerged as an important cell type in the process of bone erosion in RA. Osteoclasts are multinuclear giant cells that are differentiated from monocyte-macrophage lineage precursor cells derived from bone marrow hematopoietic cells.35 The bone matrix can be degraded through matrix enzymes synthesized and secreted by mature osteoclasts, including tartrate-resistant acid phosphatase type 5 (TRAcP), cathepsin K and MMP9. Classically, in the presence of macrophage colony-stimulating factor (M-CSF), the receptor activator of nuclear factor kappa B (NF-κB) ligand (RANKL) pathway and costimulatory signaling pathways synergistically trigger osteoclastogenesis.36,37 In the pathogenesis of RA, osteoclasts colonize the interface between the inflamed synovium and the periarticular bone surface. The abundant cytokines, particularly TNF-α, IL-1, IL-6 and IL-17, autoantibodies and RANKL in the inflammatory synovial milieu collectively accelerate bone resorption by fostering osteoclast differentiation and activation.35,38,39
Osteoblasts
Despite considerable attention being paid to osteoclasts in RA inflamed joints, increasing evidence has suggested that the synovial microenvironment in RA has the potential to induce not only osteoclast differentiation but also osteogenic inhibition. The disruption of bone homeostasis is the result of bone metabolism disorder, in terms of not only excessive bone resorption mediated by osteoclasts but also insufficient bone formation mediated by osteoblasts.35,40 Osteoblasts, which originate from mesenchymal stem cells (MSCs) in the bone marrow stroma, are indispensable for the synthesis of the bone matrix and its subsequent mineralization. Impaired osteoblast differentiation and function in RA limit the capacity of bone remodeling and result in severe and ongoing arthritic bone loss at erosion sites.40
Chondrocytes
Irreversible destruction of articular cartilage is one of the core pathological characteristics of RA.5 Articular cartilage is a complex tissue that is composed of chondrocytes and cartilage matrix produced by chondrocytes. The cartilage matrix is composed of collagen and proteoglycan aggregates, which are the structural components of joint tissue and provide biomechanical strength.41 Because there is no blood supply in articular cartilage tissue, its nutrition comes entirely from synovial fluid secreted by normal synovium. Therefore, the normal synovial fluid secreted by healthy synovium is important for the maintenance and function of physiological articular cartilage. In the pathogenesis of RA, due to abnormal synovial fluid secretion and invading synovial pannus formation, abundant proinflammatory cytokines are produced in the synovium, which increases MMP secretion by chondrocytes.42 Driven by synovial inflammatory microenvironments, anomalous ECM metabolism in coordination with deranged chondrocyte development and function leads to disruption of cartilage homeostasis.43
Crosstalk between pathogenic cell types
In the context of RA, starting with an early breach of immune tolerance, dysfunctional leukocytes in circulation massively infiltrate the synovial tissue, accompanied by uncontrolled accumulation of resident synoviocytes. This ongoing process results in the overproduction of proinflammatory mediators and therefore the chronicity of synovitis.6,44 Activated synoviocytes also secrete various proangiogenic factors to induce angiogenesis in RA.45 Sprouting angiogenesis and pathological vascular remodeling support the increased transendothelial infiltration of blood-derived leukocytes in the inflamed joints. Antigen-specific T-cell/APC interactions result in the proliferation and differentiation of T cells and the secretion of abundant lymphokines to trigger B-cell and macrophage activation. B cells differentiate into plasma cells that mediate autoantibody release and immune complex formation. These immune complexes subsequently induce macrophages to produce proinflammatory cytokines and facilitate osteoclast-mediated bone resorption.46,47 Intriguingly, expansion of RA-FLSs promotes hyperplastic transformation of the synovial lining layer and its transformation into an invasive pannus nourished by neovascularization. The pannus exhibits tumor-like behaviors that mediate structural joint destruction through invasion of cartilage and periarticular bone at the cartilage-bone interface.48,49 RANKL expressed by FLSs and osteoblasts is also associated with bone erosion because it fosters osteoclastogenesis. Furthermore, in the presence of inflammatory factors, FLSs, chondrocytes and osteoblasts can secrete MMPs to degrade articular cartilage. In light of the above pathological features, cellular interaction-dependent core immune dysfunction and altered tissue homeostasis seem to underlie the disease (Fig. 2).
Basic biology of MiRNAs
MiRNA biogenesis and function
MiRNAs are a group of evolutionarily conserved endogenous noncoding RNA molecules 22 nucleotides (nt) in length. These small RNAs have tightly regulated expression patterns and serve as posttranscriptional regulators of gene expression.50 Most miRNAs are expressed in a temporal and tissue-specific manner, meaning that miRNAs are differentially expressed in various stages of organism development and that the expression of identical miRNAs may vary in different organs and tissues.51 Genes encoding miRNAs in mammals are generally localized within the introns of protein-coding genes. However, there are also some miRNAs that originate from intergenic regions and act as independent transcription units.52,53 The process of miRNA biogenesis involves multiple steps (Fig. 3), comprising a canonical pathway dependent on Drosha processing and a noncanonical pathway in which pre-miRNAs can be alternatively generated via spliceosome machinery.50,53,54,55 Mature miRNAs mediate the inhibition of gene expression at the posttranscriptional level, which is associated with cellular function maintenance. The efficiency and mode of gene silencing depend on whether the miRNA has sufficient complementarity to the target mRNA.9,56 A perfect match between an miRNA and the 3′ UTR of its target leads to irreversible mRNA decay, while imperfect complementarity promotes the repression of translation at the initiation or elongation stages.57,58
Schematic overview of the biogenesis and mechanism of action of miRNAs. MiRNAs encoded in introns of protein-coding genes and intergenic areas are initially transcribed by RNA Pol II in the nucleus. In the canonical pathway (left), pri-miRNAs are recognized and processed by the microprocessor complex, consisting of two RNase III enzymes, Drosha and DGCR8, releasing hairpin-structured pre-miRNAs. In the noncanonical pathway (right), however, pre-miRNA hairpins are generated from introns that undergo splicing and debranching, bypassing the step of Drosha cleavage. The nuclear export of pre-miRNAs is mediated by Exportin-5 in complex with Ran-GTP. Cytoplasmic pre-miRNAs are further cleaved by the endonuclease Dicer, aided by TRBP, to yield ~22-nucleotide miRNA duplexes. One strand of the duplex (the guide strand) is retained as mature miRNA and incorporated into RISC (containing AGO proteins), whereas the other strand (the passenger strand) is degraded. Mature miRNAs fine-tune gene expression by guiding RISC interaction with complementary sites of target mRNAs. The mechanisms of miRNA-mediated gene silencing, including accelerated cleavage and restricted translation of mRNAs, seem to largely depend on the degree of complementarity between seed sequences of miRNAs and the 3’ UTRs of target mRNAs. miRNA microRNA, Pol II RNA polymerase II, pri-miRNA primary transcript, DGCR8 DiGeorge syndrome critical region gene 8, pre-miRNA precursor miRNA, TRBP TAR RNA-binding protein, RISC RNA-induced silencing complex, AGO Argonaute, 3′ UTR 3′ untranslated region
More than 60% of protein-coding genes in mammals seem to have conserved miRNA target sites.59 Each individual gene can be simultaneously targeted by multiple miRNAs, while a single miRNA species has the capacity to interact with a broad range of target genes.9,53 Thus, functional miRNAs constitute a huge and complex regulatory network involving extensive signaling cascades, thereby playing fundamental roles in diverse biological processes, including organism development, metabolism, homeostasis, and cellular behaviors such as proliferation, differentiation, migration, and apoptosis. Disturbed expression and functions of these miRNAs under specific circumstances may bring about pathological phenotypes and/or anomalous biological outcomes.9
Extracellular miRNAs and cell-to-cell communication
Numerous miRNAs can be encapsulated within cell-derived membranous structures called exosomes that can be transferred to neighboring or distant cells, thereby facilitating crosstalk between donor cells and recipient cells.60,61 Exosomes are the most well-studied class of extracellular vesicles (EVs) and have an average diameter of 50–150 nm.62 It has been well established that the transfer of metabolites and biological information between different cell types via exosomal miRNAs is crucial for the coordination of various physiological and pathological processes.63,64
The generation, trafficking, and function of exosomes depend on their different intercellular origins and can be modified by other pathological signals and stimuli that the cell may receive.62 In essence, exosome biogenesis starts with the formation of the early endosome. Subsequently, intraluminal vesicles (ILVs) are formed and accumulate through inward budding of the endosomal membrane during the maturation of multivesicular endosomes (MVEs). Finally, ILVs are secreted into the extracellular space upon fusion of MVEs with the plasma membrane.60,65 ILVs contain and carry complex biomolecules, including various types of proteins, lipids, and nucleic acids, such as DNA sequences, mRNAs, miRNAs, and other noncoding RNAs. Sorting and clustering of these exosomal cargoes into exosomes are largely dependent on particular endosomal sorting machineries.62
Interestingly, the enrichment of exosomal miRNAs is different for each cell type. To date, the precise mechanisms involved in the specific loading of miRNAs into exosomes remain elusive. Several potential pathways have been proposed to perform exosomal miRNA sorting. The presence of specific sequence motifs in certain miRNAs seems to determine whether they are incorporated into exosomes or retained in the cell in a cell-specific fashion.66 Heterogeneous nuclear ribonucleoprotein A2B1 (hnRNPA2B1) in exosomes critically regulates this process by directly binding to these exosomal miRNAs. Specifically, preferential sumoylation of hnRNPA2B1 fosters its recognition of specific short motifs in miRNAs and guides their localization into exosomes.67 hnRNPA2B1 was the first protein identified to be implicated in the selectivity of miRNA loading. Likewise, several RNA-binding proteins that control the export of miRNAs into exosomes have been described. These include Syncrip,68,69 major vault protein (MVP),70 HuR,71 and Y-box protein I (YBX1),72,73 each of which has multiple potential miRNA targets. More recently, another two RNA-binding proteins, Alyref and Fus, were discovered to strongly recognize the CGGGAG motif and target miRNAs to exosomes.66 The 3′-end of sequence-dependent miRNAs is a potential sorting signal for governing the export of miRNAs as well. 3′-Adenylated miRNAs are preferentially retained in the cell, whereas 3′-uridylated miRNAs are highly exosome-enriched.74 In addition, the correlation between miRISC and MVEs suggests another miRNA sorting machinery.75 KRAS activity can modulate the colocalization of Ago2, a key component of miRISC, with MVEs. The KRAS–MEK signaling pathway may control the loading of specific exosomal miRNAs through Ago2 phosphorylation.76 Deficiency of AGO2 has been observed to restrict the enrichment of some preferentially exported miRNAs, such as miR-451, miR-150, and miR-142-3p, in exosomes derived from multiple cell types.77
In summary, a better understanding of exosome biogenesis and how miRNAs are differentially sorted into exosomes may be useful for the artificial packing of selected miRNAs into exosomes, aiding the development of engineered exosomes for gene therapy.
Deregulated MiRNAs in RA
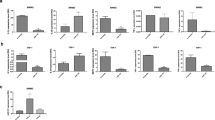
With the advances in several detection technologies, including microarrays and next-generation sequencing (NGS), aberrant changes in specific and global miRNA levels have been detected in many pathogenic cell types in RA.78 Notably, as the expression and function of miRNAs are cell type-specific,79 their overall contribution to RA progression—that is, whether they exacerbate or slow progression—is not entirely clear. Given the acknowledged role of miRNAs in fine-tuning gene expression, studying their complex effects on cellular function is bound to prove important for elucidating RA pathogenesis (Fig. 1).
MiRNA-mediated dysfunction in FLSs
Dysregulation of several miRNAs is responsible for the aberrant biological behaviors of FLSs in RA (Fig. 4). Various intracellular pathways are related to the altered expression of miRNAs, and the most significantly implicated pathways are the Wnt, NF-κB, Janus kinase/signal transducer and activator of transcription (JAK/STAT) and TLR pathways. The first two miRNAs found to be differentially expressed in RA-FLSs were miR-155 and miR-146a.80 The expression of miR-155 is prominently upregulated in RA-FLSs when they are stimulated with tumor necrosis factor (TNF)-α. In addition to its widely reported proinflammatory role in immune cells, miR-155 also upregulates the secretion of inflammatory factors from RA-FLSs.81,82 Of interest, miR-155 is assumed to protect against acquisition of a destructive phenotype by RA-FLSs by repressing the expression of MMP3 and MMP1.80,83 In this regard, further research is warranted to dissect the precise role of miR-155 in RA-FLSs and identify its direct target genes and pathways. MiR-146a, a well-described epigenetic regulator in immune cells, is also expressed at high levels in RA-FLSs upon stimulation with proinflammatory stimuli such as IL-1β.80 It was proposed that miR-146a orchestrates the inflammatory response, suppressing the proliferation, changing the metabolic state and restricting the osteoclastogenic potential of RA-FLSs. These actions are elicited by targeting TNF receptor associated factor 6 (TRAF6), an important component of the NF-κB pathway, and disrupting the receptor activator of NF-κB ligand (RANKL)/osteoprotegerin (OPG) ratio. MiR-146a deficiency has been demonstrated to be responsible for aggravated inflammatory joint damage in a model of TNF-driven arthritis.84 Likewise, miR-146a potently inhibits the secretion of inflammatory factors as well as connective tissue growth factor (CTGF) by RA-FLSs. Downregulated CTGF expression limits the proliferation of RA-FLSs and angiogenesis, thereby, to some extent, limiting pannus formation and attacking cartilage.85 The levels of miR-143 and miR-145 are also augmented in RA-FLSs compared to OA-FLSs.86 Overexpression of miR-143 enhances TNFα-induced pro-inflammatory signals by downregulating insulin-like growth factor binding protein 5 (IGFBP5). MiR-145 regulates semaphorin 3 A (SEMA3A) to render RA-FLSs susceptible to vascular endothelial growth factor 165 (VEGF165) stimulation, contributing to cell migration and invasion.86 In addition, it has been noted that miR-145-5p exacerbates RA progression by promoting nuclear translocation of p65 to activate NF-κB, which increases the levels of MMP-9.87
Deregulated miRNAs in RA-FLSs. Upregulation (red font) or downregulation (green font) of several miRNAs is observed in RA-FLSs. These deregulated miRNAs exert positive or negative regulation on the aggressive phenotypes, inflammatory mediator production, and potential to disrupt synovial joint tissues of RA-FLSs. miRNA microRNA, RA rheumatoid arthritis, FLS fibroblast-like synoviocyte, MMP matrix metalloproteinase
Elevated expression of miR-21, miR-26a-5p, miR-203, miR-221, miR-222, miR-323-3p and miR-346 in RA-FLSs has also been reported. Upregulated miR-26a-5p was found to promote the proliferation, invasion, and apoptosis tolerance of RA-FLSs through regulation of the PTEN/PI3K/AKT pathway.88 MiR-21 facilitates NF-κB nuclear translocation to activate the NF-κB pathway, resulting in enhanced proliferation of RA-FLSs.89 The robust expression of miR-203 in RA-FLSs, which relies on the regulation of hypomethylation, seems to increase the release of MMP-1 and IL-6 by controlling NF-κB pathway activity.90 Using miR-seq-based profiling, Pandis et al. were the first to identify miR-221/222 and miR-323-3p as overexpressed miRNAs in FLSs from human TNF transgenic mice and patients with RA. The predicted target genes of miR-221/222 and miR-323-3p exhibited considerable enrichment in a number of RA-related pathways. Importantly, miR-323-3p was suggested to be strongly implicated in RA pathogenesis due to its positive modulation of the WNT/cadherin signaling pathway.91 Furthermore, induced expression of miR-346 was observed in LPS-activated RA-FLSs. The expression of a nonreceptor tyrosine kinase related to the stabilization of multiple inflammatory cytokines, named Bruton’s tyrosine kinase, is indirectly repressed by miR-346 upregulation, thus mediating the inhibition of IL-18 release in RA-FLSs.92
In contrast, several pivotal miRNAs were found to be significantly reduced in RA-FLSs. Najm et al. demonstrated the anti-inflammatory and anti-erosion role of miR-17-5p in both RA-FLSs and CIA mice, which was associated with reduced IL-6 secretion, decreased immune cell infiltration and osteoclastogenesis inhibition through the JAK1-STAT3 pathway.93 Lin et al. concluded that low miR-22 levels in RA-FLSs enhance the expression of cysteine-rich protein 61 (Cyr61), which is a component of the ECM, contributing to synovial hyperplasia and pannus formation. Intriguingly, whereas wild-type (WT) p53 activates miR-22 transcription by binding to its promoter region, the mutant p53 frequently observed in RA synovial tissues lacks the ability to upregulate the expression of miR-22, leading to Cyr61 deficiency. This further supports the concept that somatic p53 mutations are one of the etiological factors of RA.94 Nakamachi et al. showed that forced expression of miR-124a in RA-FLSs led to suppression of cell proliferation and G1 phase arrest because miR-124a bound to the 3′-UTR of cyclin-dependent kinase 2 (CDK-2) and monocyte chemoattractant protein 1 (MCP-1) mRNA.95,96 Of note, the expression of miR-124a is dictated by the methylation status of its encoding gene. Consistent with what has been reported in tumor cells, the miR-124a gene promoter was found to be hypermethylated in RA synovial tissue samples from patients with RA but nonmethylated in samples from healthy subjects.97 As such, it is not surprising that 5-aza-2ʹ-deoxycytidine (5-AzadC) treatment in vitro can inhibit the proliferation of RA-FLSs and the synthesis of TNF-α via demethylation of the miR-124a gene.98 More recently, Wang et al. revealed that overexpression of forkhead box protein C1 (FoxC1) in RA-FLSs and CIA rats, which is negatively correlated with miR-141-3p expression, might promote the proliferation, migration, invasion, and inflammatory cytokine secretion of RA-FLSs. The authors also revealed that FoxC1 interacts directly with β-catenin to activate the canonical Wnt signaling pathway.99 In addition, differential expression of both miR-9 and miR-106b in RA-FLSs leads to shifts not only in the invasive properties of RA-FLSs but also in RANKL levels, which affects osteoclastogenesis.100,101
Decreased expression in RA‑FLSs was also reported for miR-19, miR-20a, miR-27a, miR-29a, miR-152, miR192, miR‑199a-3p, miR‑375, miR-34a, miR-15a/16, and miR-3926. MiR-19a/b can act as a negative regulator of the inflammatory response in RA-FLSs, which is supported by evidence showing that miR-19a/b diminishes IL-6 and MMP-3 production via direct control of TLR2 expression.102 MiR-27a inhibition in RA-FLSs significantly promotes cell migration and invasion by targeting the 3′ UTR of follistatin-like protein 1 (FSTL1) and further interfering with TLR4/NF-κB signaling.103 STAT3 and caveolin 1 (CAV1) have been identified as direct targets of miR-29a and miR-192 in RA-FLSs, respectively. MiR-29a-mediated STAT3 suppression and miR-192-mediated CAV1 inhibition can markedly restrict the expansion of RA-FLSs by reducing proliferation and triggering apoptosis. MiR-29a also has an anti-inflammatory role in RA-FLSs since it improves inflammatory cytokine levels.104,105 Moreover, decreased miR-199a-3p expression in RA-FLSs contributes to increased cell numbers. The growth-suppressive effects of miR-199a-3p were elicited by direct endogenous retinoblastoma 1 (RB1) inhibition, which resulted in an increase in caspase-3 activity and the Bax-Bcl-2 ratio.106 MiR-375 in RA-FLSs showed protective effects against synovial pathological changes that occurred as a result of decreased MMP-3 and fibronectin levels, as well as canonical Wnt signaling inactivation.107 Song et al. demonstrated substantially lower expression of miR-34a-5p in synovial tissues from patients with RA than in those from patients with OA, accompanied by robust proliferation of RA-FLSs and high TNF-α and IL-6 expression.108 Likewise, Wu et al. highlighted the therapeutic role of miR-34a-containing bone marrow mesenchymal stem cell (BMSC)-derived miR-34a in RA-FLSs. The authors’ investigation indicated that miR-34a ameliorates abnormal RA-FLS proliferation and inflammation by targeting cyclin I and further activating ATM/ATR/p53 signaling.109 Moreover, miR-20a, miR-15a/16 and miR-3926 negatively regulate the aggressive behavior of RA-FLSs and synovial inflammatory infiltration. The target genes of these miRNAs in RA-FLSs have been confirmed in various studies.110,111,112
MiRNA-mediated dysfunction in immune cells
Previous studies have proven that miRNAs function as ‘fine-tuners’ of the expression of proteins involved in the immune response. Multiple miRNAs with immunoregulatory effects ensure the development and normal function of the immune system as well as the maintenance of immune homeostasis by establishing a balance between activation and inhibition.113
Disturbed monocyte-macrophage lineage function
Research has found that the expression profiles of miRNAs are highly variable in distinct macrophage polarization states.114 As an advantage, the temporal and spatial specificity of miRNA expression patterns provides flexible adjustment capacity to meet the high plasticity of macrophages during the response to environmental stimuli. Over the past decade, the functional roles of miRNAs in many aspects of monocyte/macrophage biology, particularly in macrophage polarization, have been intensively investigated. Dysregulation of miRNA levels thus appears to contribute to RA pathogenesis by perturbing the normal function of the monocyte-macrophage lineage (Table 1).
Two extensively studied miRNAs, miR-155 and miR-146a, are strongly expressed in RA macrophages and exert proinflammatory and anti-inflammatory effects, respectively. Defective differentiation of circulating monocytes from patients with RA into M2 macrophages is associated with increased miR-155 expression.115 Conversely, miR-146a is differentially expressed in M1 macrophages and M2 macrophages and endows macrophages with the potential to preferentially differentiate toward an M2 phenotype by regulating the expression of Notch1 and PPARγ.116,117 Exposure to a variety of inflammatory stimuli or direct recognition of microbial components such as lipopolysaccharide (LPS) by TLRs regulates the expression of both miR-155 and miR-146a in macrophages in an NF-κB-dependent and temporally asymmetric manner.118,119 Mann et al. unveiled a combined positive and negative regulatory network controlling NF-κB signaling activity in which miR-155 expression is induced prior to miR-146a expression and plays a dominant role in the promotion of the inflammatory response.120 Mechanistically, miR-155 ensures robust NF-κB signaling activity via Src homology 2-containing inositol phosphatase-1 (SHIP1) and suppressor of cytokine signaling 1 (SOCS1) repression.120,121 TRAF6 and IL-1 receptor-associated kinase 1 (IRAK1), the encoded proteins of which are located downstream of TLR signaling, have been identified as target genes for posttranslational repression by miR-146a. MiR-146a, therefore, constitutes an essential part of a negative feedback loop in macrophages that attenuates the activation of NF-κB signaling and reduces the production of inflammatory factors.119 Nevertheless, since miR-146a expression in macrophages reaches saturation at sub-inflammatory levels,122 it appears that the abnormal miR-146a expression in RA is not sufficient to counteract the positive effect of miR-155 on NF-κB activity, thus contributing to the chronicity of inflammation. This idea was further verified in experimental CIA mice established by Nakasa et al., in which transfection of miR-146a, despite inducing partial relief, failed to eliminate synovial inflammation.123
MiR-155 also participates in the development of RA by promoting monocyte recruitment and retention at inflammation sites. Upregulation of miR-155 in RA monocytes not only diminishes their susceptibility to apoptosis124 but also augments proinflammatory chemokine production and decreases chemokine receptor expression.125 Mice lacking miR-155 are resistant to the development of articular inflammation.126 Furthermore, Ghani et al. revealed that the expression of miR-146a, regulated by PU.1, is indispensable for driving the myeloid cell differentiation/maturation process.127 Given the characteristic elevation of miR-146a expression in THP-1 cells in a tolerized state, miR-146a is also thought to play pivotal roles in endotoxin tolerance and endotoxin-induced cross-tolerance, suggesting that it has potential functions in preventing overreactive inflammation.128 When compared with WT controls, serum transfer arthritis (STA) K/BxN mice with miR-146a deletion show a statistically significant increase in the arthritis index.129
Increased expression levels of miR-223 have been shown in various compartments, such as plasma, serum, synovial tissues, and synovial fluid, in patients with RA. Many research teams have recognized the essential role of miR-223 in regulating macrophage polarization. For instance, miR-223 downregulation promotes the release of IL-6 and IL-1β by activating STAT3130 and improves the M1/M2 macrophage ratio by targeting TRAF6 and subsequently influencing the NF-κB signaling pathway.131 Zhuang et al. identified Pknox1 as another target gene of miR-223. Augmented miR-223 expression mediates the inhibition of Pknox1 and further fosters M2 polarization.132 Moreover, nucleotide-binding domain-(NOD)-like receptor family, pyrin domain containing 3 (NLRP3) inflammasome activity, which is reported to be associated with the initiation, progression and perpetuation of RA, is negatively controlled by miR-223 in macrophages, leading to decreased IL-1β production.133 However, an opposing perspective was raised by Ogando and coworkers. The authors validated that miR-223 is upregulated in RA-derived macrophages compared to OA-derived macrophages and reported that high miR-223 levels prevent aryl hydrocarbon receptor (AHR) activation in myeloid cells by reducing AHR nuclear translocator protein levels. Thus, miR-223 in macrophages appears to exhibit proinflammatory effects due to the repression of LPS-induced proinflammatory cytokine expression after AHR agonist treatment.134 Despite the controversy, the promising potential of miR-223 in modulating macrophage function can be seen in the growing list of its confirmed target genes. More data are needed to reveal the dual roles of miR-223 in macrophages, especially in RA disease conditions.
Forced high expression of miR-221-3p after TLR4 activation interfered with JAK3/STAT3 signaling in M2 macrophages, driving M2 macrophages to exhibit an M1 phenotype. This dysregulated balance of M1/M2 macrophage populations contributed to the proinflammatory cytokine/chemokine secretion profile in RA.135 In contrast, higher expression of miR-125a-5p was detected in M2 macrophages than in M1 macrophages. Elevated levels of miR-125a-5p after TLR2 or TLR4 activation promote alternative activation of macrophages, pointing toward an anti-inflammatory role of this miRNA.136 Administration of miR-125a-5p mimics in LPS-induced macrophages leads to diminished TRAF6 expression and TAK1 protein phosphorylation, which is correlated with an elevated proportion of M2 phenotype macrophages.137 In addition, Zhou et al. indicated that miR-144-5p overexpression suppresses the secretion of TNF-α, IL-6 and IL-8 in LPS-treated THP-1 macrophages concomitant with decreased cell activity by inhibiting TLR2 expression and impairing the NF-κB signaling pathway.138 MiR-17, miR-20a, and miR-106a potentiate phagocytosis and the inflammatory response of macrophages by posttranscriptionally reducing signal-regulatory protein α (SIRPα) synthesis.139
Clinical evidence has demonstrated that the NLRP3 inflammasome plays a crucial role in multifarious diseases, including RA.140,141 Further studies revealed that the NLRP3 expression level was associated with clinical arthritis severity and the extent of radiological destruction.142,143 In particular, activation of the NLRP3 inflammasome occurs mainly in the monocyte-macrophage lineage.144 The miRNAs that participate in RA pathogenesis through NLRP3 inflammasome activation have been broadly researched. Xie et al. stated that both miR‑33 levels and NLRP3 inflammasome activity were increased in clinical samples from patients with RA. Mechanistically, miR‑33 overexpression resulted in the accumulation of cellular ROS and enhanced NLRP3 levels, caspase‑1 activity and IL‑1β production in primary macrophages, thereby eliciting NLRP3 inflammasome activation.145 Yang et al. were the first to demonstrate that knockout of NLRP3 or intra-articular injection of miR-30a achieved significant therapeutic effects in TNFTG mice by remarkably ameliorating inflammatory joint destruction. Subsequently, functional studies further proved that miR-30a directly interacts with the 3’ UTR of NLRP3 and mediates NLRP3 inhibition.146
Decreased miR-let-7a expression was observed in monocytes from ACPA-positive RA patients. MiR-let-7a downregulation augments the inflammatory response by increasing ACPA-mediated ERK1/2 and JNK phosphorylation and inducing IL-1β release via Ras proteins.147 In addition, Zhu et al. reported that miR-let-7a can inhibit ACPA-induced macrophage activation by directly targeting HMGA2.148 In contrast, high expression of miR-let-7b provokes arthritic joint inflammation. MiR-let-7b has been recognized as a novel endogenous ligand for joint TLR-7. Interruption of miR-let-7b ligation to TLR-7 obviously impairs the M1 polarization of synovial macrophages in RA.149
Disturbed CD4+ T-cell function
Studies have shown pivotal roles of miRNAs in regulating the cellular processes of CD4+ T cells (Fig. 5).
Deregulated miRNAs in RA that affect the Th17/Treg ratio. One of the central events in RA pathology is the imbalance between Th17 cells and Tregs. In the context of RA, activated APCs capture and process autoantigens for subsequent presentation to naïve CD4+ T cells, which provokes CD4+ T-cell activation and differentiation. The differentiation and expansion of Th17 cells are enhanced, leading to overwhelming inflammation via excessive inflammatory cytokine (for example, IL-17) release. In parallel, the Treg proportion is decreased, and the suppressive properties of Tregs are impaired, which is correlated with disrupted immune homeostasis. A consequent increase in the Th17/Treg ratio accelerates RA progression. The differential expression of several miRNAs changes the frequency and function of Th17 cells (red box) and/or Tregs (green box), thereby promoting the pathogenic role of these cells in RA. miRNA microRNA, RA rheumatoid arthritis, Th17 cell T helper 17 cell, Treg regulatory T cell, APC antigen-presenting cell
The differentiation and activation of CD4+ T-cell subpopulations and their effector functions may be jointly controlled by the different miRNAs they express. Classically, RA has been thought of as a disease that is mediated by interferon-γ (IFN-γ)-producing Th1 cells. A series of early studies proposed some miRNAs as crucial regulators of the Th1 immune response. For instance, activated CD4+ T cells are observed to induce miR-155 expression, which facilitates Th1 differentiation by modifying the IFN-γRα chain.150 The miR-17-92 cluster is also involved in modulating the Th1 response, which is directed almost entirely by specific individuals within this cluster. Specifically, miR-17 and miR-19b act in synergy to promote the functions of Th1 cells, with three functional targets: TGFβRII, CREB1 and Pten.151 Conversely, Th1 differentiation is inhibited while abnormal IFN-γ production is corrected by miR-29.152
However, the Th1 cell subset is not responsible for all the mechanisms underlying RA based on previously published studies. In recent years, the role of proinflammatory Th17 cells in RA has drawn increasing attention. Patients with RA often have enhanced Th17 cell differentiation, which is at the expense of Treg differentiation, and these integrated changes favor systemic inflammation in RA.5,20,153 Jarid2 is a DNA-binding protein that recruits Polycomb repressive complex 2 (PRC2) to bind to chromatin and hence mediates silencing of cytokine genes in Th17 cells. During Th17 differentiation, miR-155 is highly expressed and contributes to the activation of Th17-related cytokine gene expression by decreasing Jarid2 expression.154 Correspondingly, mice deficient in miR-155 demonstrated decreased sensitivity to CIA and profoundly impaired Th17 polarization.126 There is a rich supply of data pointing to miR-155’s pathogenic role in both the innate and adaptive immune systems. Intriguingly, miR-155 is also essential for the maintenance of Treg homeostasis. Despite contributing to the suppressive function of Tregs, high expression of miR-155 also ensures increased Treg generation in cases of lymphodepletion by targeting SOCS1, a JAK-STAT signaling inhibitor.155,156 Given the limited number and controversial results of published studies, much uncertainty exists about the levels of miR-155 in Tregs its regulatory effect on the proportion of Tregs in RA. Elevated miR-155 expression in RA Tregs was observed by Kmiolek and coworkers,157 whereas Zhou and coworkers showed a decrease in miR-155 expression in RA Tregs.158 Further investigations focusing on specific cell subsets rather than entire populations are warranted to provide a comprehensive understanding of the role of miR-155 in RA immunity.
Another miRNA that is tightly associated with the normal status of Tregs is miR-146a. Unlike the case for miR-155, high miR-146a levels are required for Treg suppression of autoimmunity. MiR-146a in Tregs functions by directly restraining STAT1 activation rather than by impacting NF-κB signaling in myeloid cells.159 Robust expression of miR-146a, however, is restricted to Tregs of RA patients. MiR-146a deletion in Tregs of patients with active RA leads to a proinflammatory phenotype of Tregs, accompanied by intensive inflammatory cytokine secretion. Strikingly, the extent of this restriction was found to be positively correlated with disease activity, implying that miR-146a might maintain high expression levels in Tregs of healthy individuals or even in Tregs of patients with low disease activity.158
The ratio of Th17 cells to Tregs is also affected by ectopic expression of other miRNAs. The expression of miR-301a-3p in peripheral blood monocytes (PBMCs) from RA patients was higher than that in PBMCs from healthy controls. Upregulation of miR-301a-3p increased the levels of STAT3 by targeting protein inhibitor of activated STAT3 (PIAS3), resulting in a larger Th17 cell population.160 As an essential transcription factor involved in Th17 cell differentiation, STAT3 directly induces the production of IL-17, a representative cytokine of Th17 cells.161 Another miRNA, let-7g-5p, was recognized to be inversely correlated with the proportion of Th17 cells, and its expression is repressed in RA patients and CIA mice.162 A recent study proposed that let-7g-5p overexpression, which disrupts STAT3 expression, alleviates CIA symptoms and predominantly hinders the transdifferentiation of Tregs into Th17-like cells.163 Increased miR-26b-5p expression was also demonstrated to have a positive effect on CIA. The difference is that this was accomplished mainly by perturbing the conventional differentiation of Th17 cells.164 In the context of RA, the NF-κB signaling pathway activated by IL-6 and TNF-α from the inflammatory milieu triggers the sustained expression of miR-34a, reducing the levels of the transcription factor FoxP3, which is specifically expressed in Tregs and critically governs their development and function.165 Furthermore, some miRNAs display simultaneous effects on the differentiation of Th17 cells and Tregs. The disrupted Th17/Treg balance caused by aberrant miR-20a expression was suggested to be closely related to the activation of the NLRP3 inflammasome.166 MiR-21 deficiency in RA patients promotes the pathogenicity of Th17 cells via potent STAT3 expression and mediates Treg dysfunction via reduced FoxP3 levels.153 Jin et al. identified Maresin 1 (MaR1) as a mediator upstream of miR-21 that upregulates its expression. Not surprisingly, by modifying MaR1 in the CIA mouse model, the authors demonstrated that miR-21 decelerated RA progression and restored the appropriate Th17/Treg ratio.167 MiR-16 was found to maintain high levels in Th17 cells, while miR-16 levels were low in Tregs. This differential expression controls the levels of the transcription factors RORγt and FoxP3 at the transcriptional and translational levels, contributing to an elevated Th17/Treg ratio.168
Expansion of Tfh cells in RA orchestrates spontaneous germinal center (GC) formation and excessive autoantibody synthesis.169 There are limitations to existing data regarding the role of individual miRNAs in Tfh cell subsets. Among the relevant studies, the two with the strongest conclusions regarding the importance of miRNAs in the functional regulation of Tfh cells were performed on miR-146a and miR-155. Pratama et al. highlighted that upregulation of miR-146a in Tfh cells decreases their numbers and restricts Tfh cell responses, in line with the widely discussed role of miR-146a as an intracellular molecular brake in the orchestration of universal biological processes.170 In contrast, miR-155 was found to counterregulate the proliferation and differentiation of Tfh cells.171 Considering that knockdown of miR-155 completely reverses the autoimmune pathology in mice with miR-146a deletion, a miR-155-dependent mechanism is suggested to underly the uncontrolled accumulation of Tfh cells induced by loss of miR-146a.172 In view of the reported fundamental roles of miR-155 and miR-146a in Tfh cell development, whether these two important miRNAs exert their putative effects on Tfh cells in induced arthritis models or RA patients through predictable or unpredictable molecular mechanisms needs to be determined.
Disturbed B-cell function
Although many miRNAs have been implicated in the determination of B-cell fates, few have been validated to have clear effects within a specific disease model of RA. MiR-155 is the only miRNA that has been previously shown to be prevalently expressed in peripheral blood and synovial fluid CD19+ B cells and synovial tissue of RA patients. Overexpression of this miRNA induced by PU.1 inhibition was shown to positively regulate B-cell activation in RA, implying subsequent persistent production of autoantibodies.173 More recently, an in-depth analysis of miRNA expression profiles in blood CD19+ B cells revealed differences in the expression of 27 miRNAs between RA patients in remission treated with methotrexate (MTX) and healthy controls, and these miRNAs were predicted to target genes known to be crucial for B-cell development.174
Disturbed dendritic cell (DC) function
An miR-363/integrin αv/TGF-β axis associated with the promotion of Th17 cell polarization was identified in DCs from patients with RA.175 Analogously, it was proposed that forced expression of miR-34a facilitates DC activation in RA by targeting AXL. Mouse models with miR-34a deficiency are resistant to CIA and may not support the development of Th17 cells due to diminished DC-T-cell interactions in vivo.176 In addition to the suppressive effect of miR-34a on RA Tregs we discussed before, miR-34a also seems to serve as a dual regulator causing significant Th17/Treg ratio imbalance, which accelerates the progression of RA.
Disturbed neutrophil function
Rosa et al. noted impaired miRNA biogenesis in neutrophils from RA patients, as reflected by a global decline in miRNA levels that led to the pathogenic activation of neutrophils.177 Typically, Murata et al. reported that decreased expression of miR-451 in RA neutrophils contributes to their chemotaxis toward inflammation sites, which is induced by 14-3-3ζ- and Rab5a-mediated p38 MAPK phosphorylation. The authors suggested CPNE3 and Rab5a as two novel target genes directly silenced by miR-451.178 Nevertheless, as p38 MAPK signaling pathway activity showed no correlation with CPEN3, the role of CPNE3 and miR-451 in RA neutrophils requires further verification.
MiRNA-mediated dysfunction in bone-related cells
Disturbed osteoclast function
MiRNAs have been implicated in the biological regulation of osteoclast function.179 Pre-miRNAs are cleaved to duplex mature miRNAs by the endonuclease RNase III Dicer, and osteoclast-specific Dicer gene deficiency was proven to suppress osteoclastic bone resorption.180 Accordingly, dysregulated miRNA expression modulates osteoclast development and function, leading to the inhibition or promotion of bone destruction in RA (Fig. 6).
Deregulated miRNAs in RA affect osteoclastogenesis. Gross disturbances in skeletal structure and function in synovial joints of patients with RA are mostly the result of an imbalance in osteoclast-mediated bone remodeling. The discovery of RANKL signaling was indispensable for an in-depth understanding of the mechanisms underlying osteoclastogenesis and bone resorption. Dysregulated miRNA expression in RA, driven by the osteoimmune microenvironment, is capable of controlling the differentiation and maturation of osteoclasts through RANKL-dependent (left) and RANKL-independent (right) pathways, resulting in a progressive reduction in periarticular bone mass and subsequent bone erosion. miRNA microRNA, RA rheumatoid arthritis, RANKL receptor activator of nuclear factor-kappa B ligand
RANKL-dependent osteoclastogenesis signaling pathways
RANKL-dependent osteoclastogenesis signaling pathways are the main regulatory pathways related to overactivation of osteoclasts in bone disorders.37 Numerous miRNAs have been shown to be differentially expressed during osteoclast differentiation and exert crucial roles in RANKL-dependent osteoclast activation in RA.
First, the binding of RANKL to RANK largely regulates initial osteoclast precursor differentiation and is considered the trigger in osteoclast activation. The density of RANK on the cell surface and the local levels of RANKL affect the strength of RANKL/RANK signaling and downstream signal transduction.36 MiR-106b and miRNA-20a were found to inhibit RANKL expression by directly binding to the 3′UTR of RANKL mRNA, blocking osteoclast differentiation.181,182 One of our previous studies revealed that inhibition of miR-106b substantially repressed mature osteoclast formation in vitro and alleviated the pathologic process of arthritis in CIA mice, accompanied by a decreased RANKL/OPG ratio.183 Kong et al. also demonstrated that miR-20a agomir injection impeded osteoclast proliferation and differentiation and thus improved bone erosion in a CIA mouse model.182
The binding of RANKL to RANK induces the recruitment of the adaptor protein TRAF6, and this process is controlled by miR-125a and miR-146a.119,184 MiR-125a expression is dramatically downregulated during osteoclast differentiation. Overexpression of miR-125a restricted osteoclastogenesis because miR-125a targets TRAF6, whereas the transcription of miR-125a was negatively regulated by the binding of NFATc1 to the promoter of miR-125a, suggesting the existence of a TRAF6/NFATc1/miR-125a regulatory feedback loop.184 In a CIA rat model, miR-125a treatment effectively attenuated arthritis severity.185 Moreover, miR-146 was reported to blunt TLR and cytokine signaling by base-pairing with sequences in the 3′ UTRs of TRAF6.119
The RANK/TRAF6 signaling cascade activates the MAPK pathway, potentially leading to additional miRNA targets. Forced expression of miR-20a inhibited the proliferation and differentiation potential of RA osteoclasts via the TLR4/p38 pathway.182 Abnormally low expression of miR-143-3p was noted in both clinical and experimental samples from RA patients and arthritis animals. Importantly, treatment with MAKP inhibitors elevated miR-143-3p levels in osteoclasts from adjuvant-induced arthritis (AIA) rats. Upregulated miR-143-3p expression was found to mediate osteoclastogenesis suppression by diminishing the expression of the early-stage protein ERK1/2 and the late-stage protein JNK in MAPK signaling.186 In the study, the authors first investigated the differential expression of miR-143-3p in vivo and further verified the regulatory mechanism of osteoclast differentiation in vitro. The limitation is that the detailed therapeutic effects of miR-143-3p administration on bone erosion in arthritic animal models were not investigated.
Another pivotal downstream signaling pathway activated by the RANK/TRAF6 signaling cascade is the NF-κB pathway. The activation of NF-кB signaling is involved in the regulation of a wide range of pathological processes during RA, such as synovitis and immune dysfunction. de la Rica et al. demonstrated that NF-κB directs monocyte-to-osteoclast conversion through an miRNA-dependent mechanism. NF-κB p65 can bind the promoters of two miRNA clusters: miR-212/132 and miR-99b/let-7e/125a. The NF-κB-directed activation of these miRNAs downregulates the expression of certain monocyte-specific and immunomodulatory genes, mediating proper osteoclast differentiation.187 Of note, contradictory outcomes emerged when validating the efficacy of these two NF-κB-associated miRNA clusters in animal models of RA. Articular knockdown of miR-132 in AIA murine models ameliorated bone erosion in the joints,188 while miR-125a overexpression retarded arthritis progression in CIA rats.185 These contradictory results might be explained by the complicated molecular mechanisms of miRNAs in different cell types and cell developmental stages. However, whether miR-132 and miR-125a are associated with NF-κB pathway activity in osteoclasts in the setting of RA is still unclear. Although growing data have shown a significant correlation between NF-кB signaling and deregulated miRNA levels in RA, strikingly, there is a lack of knowledge on whether specific miRNAs affect osteoclast differentiation during RA via NF-κB modulation. Follow-up investigations should take into account the cell specificity and condition specificity of the cellular signaling pathway.
NFATc1 has recently been described as a direct target of miRNAs. Consistent with previous findings regarding the NFATc1-dependent suppressive effects of miR-124 on osteoclastogenesis,189 forced expression of miR-124 in the ankle joints of AIA rats attenuated bone erosion, and direct binding of miR-124 to the 3’UTR of NFATc1 was identified.190 Interestingly, despite the inability of miR-124 to target RANKL mRNA, the mRNA and protein expression of RANKL was reduced in the ankle tissues of rats treated with pre-miR-124.190 MiR-146a also exerts inhibitory effects on osteoclast differentiation of human PBMCs in a dose-dependent manner by reducing NFATc1, c-Jun, and TRAP expression. Intravenous administration of double-stranded miR-146a resulted in the suppression of cartilage and bone destruction in CIA mice.123,191 Further studies should be performed to validate the direct binding of miR-146a to the 3′ UTR of NFATc1 mRNA.
OPG binds with strong affinity to RANKL, which limits bone resorption by competitively preventing RANKL from binding to its receptor RANK. MiR-145-5p not only strongly participates in RA-FLS pathology as mentioned above but also exerts destructive functions in osteoclasts. Augmented miR-145-5p expression in osteoclasts leads to a significant increase in RANK and RANKL expression and a decrease in OPG levels in a time-dependent manner. OPG has been identified as a target of miR-145-5p in osteoclasts.192 Intravenous injection of an miR-145-5p agomir resulted in more severe joint damage in a CIA model.192
RANKL-independent osteoclastogenesis signaling pathways
Recent discoveries have also revealed the essential role of alternative pathways in RANKL-independent osteoclastogenesis. TNF, as an important cytokine in the pathophysiology of RA, can also elicit osteoclast differentiation.193 Ohnuma et al. explored the potential role of miR-124 in the RANKL-independent osteoclastogenesis pathway. The researchers suggested that miR-124 represses osteoclast differentiation in response to stimulation with TNF-α, IL-6, and M-CSF. The expression levels of osteoclast-specific genes and NFATc1 protein were reduced in miR-124 mimic-transfected osteoclasts.194 Miller et al. demonstrated that miR-182 is also a negative regulator of TNF-α-mediated osteoclastogenesis via regulation of Foxo3 and Maml1, which may reveal an effective therapeutic approach in RA.195 More in vivo experimental evidence is still urgently needed for translation of relevant miRNA-related strategies into the clinic. With the application of anti-TNF drugs in clinical practice, future studies may focus on whether miRNA-based therapies can act synergistically with anti-TNF therapy in RA.
The mitochondrial apoptosis pathway can regulate the proliferation and activity of osteoclasts.196 Reduced miR-96-5p expression in osteoclasts mediates the regulation of ATP binding cassette subfamily A member 1 (Abca1), contributing to an increased level of mitochondria-dependent apoptosis in vitro and attenuated arthritic bone loss in vivo.197
During osteoclast differentiation, miR-146a hinders osteoclast progenitor differentiation through a marked decrease in Relb transcript levels. Targeted delivery of miR-146a into Ly6Chigh monocytes was shown to effectively repress monocyte-to-osteoclast conversion and prevent joint destruction in CIA mice.129
Disturbed osteoblast function
MiRNAs have emerged as major regulators of osteogenesis.179 Maeda et al. identified 22 deregulated miRNAs in the inflamed synovium of serum-transfer arthritis K/BxN mice related to bone formation, and the putative candidate targets are components of the Wnt and bone morphogenetic protein (BMP) pathways.198 MiR-218 establishes a positive feedback loop in osteoblasts to promote osteogenic differentiation.199 Specifically, miR-218 is abundantly expressed in response to canonical Wnt signaling. By suppressing multiple Wnt antagonists, such as DKK proteins, overexpression of miR-218 in turn reinforces Wnt signaling. Of interest, it appears that activation of Wnt signaling by miR-218 also increases the osteogenic differentiation of RA-FLSs, which contributes to the repair of bone erosion in RA.200 A limitation of current studies on miRNAs regulating osteoblast-mediated bone formation in RA is that most of them have only validated diverse miRNA–mRNA interactions using in vitro experiments. Sufficient evidence to prove the therapeutic efficacy of these miRNAs in RA patients and/or animal models is still lacking, making clinical translation challenging.
It has been proposed that Akt signaling cascade activation leads to miR-93, miR-129-3p, and miR-590-3p depletion in RA osteoblasts.201,202,203 Downregulated miR-93 expression mediates induction of oncostatin M, which is a pleiotropic cytokine of the IL-6 family that participates in RA pathogenesis.201 Loss of miR-129-3p in osteoblasts directly induces IL-17 protein synthesis at the posttranscriptional level and facilitates monocyte migration during RA.202 As precursors of macrophages, osteoclasts and DCs, monocytes tend to commit to a certain differentiation pathway based on the surrounding conditions upon migration into localized tissues. Massive monocyte infiltration in inflamed joints of RA is therefore sufficient to support the inflammatory reaction and bone destruction. Hyperplastic pannus nourished by neovascularization is a hallmark of RA, and prior research has revealed coupling of osteogenesis and angiogenesis.204 Due to the direct interaction of miR-590-3p with IL-18 mRNA, a low level of miR-590-3p in osteoblasts upregulates IL-18 expression and subsequently promotes the migration and tube formation of endothelial progenitor cells (EPCs).203 In addition, as Cyr61 is abundantly expressed in the synovial fluid of RA patients. This ECM protein is capable of inhibiting miR-126 and miR-518-5p expression in osteoblasts via the protein kinase C-alpha (PKC-α) and MAPK signaling cascades, respectively. In vitro functional research has clarified that miR-126 and miR-518-5p directly bind to the 3′ UTR of VEGF and the chemokine MCP-1 (also known as CCL2), respectively, leading to restriction of EPC angiogenesis and monocyte migration. Knockdown of Cyr61 in osteoblasts effectively prevents CIA progression and reduces disease severity.205,206 Similar to miR-126, in both in vitro and in vivo experiments, miR-16-5p was demonstrated to affect angiogenesis of EPCs by decreasing VEGF levels in osteoblasts. The expression of miR-16-5p in osteoblasts is regulated by c-Src-dependent FAK signaling.207
These findings characterize the tightly regulated expression patterns and pleiotropic effects of miRNAs. Aberrant miRNA expression profiles in RA osteoblasts control structural joint damage from multiple angles, impacting not only osteoblast differentiation but also cell chemotaxis and angiogenesis.
Disturbed chondrocyte function
An investigation launched by Chen et al. utilized NGS technologies and bioinformatics analyses to identify the expression profile of miRNAs in RA chondrocytes and to screen a series of candidate target genes.208 Overexpression of the long noncoding RNAs (lncRNAs) ZNF667-AS1, HOTAIR and MEG3 in LPS-induced chondrocytes stimulated chondrogenic proliferation and alleviated inflammation by regulating the production of inflammatory factors such as IL-6, IL-17 and IL-23, as well as the proliferation-related protein Ki67. Importantly, the protective effects of these three lncRNAs were effectively abrogated by miR-523-3p, miR-138, and miR-141, which participate in the activation of the JAK/STAT, NF-κB and AKT/mTOR signaling pathways, respectively.209,210,211 In a study in primary articular chondrocytes, miR-23a diminished IL-17-dependent proinflammatory mediator secretion, which was first proven to be mediated by direct interaction with IKKα, a kinase of the NF-κB pathway.212 Due to its induction of abnormal levels of type II collagen, MMP-13, COX-2, and IL-6, forced expression of miR-498 was suggested to alter the cellular signature via mTOR silencing in LPS-treated chondrocytes and thus aggravate ECM degradation.213 In addition, in vitro experiments conducted by Zhou et al. indicated that the arrest of chondrocyte apoptosis as a result of miR-27b-3p overexpression was related to the control of HIPK2 expression.214 Rats treated with interventions targeting miR-26a were shown to be resistant to arthritis in an experimental arthritis model established by Jiang et al. The authors also highlighted that high amounts of miR-26a regulate chondrogenic proliferation and apoptosis to increase cartilage destruction via CTGF repression.215
MiR-140, specifically expressed in cartilage, is fundamental to the maintenance of cartilage function and homeostasis.216 The regulatory role of miR-140 in chondrocytes and the underlying molecular mechanisms have been broadly investigated in OA.217,218,219 The aberrant synovial microenvironment governs cartilage damage in the context of RA, and an RA-FLS-based function of miR-140 has been identified.220 Consequently, it is reasonable to assume that miR-140 is also implicated in chondrocyte fate determination in RA. According to the study by Chen et al. mentioned above, deregulated miR-140-3p in RA chondrocytes might target FGF9, controlling the cell cycle to affect chondrogenic proliferation.208 This finding is in line with the results from several miRNA prediction databases, but further experimental verification is needed.
Roles of exosomal MiRNAs in RA progression
Growing data have revealed the tight association of exosomal miRNAs with RA pathogenesis.60 In addition to being recognized as a novel class of endocrine factors for cellular self-regulation, miRNAs might also serve as paracrine messengers to establish intercellular communication in the context of RA. Recent studies exploring the cellular sources of exosomes in RA have focused on MSCs, synoviocytes, and leukocytes (Table 2).
FLS-derived exosomal miRNAs
Synovium-derived miRNAs not only orchestrate synovial inflammation but also have a regulatory role in bone and cartilage erosion in RA.198 MiRNAs secreted from cells of synovial origin potentially tune gene expression within osteoblasts to govern osteogenesis in RA through exosomal transfer. For instance, overexpression of synovium-derived miR-221-3p impedes osteoblast maturation in RA by modulating Dickkopf2 (DKK2) protein levels to affect Wnt signaling activity.198 Exosomes secreted from RA-FLSs carrying miR-486-5p were demonstrated to potentiate the proliferation, differentiation, and mineralization of osteoblasts and thus accelerate bone repair and ameliorate CIA symptoms. Mechanistically, direct interplay between miR-486-5p and TOB1 upregulates BMP/Smad signaling.221 Moreover, given that RA-FLS-derived exosomes containing miR-106b significantly disrupts the normal cellular processes of chondrocytes by targeting the PDK4 gene, miR-106b suppression has enormous potential in attenuating cartilage injury in CIA and RA progression.222
The manipulation of synovium-derived exosomes with certain miRNAs may direct T-cell differentiation. Exosomes secreted from FLSs with forced expression of miR-424 under hypoxic conditions were shown to disrupt the Th17/Treg balance, increasing the proportion of Th17 cells and simultaneously inhibiting Treg differentiation via FOXP3 inhibition.223
MSC-derived exosomal miRNAs
Several exosomal miRNAs have been reported to move from MSCs to FLSs to protect against the pathological processes of RA, including miR-320a, miR-150-5p, miR-124a, and miR-140-3p.224,225,226,227 The loading of miR-320a into MSC-derived exosomes is highly effective. After exosomes are taken up by RA-FLSs, the high abundance of exosomal miR-320a substantially restricts the invasive properties of RA-FLSs through interaction of miR-320a with CXCL9.224 MiR-150-5p expression in FLSs of RA patients was found to be decreased compared with that in FLSs of OA patients. MSC-derived exosomes carrying miR-150-5p were obtained after transfection of MSCs with a plasmid encoding miR-150-5p. Exosomal miR-150-5p negatively regulated MMP14 and VEGF, contributing to suppression of the migration and invasion of RA-FLSs and the decreased tube formation by HUVECs.225 Coincubation of exosomes derived from human MSCs with miR-124a overexpression with RA-FLSs resulted in a significant decline in the number and reduced migration of RA-FLSs.226 However, the underlying mechanisms and targets of exosomal miR-124a in the modulation of the aggressive behavior of FLSs in RA were not elucidated in this work. Exosomal miR-140-3p from human umbilical cord MSCs has recently been proven to inhibit RA-FLS growth via SGK1 silencing. Administration of MSC-derived exosomes containing miR-140-3p to a CIA rat model decreased chondrocyte apoptosis and the progression of joint injury.227
MSC-derived exosomes also play an immunomodulatory role in RA pathogenesis. Exosomes derived from miR-146a/miR155-transduced MSCs significantly expand Treg populations, which may ultimately lead to the restoration of appropriate T-cell responses in the chronic inflammatory microenvironment of RA.228
T-cell-derived exosomal miRNAs
T-cell-derived exosomes appear to promote the intercellular transfer of miR-204-5p and miR-132. Wu et al. identified human T cells and FLSs as parental and recipient cells of RA-associated exosomal miR-204-5p, respectively. The delivery of miR-204-5p to RA-FLSs repressed cell activation, proliferation, and invasion as well as inflammatory cytokine production.229 MiR-132 was found to be packaged into exosomes produced by Th17 cells and released into osteoclast precursors to promote their differentiation into mature osteoclasts via COX2 downregulation. Interestingly, cigarette smoke (CS) can act as an important agonist for AHR, and AHR signaling can result in further forced expression of miR-132 in patients with RA. Increased osteoclastogenesis in arthritic mice with or without CS exposure was blocked by local knockdown of miR-132 in joints.188 Unfortunately, despite the high miR-132 expression induced by AHR activation in RA Th17 cells, the authors failed to reveal an intrinsic role of miR-132 in Th17 cell differentiation. In addition to its effect on osteoclastogenesis, Th-17-derived exosomal miR-132 is likely to have promising, albeit unproven, applications for the regulation of arthritis in vivo.
MiRNA-based diagnostic strategies for RA
Early diagnosis and treatment of RA are the key stopping the progression of the disease and reducing the rate of disability. Owing to the insufficient sensitivity and specificity of the early predictive indicators such as RF and ACPA, the diagnosis mainly depends on the integrated consideration of medical history, clinical manifestations, physical examination results, and radiological characteristics. In many cases, by the time a patient is conclusively diagnosed with RA, irreversible joint damage has already occurred, and the optimal time for treatment has been missed. Targeting characteristic miRNA expression profiles in RA may provide breakthroughs for early diagnosis and therapy.
As mentioned above, miRNAs that are formed and function inside cells can be released into circulating body fluids. Whether these miRNAs are retained or released depends on cell-type-specific miRNA sorting motifs.66 Circulating miRNAs of distinct cellular sources are encapsulated in EVs such as exosomes and microvesicles and transferred between cells, thereby establishing intercellular communication.60,66 Likewise, some proteins, such as those in Ago2 complexes and lipoproteins, may serve as vesicle-free carriers that transfer extracellular miRNAs to recipient cells.230,231 Because they are contained within these carriers as cargo, circulating miRNAs are resistant to degradation by endogenous RNases and thus present in body fluids in a stable form, making it relatively simple to noninvasively extract samples for detection.232 Moreover, miRNA expression can be easily detected and accurately quantified by current available techniques, including quantitative reverse transcriptase polymerase chain reaction (qRT‒PCR), in situ hybridization, microarray, and small RNA sequencing. Accordingly, circulating miRNAs are attractive and promising biomarkers for clinical applications in RA, such as aiding early diagnosis, monitoring of the disease course, and prediction of treatment response.
Contribution to prevention and precise diagnosis
Differential expression profiles of serum miRNAs in RA patients compared to healthy individuals were identified through bioinformatics methods in a multicenter study by Luque-Tévar et al. Among the 223 deregulated serum miRNAs that were assessed for enriched biological functions, the authors further identified a set of miRNAs (miR-106a-5p, miR-143-5p, miR-148b-3p, miR-199a-5p, and miR-346) that seem to target critical genes implicated in RA pathogenesis. Serum levels of miR-106a-5p, miR-148b-3p and miR-199a-5p were decreased whereas serum miR-143-5p and miR-346 levels were increased in patients with RA compared to healthy individuals.233 In addition, patients with established rather than early RA exhibited higher expression of serum miR-223 than unaffected controls.234,235 A study by Ouboussad et al. suggested a pronounced increase in the expression levels of serum miR-22 after progression from the preclinical state to early RA. Serum miR-22 is therefore deemed an important predictor of the onset and development of RA.236 Utilizing the FirePlex multiplex assay, Cunningham et al. identified miR-24-3p, miR-126-3p, miR-130a-3p, miR-221-3p, miR-431-3p, and let-7d-5p as potential biomarkers for RA in a fixed panel of miRNAs linked to immune cell dysfunction. The differential expression of these 6 miRNAs in serum distinguishes RA patients and ACPA-positive at-risk individuals afflicted with arthralgia from healthy subjects.237 In addition, circulating miR-103a-3p has been identified as a robust biomarker that can be used to identify populations at risk of imminent RA for timely intervention. Whole-blood miR-103a-3p levels in ACPA-positive, asymptomatic, first-degree relatives were comparable to those in RA patients but increased ~7.6-fold compared with those in healthy subjects.238
The levels of plasma miR-22-3p, miR-23b, miR-24-3p, miR-26a-5p, miR-125a-5p, miR-146a-5p, miR-155, and let-7a-5p were elevated in patients with RA in comparison with healthy controls.239,240,241,242 Among these, the combination of miR-24-3p, miR-26a-5p, and miR-125a-5p showed the greatest diagnostic power for RA.241 The upregulation of miR-155 in the plasma of patients with RA seems to agree with the widely reported high expression of miR-155 in diverse RA-associated cells. Unfortunately, another research group revealed lower plasma miR-155 concentrations in RA patients than in healthy subjects.243 Indeed, due to the relatively low plasma miR-155 levels,241 findings regarding changes in the level of miR-155 in the context of RA have a lower likelihood of being reproduced across studies. MiR-155 expression in serum samples has been shown to be higher in patients with RA than in unaffected controls in at least two independent investigations.235,244 Taha et al. later proposed the conflicting view that serum miR-155 expression does not differ between RA patients and controls.234 Overall, the intrinsic and extrinsic factors affecting circulating miRNA levels have not been fully characterized. The sex, age, race, and lifestyle of the subjects, techniques employed for miRNA analysis, study design and stage of disease may all contribute to the differences in circulating miRNA signatures.
Circulating exosomal miRNAs have emerged as attractive candidate markers for RA diagnosis. Wu et al. performed global miRNA profiling of plasma exosomes in patients with RA and identified 14 exosomal miRNAs with disturbed expression. The authors subsequently highlighted a decrease in exosomal miR-204a-5p levels in RA patients in comparison to healthy subjects. This finding was confirmed in both the replication and validation groups, which had larger numbers of independent samples.229 Likewise, exosome-derived miR-548a-3p and miR-6089, both of which perturb the proliferation and activation of THP-1 macrophages by targeting TLR4, are decreased in serum samples from patients with RA.245,246 Focusing on the unique miRNA profiles in serum exosomes from patients with early-stage untreated RA, Rodriguez-Muguruza et al. developed a biomarker panel comprising serum exo-miR-25-3p, exo-miR-451a, and soluble TNF-like weak inducer of apoptosis (TWEAK), an inflammatory cytokine involved in the pathogenesis of RA, that may exhibit superior diagnostic utility over ACPA positivity.247 More reproducible studies with expanded sample sizes and designs optimized to assess panel specificity are needed in the future. Two important confounding factors affecting the utility of exosomal miRNAs as biomarkers for RA are the diverse approaches used to isolate miRNAs from body fluids and the difficulty in identifying their cellular origin. With more precise and standardized techniques for separating miRNA-containing exosomes and methods for recognizing exosomes based on their cellular origin and surface protein patterns, exosomal miRNAs will hopefully be developed into robust diagnostic tools for RA.
The final remaining challenge for miRNAs to be used as diagnostic biomarkers in clinical practice is that accurate diagnosis also requires discrimination of RA from other inflammatory autoimmune disorders. Substantial upregulation of miR-22-3p, miR-24-3p, miR-96-5p, miR-134-5p, miR-140-3p, and miR-627-5p, independent of disease activity and seropositivity, was noted and validated in plasma samples from RA patients compared to heathy controls. These differentially expressed miRNAs, however, failed to distinguish RA from systemic lupus erythematosus (SLE) due to their shared autoimmune properties.248 Jin et al. discovered that miR-124, miR-448, and miR-551b can serve as reliable diagnostic markers for RA. The expression of these 3 miRNAs in serum samples from patients with RA, however, was comparable to that in patients with SLE, Sjogren’s syndrome, and ulcerative colitis (UC).249 Inspiringly, in the past 5 years, several circulating miRNAs have been proposed as suitable biomarkers for the differential diagnosis of RA from related inflammatory autoimmune diseases. For instance, plasma concentrations of miR-16, miR-19b, miR-23a, miR-27a, miR-92a, and miR-223 were significantly higher in patients with RA than in patients with SLE.250 In a recent study, Cheleschi et al. first demonstrated the remarkable ability of serum miR-140 levels to discriminate RA from psoriatic arthritis (PsA), suggesting miR-140 as a specific biomarker to distinguish RA from PsA.251
Furthermore, it is of practical value to discuss the ability of cell-free miRNAs to differentiate RA from OA. Reduced miR-132 expression was observed in the plasma of patients with RA and patients with OA; however, the change in miR-132 levels was not sufficient to distinguish these two populations.252 When compared with OA patients, RA patients demonstrated considerably higher serum miR-125b levels, whereas serum miR-150-5p expression was dramatically diminished.225,253 Intriguingly, extracellular miRNAs are also stably present in the synovial fluid of patients with arthritis. Synovial fluid miRNAs, primarily derived from synovial tissue and infiltrating cells within the articular cavity, possess distinct expression patterns from peripheral blood miRNAs.252 In patients with RA, the expression of miR-16, miR-132, miR-146a, and miR-223 in synovial fluid was reduced compared to that in plasma,252 whereas synovial fluid miR-146b concentrations were increased compared to those in serum.254 Synovial fluid levels of miR-16, miR-146a, miR-155, miR-221-3p, and miR-223 were found to be upregulated in RA patients compared with OA patients.135,252
Disease activity and prognosis monitoring
Cell-free miRNAs have been suggested to be promising markers of disease activity, given their marked correlation with a number of clinical parameters in RA. Serum miR-155 expression and serum miR-210 expression in RA were upregulated and downregulated, respectively, and these changes were associated with an increase in TNF-α and IL-1β levels.244 A meta-analysis conducted by Bae et al. indicated that forced expression of miR-146a in synovial fluid may result in high erythrocyte sedimentation rates (ESR).255 In patients with RA, sustained plasma miR-23b expression suggested active disease and was accompanied by elevated DAS28, ESR, and C-reactive protein (CRP) levels, regardless of the RF or ACPA level.242 The level of circulating miR-16 in RA was proven to be significantly negatively correlated with the Disease Activity Score for 28 joints (DAS28).252 Notably, the baseline levels of serum miR-16 in patients with early RA, as well as the variations in serum miR-16 levels within the first 3 months following initiation of treatment, appear to have potential in the prediction of disease activity and outcome.235 Ciechomska et al. identified circulating miR-146b as a reliable marker of disease activity with improved specificity. The greater the increase in the concentration of serum miR-146b was, the lower the DAS28 based on CRP (DAS28-CRP) and ESR (DAS28-ESR).254 Moreover, an inverse correlation between the levels of circulating exosomal miR-204-5p and ESR, CRP, and RF but not DAS28 and ACPA was noted recently.229
Patients with a prolonged course of RA are prone to suffer from extra-articular comorbidities, contributing to a bleak prognosis. These comorbidities modify miRNA expression profiles in individuals with RA and are one of the challenges in miRNA biomarker development. The differential expression of serum miR-9-5p in RA patients with peripheral neuropathy suggests that serum miR-9-5p expression can be used as a predictor of RA-associated peripheral neuropathy.256 Compared to those in RA patients without lung involvement, the plasma miR-7-5p and miR-214-5p levels were found to be upregulated in RA patients with interstitial lung disease.257 Using NGS and random forest analysis, Ormseth et al. identified 8 candidate plasma miRNAs that could be used to monitor the progression of coronary artery calcification in patients with RA, which may support the early identification and management of coronary atherosclerosis.258 Furthermore, the serum exosomal miR-155 level is high in RA patients with chronic hepatitis C virus (HCV) infection compared with those without infection.259
Therapeutic response prediction
That miRNAs can be used to predict response to medication offers the opportunity to optimize individualized protocols for more RA patients, which can increase the rate of remission and/or decrease disease activity by allowing patients to switch to alternative therapies. In serum or PBMCs from patients with RA after 3 months of conventional disease-modifying anti-rheumatic drug (cDMARD) treatment, higher baseline levels of miR-15a, miR-16, miR-125b, and miR-223 are associated with a better therapeutic response (DAS28 < 3.2).110,235,260 In contrast, high pretherapy concentrations of miR-132-3p, miR-146a-5p, and miR-155-5p in whole blood of RA patients are indicators of inadequate response (DAS28 ≥ 3.2) to MTX therapy.261
Biological DMARDs, as the preferred alternative treatment for patients who fail cDMARD therapy, have been confirmed to modify the circulating miRNA expression profile in RA.233 In patients with RA who responded well to TNF inhibitor (TNFi)/cDMARD combination treatment, serum miR-23a-3p and miR-223-3p expression was decreased at baseline and significantly increased after 3 months of treatment.262 The circulating miR-22, miR-27-3p and miR-886-3p concentrations in patients with RA prior to the initiation of adalimumab/MTX combination therapy are pivotal predictors of a good clinical response. Specifically, RA patients with high plasma miR-27a-3p expression or a combination of decreased serum miR-22 levels and increased miR-886-3p levels are more likely to achieve clinical remission.263,264 Notably, adalimumab/MTX combination therapy has been demonstrated to suppress miR-27a-3p expression in the first 3 months after initiation, whereas in nonresponders, an increase rather than a decrease in miR-27a-3p levels has been observed. In addition, reduced miR-5196 expression and elevated circulating miR-146 levels are considered potential markers of good response to TNFi therapy.262,265,266
Treatment-naïve patients with RA who have high serum miR-125b expression levels tend to have disease that can be adequately controlled by rituximab therapy.253 Importantly, considering that increased HCV activity is associated with rituximab response, serum exosomal miR-155, as a suppressor of HCV replication in hepatocytes, may be applied to monitor the HCV load in RA patients during the duration of rituximab therapy.259
Serum miR-432-5p concentrations were decreased in patients with RA in remission but surprisingly upregulated in nonremission patients and patients who relapsed following 5 years of tofacitinib therapy in a trial.267 Nasonov et al. recently published the first randomized controlled phase III trial of olokizumab (OKZ), a monoclonal antibody targeting IL-6, in RA, in which they evaluated the efficacy and safety of OKZ in RA patients who failed MTX monotherapy.268 Furthermore, baseline plasma levels of miR-26b, miR-29, miR-451, and miR-522 in RA patients appear to be predictors of response to OKZ therapy, although the findings were derived from a Russian population.269
MiRNA-based therapeutic strategies for RA
Modern therapies for RA aim to achieve remission or at least low disease activity within 6 months, which is fulfilled by a treat-to-target strategy that involves rigorous disease control and individualized management.270 To date, the first-line medication for RA patients has been cDMARDs, primarily MTX, supplemented with glucocorticoids and/or nonsteroidal anti-inflammatory drugs. The emergence of novel DMARDs in the past 2 decades, including small molecule inhibitors and biologic agents, has provided additional opportunities for improving the course and prognosis of this disease.7,14 Detailed classifications and descriptions of modern pharmacologic therapies for RA and key cells and receptors/pathways targeted by these strategies have been thoroughly reviewed elsewhere.7,271 Nevertheless, current treatment strategies sometimes produce only partial responses accompanied by inevitable side effects or require long-term use due to the high likelihood of relapse following therapy discontinuation.270 Considering the challenge of re-establishing immune homeostasis and remodeling after joint destruction, achieving sustained remission or retaining low disease activity remains elusive. Therefore, the continued development of safe and efficacious new RA therapeutic agents with the aim of preventing disease progression and improving patient quality of life is warranted.
Modulation of miRNAs by modern RA pharmacologic therapies
As RA is a multifactorial and stage-specific inflammatory disease, the onset, progression, and persistence of RA inevitably require the involvement of a range of cells and cytokines.271 Focusing on the effects of modern pharmacologic therapies on miRNAs in RA will help further elucidate definitive molecular or cellular nodes in complex miRNA regulatory networks that subserve disease and aid the development of novel therapeutic strategies.
MTX is widely used as an anchor drug for RA treatment. Intriguingly, administration of MTX alters the global expression profile of miRNAs in RA-FLSs, resulting in sustained expression of miR-877-3p. Dysregulation of miR-877-3p expression limited the growth and migration of RA-FLSs by significantly reducing GM-CSF and CCL3 transcript levels.272
Upregulation of miR-29b in RA may be associated with TNF-α induction. Repressed miR-29 expression in RA patients treated with infliximab contributes to enhanced resistance of PBMCs to apoptosis by inactivating HBP1 signaling.273 Administration of the TNFi adalimumab to patients with RA downregulated miR-155 levels in macrophages and consequently reversed the defect in M2 polarization.115 Low global miRNA levels in RA neutrophils were observed, which may have been mediated by the release of excessive levels of inflammatory cytokines, such as TNF-α and IL-6, and/or a defect in miRNA processing as a result of DICER inadequacy. Treatment with the anti-TNF agent infliximab and the anti-IL-6 agent tocilizumab dramatically reversed the global downregulation of miRNAs in RA neutrophils, improved miRNA processing and decreased the production of various cytokines and chemokines, which blocked the activation of RA neutrophils.177 Moreover, tocilizumab was found to upregulate serum miR-146a-5p levels in treated RA patients. Conversely, treatment of an in vitro coculture system including monocytes and fibroblasts with tocilizumab downregulated miR-146a-5p expression.274 One explanation is that the serum may reflect the state of synovial joints, where interactions between multiple cell types may produce different results than those derived from in vitro cell lines. Despite these inconsistencies, tocilizumab treatment has been demonstrated to regulate miR-146a-5p expression, which is partially involved in the restriction of angiogenic potential in RA patients.274
Abatacept is a T-cell costimulation modulator consisting of an extracellular domain of human cytotoxic T-lymphocyte-associated antigen 4 (CTLA-4) and a modified Fc portion of human IgG1.271 According to Hayakawa et al., plasma miR-766-3p levels were lower in abatacept-treated RA patients than in untreated patients. The authors focused on not only the possibility of applying plasma miR-766-3p as a biomarker for RA but also its function in different cell lines in suppressing the inflammatory response through indirect NF-κB signaling inactivation.275
MiRNA-based gene therapies in RA
Overall findings regarding the involvement of miRNAs in mediating pathogenic processes in RA suggest enormous potential for their therapeutic manipulation. Although a considerable number of miRNAs differentially expressed in RA have been discovered over the years, and some corresponding preclinical studies have been conducted, the development of miRNA therapeutics is in its infancy. It is still difficult to translate these preliminary investigations into clinical practice for RA therapy. Bare miRNAs in body fluids are bound to be rapidly degraded by RNases, which prevents them from reaching the target sites to perform functions. In addition to having poor biostability, due to their large molecular size and strong anionic charge, miRNAs are unable to pass through the plasma membrane, and therapeutic miRNAs are prone to off-target effects.276 A specific miRNA may exert diverse effects in RA pathogenesis, depending on the cell type in which it is expressed. Certain miRNAs are protective against RA, regardless of their expression in clinical samples. Hence, there are two main challenges in developing miRNA-based therapies for RA: (i) developing a proper and comprehensive understanding of miRNA function, which will facilitate the selection of miRNA candidates with maximum target efficacy and knowledge of whether to inhibit or increase their endogenous expression; and (ii) rationally designing optimal miRNA delivery vehicles that allow tissue- or cell-specific targeting, which will lead to outstanding biostability and cellular uptake as well as minimal toxicity and low probability of off-target effects of miRNA candidates.
MiRNA inhibition and replacement therapy
The central action of miRNAs is to restrain gene expression by binding and silencing specific mRNAs, which in turn modulates cellular function. Accordingly, miRNA-based gene therapies aim to artificially interfere with target miRNA activity in RA, managing RA-related gene expression and protein synthesis via the silencing of overexpressed pathogenic miRNAs and increasing the expression of underexpressed beneficial miRNAs.
Currently, clinical studies of miRNA-based therapeutic strategies mainly use inhibitors and mimics of miRNAs. MiRNA inhibitors are synthetic single-stranded RNA molecules consisting of nt complementary to endogenous miRNAs, whereas miRNA mimics are synthetic double-stranded RNAs that bypass the synthesis process of endogenous mature double-stranded miRNAs and achieve high expression of specific miRNAs in a short duration. Synthetic double-stranded miRNAs and antisense oligonucleotides are frequently used for in vitro functional assays of miRNAs, but their chemical modification is required to increase affinity, biostability and tissue and cellular uptake when they are applied in vivo to modulate deregulated miRNAs.277 In most cases, specifically chemically modified miRNA agomirs and antagomirs are recommended for use in validation in in vivo animal experiments. Injection of chemically modified miR-708-5p mimics into the tail vein of rats with CIA ameliorated arthritis scores and ankle histopathology by blocking Wnt3a/β-catenin signaling activation.278 In contrast, intravenous administration of an miR-145-5p agomir resulted in reduced OPG expression in ankle joints and a consequent increase in the severity of joint structural destruction in mice with CIA.192 Moreover, CIA mice injected intravenously with an miR-34a antagomir exhibited improved arthritis phenotypes and symptoms, accompanied by a restored Treg proportion.165
Systemic administration of miRNAs tends to result in unsatisfactory biodistribution and short-term serum miRNA levels owing to preferential accumulation in the liver and rapid renal clearance.277 Thus, local administration seems to be more appropriate for delivering miRNAs in RA to achieve sustained therapeutic concentrations in the inflamed joints, avoiding repeated injections or a single administration of high doses. AIA mice injected intra-articularly with an miR-132 antagomir demonstrated a significant reduction in the number of periarticular osteoclasts.188 Consistent with the in vitro identification of the miR-141-3p/FoxC1/β-catenin axis regulating the pathological behaviors of RA-FLSs, manipulation of miR-141-3p overexpression in the joint cavity suppressed the expression of FoxC1 and β-catenin in the CIA rat model. Synovial hyperplasia, inflammation, and cartilage degradation, as expected, were inhibited in CIA rats treated with miR-141-3p agomir.99 Following injection of pre-miR-124 into a single ankle joint of rats with AIA, apart from accumulation at the injection site, mature miR-124 was also distributed to distant joints, leading to alleviated arthritic tissue destruction in multiple joints.190
Another approach of miRNA replacement and inhibition therapy in RA is to use lentiviruses as vectors for in vivo gene transfer. Lentivirus vector systems are ideally suited to mediate miRNA interference in RA due to their wide range of transduced tissues and efficient delivery of miRNAs. Wu et al. demonstrated that intra-articular administration of an miR-204-5p-overexpressing lentivirus could dramatically mitigate the development of CIA.229 Intra-articular injection of a pre-miR-140-containing lentivirus ameliorated synovial inflammation and pannus formation, which was in part due to the high abundance of FLSs mediated by miR-140 in the synovium of arthritis mice.279 In one of our previous studies, we delivered miR-106b inhibitors into mice with established CIA via orbital injection of lentiviral vectors to neutralize endogenous miR-106b expression in inflamed joints. Lentivirus-mediated miR-106b diminution decreased the ratio of RANKL to OPG and interfered with inflammatory cytokine production at the local and systemic levels.183 Building on the recognition of miR-106b as a putative target for RA therapy, other researchers have further concluded that miR-106b inhibition in the CIA mouse model contributes to the rebalancing of the RANKL/RANK/OPG system and restoration of cartilage degradation, which is the result of PDK4 upregulation.222 However, some concerns with lentivirus vector-mediated miRNA interference have also been uncovered, including off-target effects, innate immune responses, and alterations in endogenous miRNA pathways. Future research efforts will focus on finding more feasible solutions to mitigate these potential risks and enable the clinical application of this technology.
MSC-derived exosomes as carriers for miRNAs
The endogenous properties of exosomes enable them to be natural delivery vehicles for miRNAs and improve cell-free therapeutic strategies for RA.280,281 Among the various cell-derived exosomes, those of MSC origin exert excellent immunomodulatory effects and thus are likely to be the most ideal agents for the delivery of therapeutic miRNAs in autoimmune diseases.
CIA rats treated with bone marrow-derived mesenchymal stem cell (BMSC)-derived exosomes carrying miR-192-5p showed a delayed inflammatory response and a decrease in osteoclast numbers.282 Exosomal miR-21 secreted from BMSCs was found to be incorporated into mouse FLSs, inhibiting inflammatory cytokine generation via the miR-21/TET1/KLF4 axis. BMSC-derived exosomal miR-223 was also found to exert anti-inflammatory activity in macrophages by restraining activation of the NLRP3 inflammasome. Administration of exosomal miR-21 in mice with CIA apparently attenuated inflammatory infiltration in synovial tissues, whereas the in vivo efficacy of exosomal miR-223 has not been discussed.283,284 It has been proven that MSC-secreted exosomes can transfer miR-320a and miR-150 into RA-FLSs, thereby altering their pathological properties. Tail vein injection of MSC-derived exosomes containing miR-320a into CIA mice alleviated arthritis severity, which was correlated with suppressed CXCL9 levels.224 MSC-derived exosomal miR-150 interfered with the migration, invasion, and tube formation of RA-FLSs through a mechanism involving MMP14 and VEGF. Thus, intraperitoneal administration of exosomal miR-150 in mice with CIA led to the amelioration of synovial hyperplasia and angiogenesis.225 Recently, Huang et al. reported that injection of miR-140-3p packaged in human umbilical cord mesenchymal stem cell-derived exosomes into CIA rats had therapeutic effects, including reducing the inflammatory response, oxidative stress, synoviocyte proliferation and cartilage damage.227
Notably, exosomal surfaces can be finely edited by bioengineering to augment their stability, bioactivity and efficiency in targeting specific cells and tissues. Based on metabolic glycoengineering-mediated click chemistry, You et al. developed dextran sulfate-decorated exosomes (DS-EXOs) secreted from surface-modified adipose-derived stem cells to target macrophages in RA. Let-7b-5p and miR-24-3p encapsulated in the engineered DS-EXOs were internalized by activated macrophages and drove M1 macrophages to polarize toward the M2 phenotype. Intravenous injection of DS-EXOs into CIA mice at a lower dose than native exosomes resulted in increased accumulation in the inflamed joints and consequent relief of arthritis symptoms, ultimately decelerating CIA progression.285
Despite these preclinical studies showing the therapeutic potential of MSC-derived exosomes in RA, there are still some gaps regarding safety and efficacy that limit their clinical application.286 The underlying transport and cellular mechanisms that underly the observed clinical benefits of exosomal miRNAs require in-depth characterization. The current technologies of exosome isolation and purification are relatively immature. Furthermore, the optimal administration regimen for exosomes, including route and dose, is unknown. The poor in vivo biodistribution of MSC-derived exosomes after systemic administration, which affects the efficacy of practical applications, needs to be addressed.
Synthetic materials as miRNA delivery vehicles
Although liposomes are widely used as in vitro transfection reagents due to their low immunogenicity, their in vivo application is often restricted by toxicity linked to compound composition and structure.287 Encouragingly, several cationic lipid-based vectors have been successfully utilized to improve the in vivo delivery of therapeutic miRNAs in murine models of RA.
Injection of lipoplexes encapsulating miR-17 mimics into the ankle joints protected CIA mice from synovitis, cartilage degradation and bone erosion. Forced miR-17 expression markedly inhibited pro-inflammatory cytokine (for example, IL-6) secretion, synovial immune cell infiltration, and osteoclast formation by blocking JAK1/STAT3 signaling.93 Sujitha et al. employed poly-ethylene glycol (PEG)-grafted polyethylenimine (PEI) to deliver miR-23a, a functional miRNA associated with Wnt1/β-catenin signaling modulation. MiR-23a incorporated within PEGylated PEI polyplexes was systemically administered to AIA rats, in which it achieved multifaceted benefits in terms of inducing disease remission.288 A previous study validated the role of miR-124 in impeding osteoclast differentiation via direct interaction with NFATc1 mRNA.190 To enable MTX and miR-124 combination therapy, methotrexate-conjugated polymer hybrid micelles (M-PHMs) with efficient cellular uptake, optimized transfection efficiency and decreased cytotoxicity were developed. M-PHMs were coloaded with miR-124 to target activated macrophages in inflamed joints. The application of M-PHM/miR-124 complexes led to significant alleviation of arthritis symptoms through anti-inflammatory and anti-bone erosion effects in the AIA rat model.289
Of note, lipoplexes containing miR-146a mimics formulated with DMAPAP/DOPE cationic liposomes efficiently delivered miR-146a to Ly6Chigh monocytes and diminished inflammation-induced local bone loss; however, these effects failed to ameliorate the overwhelming inflammation in both CIA and STA mouse models.129 Likewise, CIA mice injected intravenously with double-stranded miR-146a/atelocollagen complexes were resistant to bone erosion but not synovitis.123 These results are consistent with the overexpression of miR-146a noted in various cells and tissues of RA, and miR-146a seems to lose its normal function as a sensor to orchestrate the inflammatory response.
MiRNA-loaded nanoparticles (NPs) are another well-tested carrier for miRNAs. Poly(lactide-co-glycolide) (PLGA) NPs and poly (cyclohexane-1,4-diylacetone dimethylene ketal) (PCADK) NPs have been applied to codeliver miR-124 and ketoprofen into rats with AIA. When administered systemically, miR-124 and ketoprofen encapsulated in these two delivery systems effectively decelerated the progress of AIA. Compared to PLGA NPs, acid-responsive PCADK NPs achieved accelerated release of payloads in response to the slightly acidic microenvironment of inflamed joints, suggesting superior therapeutic efficacy.290,291
Based on evidence indicating overexpression of miR-30-5p in synovial tissue samples from patients with RA, Zhang et al. fabricated cell membrane-penetrating peptide (CADY)-modified mesoporous silica nanoparticle (MSN)/miR-30-5p inhibitor complexes for targeted delivery into RA-FLSs. The reported systemic delivery strategy using these miR-30-5p inhibitor-containing complexes in coordination with a near-infrared response MSN-based vehicle that governs triptolide (TP) release is promising for RA treatment.292
Moreover, after repeated systemic administration in zymosan A-induced arthritis (ZIA) mice, passive accumulation of the formulated nanocomplexes (NCs) loaded with miR-21 and IL-4 in inflamed joints was observed owing to their prolonged serum half-life (t1/2). MiR-21 cooperates with IL-4 to orchestrate the composition of the osteoimmune microenvironment in ZIA, eliciting pronounced anti-inflammatory and tissue repair outcomes via NF-κB signaling inactivation and M2 polarization promotion. Intriguingly, the NC-mediated codelivery of miR-21 and IL-4 to different target sites was carried out in a hierarchical manner under spatiotemporal control. Specifically, instructed by the micro-acidic inflammatory surroundings, the poly-l-lysine-cis-aconitic anhydride (PLL-CA) in the outer layer of the NCs undergoes charge reversal. The switch of PLL-CA from negative to positive first leads to the liberation of surface-adsorbed IL-4 to the extracellular compartments of the inflamed synovium, followed by exposure of the cationic membrane-penetrating, helical polypeptide (PG)/miR-21 inner core, which allows intracellular transport into macrophages.293
Conclusions and perspectives
The recognition of miRNAs as fine-tuners of gene expression and the abundance of intensive relevant research have updated our fundamental understanding of RA pathogenesis. A plethora of deregulated miRNAs are implicated in regulating the differentiation, development, homeostasis and function of multiple cell types in different tissues and developmental stages, thereby contributing to tissue inflammation and damage in RA. The miRNA-mediated crosstalk between diverse cell lines in RA also represents an important part of the disease process. Notably, the complicated interactions and integration of multilayer signaling between miRNAs and other epigenetic factors are indispensable for RA pathogenesis but remain largely unknown. Many mechanistic studies concerning miRNA–mRNA interactions have identified a growing list of miRNA target genes in FLSs, immune cells and bone-related cells, allowing a glimpse of the potential functional scope of these miRNAs in inflammatory cascades, the immune response and bone homeostasis. Nevertheless, there is currently a lack of sufficient in vivo experimental evidence to address the pleiotropic role of miRNAs in RA disease states. More studies should be conducted in animal models of inducible arthritis, preferably utilizing conditional gene knockout/knock-in strategies, to further deduce the function of specific miRNAs in distinct cell subsets of RA. Furthermore, the initiation, development and persistence of RA rely on the synergistic actions of miRNAs, which constitute a complex network of intracellular signaling modulation. Thus, although the capacity of a single miRNA to target a broad range of genes simultaneously is exciting and being extensively researched, the combinatorial effects of global miRNAs on gene expression as well as pathological phenotype in RA warrant further exploration.
Accurate early diagnosis and feasible new drug therapies that facilitate the achievement of remission are urgently needed for patients with RA. Circulating miRNAs have been proposed as novel biomarkers for RA. Given that the field is still in its infancy, more large-scale cohort studies are required to verify the superiority and clinical utility of miRNAs as diagnostic, predictive and prognostic markers in RA.
In addition, a profound understanding of miRNA deregulation contributes to the clinical development of miRNA-based therapies for RA. The ability of miRNAs to target multiple genes and pathways renders them promising therapeutic targets. Unfortunately, this feature has also conferred significant risk of off-target effects and unpredictable side effects owing to the systemic delivery of a particular miRNA in the majority of preclinical studies. In some cases, simultaneous regulation of the expression and function of multiple miRNAs might lead to better clinical outcomes in RA patients. However, the widespread effects of miRNAs on several targets in different tissues is potentially capable of shutting down entire pathways. The remaining questions regarding miRNA tissue distribution and systemic toxicity imply that before using miRNAs in clinical practice, substantial work is still required to comprehensively appraise the therapeutic potential of miRNAs and to obtain optimized protocols. The design of safe and effective delivery systems that enable miRNA aggregation in inflamed joints and specific target cell types in RA remains the key to bringing miRNAs into the clinic. MiRNA-mediated gene regulation is highly sensitive to the cell type and context. Strategies targeting monocytes and MSC-based strategies appear to be ideal therapeutic approaches since these cells serve as the main source of pathogenic cells in the development of RA. Exosomal miRNAs driving intercellular communication have shown substantial potential as candidates for RA treatment. In addition to linking miRNAs to RA therapy from a biological perspective, encouraging results have been achieved using nanomaterial-based engineering strategies for the local delivery of miRNAs into joint tissues.
In conclusion, comprehensive insights into the pathogenesis of RA gained from a multidisciplinary approach will continue to accelerate the clinical utilization of miRNAs as novel biomarkers and therapeutic targets, thus moving toward a new era of individualized management for patients with RA.
References
Crowson, C. S. et al. Impact of risk factors associated with cardiovascular outcomes in patients with rheumatoid arthritis. Ann. Rheum. Dis. 77, 48–54 (2018).
Safiri, S. et al. Global, regional and national burden of rheumatoid arthritis 1990–2017: a systematic analysis of the Global Burden of Disease study 2017. Ann. Rheum. Dis. 78, 1463–1471 (2019).
Fu, J. et al. Shared epitope-aryl hydrocarbon receptor crosstalk underlies the mechanism of gene-environment interaction in autoimmune arthritis. Proc. Natl. Acad. Sci. USA 115, 4755–4760 (2018).
Ospelt, C. et al. Epigenetics in the pathogenesis of RA. Semin Immunopathol. 39, 409–419 (2017).
McInnes, I. B. & Schett, G. The pathogenesis of rheumatoid arthritis. N. Engl. J. Med. 365, 2205–2219 (2011).
Scott, D. L. et al. Rheumatoid arthritis. Lancet 376, 1094–1108 (2010).
McInnes, I. B. & Schett, G. Pathogenetic insights from the treatment of rheumatoid arthritis. Lancet 389, 2328–2337 (2017).
Lee, R. C. et al. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993).
Bushati, N. & Cohen, S. M. microRNA functions. Annu Rev. Cell Dev. Biol. 23, 175–205 (2007).
Lee, Y. S. & Dutta, A. MicroRNAs in cancer. Annu Rev. Pathol. 4, 199–227 (2009).
Zhou, S. S. et al. miRNAS in cardiovascular diseases: potential biomarkers, therapeutic targets and challenges. Acta Pharm. Sin. 39, 1073–1084 (2018).
Ali, S. A. et al. The non-coding RNA interactome in joint health and disease. Nat. Rev. Rheumatol. 17, 692–705 (2021).
Dakin, S. G. et al. Pathogenic stromal cells as therapeutic targets in joint inflammation. Nat. Rev. Rheumatol. 14, 714–726 (2018).
Aletaha, D. & Smolen, J. S. Diagnosis and management of rheumatoid arthritis: a review. JAMA 320, 1360–1372 (2018).
Bottini, N. & Firestein, G. S. Duality of fibroblast-like synoviocytes in RA: passive responders and imprinted aggressors. Nat. Rev. Rheumatol. 9, 24–33 (2012).
Mousavi, M. J. et al. Transformation of fibroblast-like synoviocytes in rheumatoid arthritis; from a friend to foe. Auto. Immun. Highlights 12, 3 (2021).
Firestein, G. S. et al. Somatic mutations in the p53 tumor suppressor gene in rheumatoid arthritis synovium. Proc. Natl. Acad. Sci. USA 94, 10895–10900 (1997).
Tu, J. et al. A Tale of Two Immune Cells in Rheumatoid Arthritis: The Crosstalk Between Macrophages and T Cells in the Synovium. Front. Immunol. 12, 655477 (2021).
Kondo, Y. et al. Review: Transcriptional Regulation of CD4+ T Cell Differentiation in Experimentally Induced Arthritis and Rheumatoid Arthritis. Arthritis Rheumatol. 70, 653–661 (2018).
Niu, Q. et al. Disturbed Th17/Treg balance in patients with rheumatoid arthritis. Rheumatol. Int. 32, 2731–2736 (2012).
Wehrens, E. J. et al. T cells out of control-impaired immune regulation in the inflamed joint. Nat. Rev. Rheumatol. 9, 34–42 (2013).
Wu, F. et al. B Cells in Rheumatoid ArthritisPathogenic Mechanisms and Treatment Prospects. Front. Immunol. 12, 750753 (2021).
Ronnelid, J. et al. Autoantibodies in Rheumatoid Arthritis - Laboratory and Clinical Perspectives. Front. Immunol. 12, 685312 (2021).
Kennedy, A. et al. Macrophages in synovial inflammation. Front. Immunol. 2, 52 (2011).
Hannemann, N. et al. New insights into macrophage heterogeneity in rheumatoid arthritis. Jt. Bone Spine 88, 105091 (2021).
Hashimoto, D. et al. Tissue-resident macrophages self-maintain locally throughout adult life with minimal contribution from circulating monocytes. Immunity 38, 792–804 (2013).
Lavin, Y. et al. Regulation of macrophage development and function in peripheral tissues. Nat. Rev. Immunol. 15, 731–744 (2015).
Huang, Q.-Q. et al. Critical role of synovial tissue-resident macrophage niche in joint homeostasis and suppression of chronic inflammation. Sci. Adv. 7, eabd0515 (2021).
Das, A. et al. Monocyte and macrophage plasticity in tissue repair and regeneration. Am. J. Pathol. 185, 2596–2606 (2015).
Funes, S. C. et al. Implications of macrophage polarization in autoimmunity. Immunology 154, 186–195 (2018).
Wang, Y. et al. Is macrophage polarization important in rheumatoid arthritis? Int. Immunopharmacol. 50, 345–352 (2017).
Cascao, R. et al. Neutrophils in rheumatoid arthritis: More than simple final effectors. Autoimmun. Rev. 9, 531–535 (2010).
Iberg, C. A. et al. Dendritic cells as inducers of peripheral tolerance. Trends Immunol. 38, 793–804 (2017).
Iwasaki, A. & Medzhitov, R. Control of adaptive immunity by the innate immune system. Nat. Immunol. 16, 343–353 (2015).
Schett, G. & Gravallese, E. Bone erosion in rheumatoid arthritis: mechanisms, diagnosis and treatment. Nat. Rev. Rheumatol. 8, 656–664 (2012).
Asagiri, M. & Takayanagi, H. The molecular understanding of osteoclast differentiation. Bone 40, 251–264 (2007).
Wada, T. et al. RANKL-RANK signaling in osteoclastogenesis and bone disease. Trends Mol. Med. 12, 17–25 (2006).
Amarasekara, D. S. et al. Regulation of Osteoclast Differentiation by Cytokine Networks. Immune Netw. 18, e8 (2018).
Hashizume, M. et al. IL-6 trans-signalling directly induces RANKL on fibroblast-like synovial cells and is involved in RANKL induction by TNF-alpha and IL-17. Rheumatol. (Oxf.) 47, 1635–1640 (2008).
Baum, R. & Gravallese, E. M. Bone as a target organ in rheumatic disease: impact on osteoclasts and osteoblasts. Clin. Rev. Allergy Immunol. 51, 1–15 (2016).
Otero, M. & Goldring, M. B. Cells of the synovium in rheumatoid arthritis. Chondrocytes. Arthritis Res. Ther. 9, 220 (2007).
Li, Z. et al. Omentin-1 prevents cartilage matrix destruction by regulating matrix metalloproteinases. Biomed. Pharmacother. 92, 265–269 (2017).
Goldring, M. B. & Marcu, K. B. Cartilage homeostasis in health and rheumatic diseases. Arthritis Res. Ther. 11, 224 (2009).
Falconer, J. et al. Review: synovial cell metabolism and chronic inflammation in rheumatoid arthritis. Arthritis Rheumatol. 70, 984–999 (2018).
Elshabrawy, H. A. et al. The pathogenic role of angiogenesis in rheumatoid arthritis. Angiogenesis 18, 433–448 (2015).
Laurent, L. et al. Fcgamma receptor profile of monocytes and macrophages from rheumatoid arthritis patients and their response to immune complexes formed with autoantibodies to citrullinated proteins. Ann. Rheum. Dis. 70, 1052–1059 (2011).
Negishi-Koga, T. et al. Immune complexes regulate bone metabolism through FcRgamma signalling. Nat. Commun. 6, 6637 (2015).
Bartok, B. & Firestein, G. S. Fibroblast-like synoviocytes: key effector cells in rheumatoid arthritis. Immunol. Rev. 233, 233–255 (2010).
Bottini, N. & Firestein, G. S. Duality of fibroblast-like synoviocytes in RA: passive responders and imprinted aggressors. Nat. Rev. Rheumatol. 9, 24–33 (2013).
Ha, M. & Kim, V. N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524 (2014).
Chen, J. F. et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat. Genet. 38, 228–233 (2006).
Di Leva, G. et al. MicroRNAs in cancer. Annu. Rev. Pathol. 9, 287–314 (2014).
Krol, J. et al. The widespread regulation of microRNA biogenesis, function and decay. Nat. Rev. Genet. 11, 597–610 (2010).
Han, J. et al. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027 (2004).
Diederichs, S. & Haber, D. A. Dual role for argonautes in microRNA processing and posttranscriptional regulation of microRNA expression. Cell 131, 1097–1108 (2007).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Wilson, R. C. & Doudna, J. A. Molecular mechanisms of RNA interference. Annu. Rev. Biophys. 42, 217–239 (2013).
Guo, H. et al. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466, 835–840 (2010).
Friedman, R. C. et al. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Alghamdi, M. et al. Circulating extracellular vesicles and rheumatoid arthritis: a proteomic analysis. Cell Mol. Life Sci. 79, 25 (2021).
Li, D. et al. Osteoclast-derived exosomal miR-214-3p inhibits osteoblastic bone formation. Nat. Commun. 7, 10872 (2016).
van Niel, G. et al. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19, 213–228 (2018).
Zhu, S. et al. Coupling factors and exosomal packaging microRNAs involved in the regulation of bone remodelling. Biol. Rev. Camb. Philos. Soc. 93, 469–480 (2018).
Ji, C. & Guo, X. The clinical potential of circulating microRNAs in obesity. Nat. Rev. Endocrinol. 15, 731–743 (2019).
Kowal, J. et al. Biogenesis and secretion of exosomes. Curr. Opin. Cell Biol. 29, 116–125 (2014).
Garcia-Martin, R. et al. MicroRNA sequence codes for small extracellular vesicle release and cellular retention. Nature 601, 446–451 (2022).
Villarroya-Beltri, C. et al. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat. Commun. 4, 2980 (2013).
Santangelo, L. et al. The RNA-Binding Protein SYNCRIP Is a Component of the Hepatocyte Exosomal Machinery Controlling MicroRNA Sorting. Cell Rep. 17, 799–808 (2016).
Hobor, F. et al. A cryptic RNA-binding domain mediates Syncrip recognition and exosomal partitioning of miRNA targets. Nat. Commun. 9, 831 (2018).
Teng, Y. et al. MVP-mediated exosomal sorting of miR-193a promotes colon cancer progression. Nat. Commun. 8, 14448 (2017).
Mukherjee, K. et al. Reversible HuR-microRNA binding controls extracellular export of miR-122 and augments stress response. EMBO Rep. 17, 1184–1203 (2016).
Shurtleff, M. J. et al. Y-box protein 1 is required to sort microRNAs into exosomes in cells and in a cell-free reaction. Elife 5, e19276 (2016).
Liu, X. M. et al. Selective sorting of microRNAs into exosomes by phase-separated YBX1 condensates. Elife 10, e71982 (2021).
Koppers-Lalic, D. et al. Nontemplated nucleotide additions distinguish the small RNA composition in cells from exosomes. Cell Rep. 8, 1649–1658 (2014).
Gibbings, D. J. et al. Multivesicular bodies associate with components of miRNA effector complexes and modulate miRNA activity. Nat. Cell Biol. 11, 1143–1149 (2009).
McKenzie, A. J. et al. KRAS-MEK signaling controls Ago2 sorting into Exosomes. Cell Rep. 15, 978–987 (2016).
Guduric-Fuchs, J. et al. Selective extracellular vesicle-mediated export of an overlapping set of microRNAs from multiple cell types. BMC Genom. 13, 357 (2012).
Git, A. et al. Systematic comparison of microarray profiling, real-time PCR, and next-generation sequencing technologies for measuring differential microRNA expression. RNA 16, 991–1006 (2010).
Correia de Sousa M. et al. Deciphering miRNAs’ Action through miRNA Editing. Int. J. Mol. Sci. 20, 6249 (2019).
Stanczyk, J. et al. Altered expression of MicroRNA in synovial fibroblasts and synovial tissue in rheumatoid arthritis. Arthritis Rheum. 58, 1001–1009 (2008).
Xie, Z. et al. PU.1 attenuates TNFalphainduced proliferation and cytokine release of rheumatoid arthritis fibroblastlike synoviocytes by regulating miR155 activity. Mol. Med. Rep. 17, 8349–8356 (2018).
Liu, N. et al. Paeonol protects against TNF-alpha-induced proliferation and cytokine release of rheumatoid arthritis fibroblast-like synoviocytes by upregulating FOXO3 through inhibition of miR-155 expression. Inflamm. Res. 66, 603–610 (2017).
Long, L. et al. Upregulated microRNA-155 expression in peripheral blood mononuclear cells and fibroblast-like synoviocytes in rheumatoid arthritis. Clin. Dev. Immunol. 2013, 296139 (2013).
Saferding, V. et al. MicroRNA-146a governs fibroblast activation and joint pathology in arthritis. J. Autoimmun. 82, 74–84 (2017).
Sun, W. et al. Resolvin D1 suppresses pannus formation via decreasing connective tissue growth factor caused by upregulation of miRNA-146a-5p in rheumatoid arthritis. Arthritis Res. Ther. 22, 61 (2020).
Hong, B. K. et al. MicroRNA-143 and -145 modulate the phenotype of synovial fibroblasts in rheumatoid arthritis. Exp. Mol. Med. 49, e363 (2017).
Wang, X. et al. Elevated microRNA1455p increases matrix metalloproteinase9 by activating the nuclear factorkappaB pathway in rheumatoid arthritis. Mol. Med. Rep. 20, 2703–2711 (2019).
Huang, Z. et al. MiR-26a-5p enhances cells proliferation, invasion, and apoptosis resistance of fibroblast-like synoviocytes in rheumatoid arthritis by regulating PTEN/PI3K/AKT pathway. Biosci. Rep. 39, BSR20182192 (2019).
Chen, Y. et al. MicroRNA-21 Promotes Proliferation of Fibroblast-Like Synoviocytes through Mediation of NF-kappaB Nuclear Translocation in a Rat Model of Collagen-Induced Rheumatoid Arthritis. Biomed. Res. Int. 2016, 9279078 (2016).
Stanczyk, J. et al. Altered expression of microRNA-203 in rheumatoid arthritis synovial fibroblasts and its role in fibroblast activation. Arthritis Rheum. 63, 373–381 (2011).
Pandis, I. et al. Identification of microRNA-221/222 and microRNA-323-3p association with rheumatoid arthritis via predictions using the human tumour necrosis factor transgenic mouse model. Ann. Rheum. Dis. 71, 1716–1723 (2012).
Alsaleh, G. et al. Bruton’s tyrosine kinase is involved in miR-346-related regulation of IL-18 release by lipopolysaccharide-activated rheumatoid fibroblast-like synoviocytes. J. Immunol. 182, 5088–5097 (2009).
Najm, A. et al. MicroRNA-17-5p reduces inflammation and bone erosions in mice with collagen-induced arthritis and directly targets the JAK/STAT pathway in rheumatoid arthritis fibroblast-like synoviocytes. Arthritis Rheumatol. 72, 2030–2039 (2020).
Lin, J. et al. A novel p53/microRNA-22/Cyr61 axis in synovial cells regulates inflammation in rheumatoid arthritis. Arthritis Rheumatol. 66, 49–59 (2014).
Nakamachi, Y. et al. MicroRNA-124a is a key regulator of proliferation and monocyte chemoattractant protein 1 secretion in fibroblast-like synoviocytes from patients with rheumatoid arthritis. Arthritis Rheum. 60, 1294–1304 (2009).
Kawano, S. & Nakamachi, Y. miR-124a as a key regulator of proliferation and MCP-1 secretion in synoviocytes from patients with rheumatoid arthritis. Ann. Rheum. Dis. 70, i88–i91 (2011).
Zhou, Q. et al. Research of the methylation status of miR-124a gene promoter among rheumatoid arthritis patients. Clin. Dev. Immunol. 2013, 524204 (2013).
Zhou, Q. et al. Demethylation of microRNA-124a genes attenuated proliferation of rheumatoid arthritis derived fibroblast-like synoviocytes and synthesis of tumor necrosis factor-alpha. PLoS One 11, e0164207 (2016).
Wang, J. et al. Identification of a novel microRNA-141-3p/Forkhead box C1/beta-catenin axis associated with rheumatoid arthritis synovial fibroblast function in vivo and in vitro. Theranostics 10, 5412–5434 (2020).
Liu, L. et al. MicroRNA-106b Overexpression suppresses synovial inflammation and alleviates synovial damage in patients with rheumatoid arthritis. Mod. Rheumatol. 32, 1054–1063 (2021).
Lee, W. S. et al. MicroRNA-9 ameliorates destructive arthritis through down-regulation of NF-kappaB1-RANKL pathway in fibroblast-like synoviocytes. Clin. Immunol. 212, 108348 (2020).
Philippe, L. et al. TLR2 expression is regulated by microRNA miR-19 in rheumatoid fibroblast-like synoviocytes. J. Immunol. 188, 454–461 (2012).
Shi, D. L. et al. MicroRNA-27a inhibits cell migration and invasion of fibroblast-like synoviocytes by targeting follistatin-like protein 1 in rheumatoid arthritis. Mol. Cells 39, 611–618 (2016).
Liu, J. et al. MicroRNA-29a inhibits proliferation and induces apoptosis in rheumatoid arthritis fibroblast-like synoviocytes by repressing STAT3. Biomed. Pharmacother. 96, 173–181 (2017).
Li, S. et al. MicroRNA-192 suppresses cell proliferation and induces apoptosis in human rheumatoid arthritis fibroblast-like synoviocytes by downregulating caveolin 1. Mol. Cell Biochem. 432, 123–130 (2017).
Wangyang, Y. et al. MiR-199a-3p inhibits proliferation and induces apoptosis in rheumatoid arthritis fibroblast-like synoviocytes via suppressing retinoblastoma 1. Biosci. Rep. 38, BSR20180982 (2018).
Miao, C. G. et al. miR-375 regulates the canonical Wnt pathway through FZD8 silencing in arthritis synovial fibroblasts. Immunol. Lett. 164, 1–10 (2015).
Song, A. F. et al. MiR-34a-5p inhibits fibroblastlike synoviocytes proliferation via XBP1. Eur. Rev. Med. Pharm. Sci. 24, 11675–11682 (2020).
Wu, H. et al. miR-34a in extracellular vesicles from bone marrow mesenchymal stem cells reduces rheumatoid arthritis inflammation via the cyclin I/ATM/ATR/p53 axis. J. Cell Mol. Med. 25, 1896–1910 (2021).
Wei, H. et al. MicroRNA-15a/16/SOX5 axis promotes migration, invasion and inflammatory response in rheumatoid arthritis fibroblast-like synoviocytes. Aging (Albany NY) 12, 14376–14390 (2020).
Fu, D. et al. MiR-3926 inhibits synovial fibroblasts proliferation and inflammatory cytokines secretion through targeting toll like receptor 5. Gene 687, 200–206 (2019).
Xie, Z. et al. MiR-20a inhibits the progression of human arthritis fibroblast-like synoviocytes and inflammatory factor expression by targeting ADAM10. Environ. Toxicol. 35, 867–878 (2020).
Baltimore, D. et al. MicroRNAs: new regulators of immune cell development and function. Nat. Immunol. 9, 839–845 (2008).
Cobos Jimenez, V. et al. Next-generation sequencing of microRNAs uncovers expression signatures in polarized macrophages. Physiol. Genom. 46, 91–103 (2014).
Paoletti, A. et al. Monocyte/macrophage abnormalities specific to rheumatoid arthritis are linked to miR-155 and are differentially modulated by different TNF inhibitors. J. Immunol. 203, 1766–1775 (2019).
Xu, J. et al. NOTCH reprograms mitochondrial metabolism for proinflammatory macrophage activation. J. Clin. Investig. 125, 1579–1590 (2015).
Huang, C. et al. MiR-146a modulates macrophage polarization by inhibiting Notch1 pathway in RAW264.7 macrophages. Int Immunopharmacol. 32, 46–54 (2016).
O’Connell, R. M. et al. MicroRNA-155 is induced during the macrophage inflammatory response. Proc. Natl. Acad. Sci. USA 104, 1604–1609 (2007).
Taganov, K. D. et al. NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 103, 12481–12486 (2006).
Mann, M. et al. An NF-kappaB-microRNA regulatory network tunes macrophage inflammatory responses. Nat. Commun. 8, 851 (2017).
Du, F. et al. MicroRNA-155 deficiency results in decreased macrophage inflammation and attenuated atherogenesis in apolipoprotein E-deficient mice. Arterioscler Thromb. Vasc. Biol. 34, 759–767 (2014).
Schulte, L. N. et al. Differential activation and functional specialization of miR-146 and miR-155 in innate immune sensing. Nucleic Acids Res. 41, 542–553 (2013).
Nakasa, T. et al. The inhibitory effect of microRNA-146a expression on bone destruction in collagen-induced arthritis. Arthritis Rheum. 63, 1582–1590 (2011).
Rajasekhar, M. et al. MicroRNA-155 contributes to enhanced resistance to apoptosis in monocytes from patients with rheumatoid arthritis. J. Autoimmun. 79, 53–62 (2017).
Elmesmari, A. et al. MicroRNA-155 regulates monocyte chemokine and chemokine receptor expression in Rheumatoid Arthritis. Rheumatol. (Oxf.) 55, 2056–2065 (2016).
Kurowska-Stolarska, M. et al. MicroRNA-155 as a proinflammatory regulator in clinical and experimental arthritis. Proc. Natl. Acad. Sci. USA 108, 11193–11198 (2011).
Ghani, S. et al. Macrophage development from HSCs requires PU.1-coordinated microRNA expression. Blood 118, 2275–2284 (2011).
Chan, E. K. et al. MicroRNA-146a in autoimmunity and innate immune responses. Ann. Rheum. Dis. 72, ii90–ii95 (2013).
Ammari, M. et al. Delivery of miR-146a to Ly6C(high) Monocytes Inhibits Pathogenic Bone Erosion in Inflammatory Arthritis. Theranostics 8, 5972–5985 (2018).
Chen, Q. et al. Inducible microRNA-223 down-regulation promotes TLR-triggered IL-6 and IL-1beta production in macrophages by targeting STAT3. PLoS One 7, e42971 (2012).
Xue, Y. L. et al. Long non-coding RNA MEG3 inhibits M2 macrophage polarization by activating TRAF6 via microRNA-223 down-regulation in viral myocarditis. J. Cell Mol. Med. 24, 12341–12354 (2020).
Zhuang, G. et al. A novel regulator of macrophage activation: miR-223 in obesity-associated adipose tissue inflammation. Circulation 125, 2892–2903 (2012).
Zhang, Q. B. et al. MicroRNA-223 Suppresses IL-1beta and TNF-alpha Production in Gouty Inflammation by Targeting the NLRP3 Inflammasome. Front. Pharm. 12, 637415 (2021).
Ogando, J. et al. Notch-regulated miR-223 targets the aryl hydrocarbon receptor pathway and increases cytokine production in macrophages from rheumatoid arthritis patients. Sci. Rep. 6, 20223 (2016).
Quero, L. et al. MiR-221-3p drives the shift of M2-macrophages to a pro-Inflammatory function by suppressing JAK3/STAT3 activation. Front. Immunol. 10, 3087 (2019).
Banerjee, S. et al. MiR-125a-5p regulates differential activation of macrophages and inflammation. J. Biol. Chem. 288, 35428–35436 (2013).
Wang, W. & Guo, Z. H. Downregulation of lncRNA NEAT1 ameliorates LPS-induced inflammatory responses by promoting macrophage M2 polarization via miR-125a-5p/TRAF6/TAK1 axis. Inflammation 43, 1548–1560 (2020).
Zhou, G. et al. Role and mechanism of miR-144-5p in LPS-induced macrophages. Exp. Ther. Med. 19, 241–247 (2020).
Zhu, D. et al. MicroRNA-17/20a/106a modulate macrophage inflammatory responses through targeting signal-regulatory protein alpha. J. Allergy Clin. Immunol. 132, 426–436.e428 (2013).
Jo, E. K. et al. Molecular mechanisms regulating NLRP3 inflammasome activation. Cell Mol. Immunol. 13, 148–159 (2016).
Bai, B. et al. NLRP3 inflammasome in endothelial dysfunction. Cell Death Dis. 11, 776 (2020).
Choulaki, C. et al. Enhanced activity of NLRP3 inflammasome in peripheral blood cells of patients with active rheumatoid arthritis. Arthritis Res. Ther. 17, 257 (2015).
Zhang, Y. et al. NLRP3 inflammasome plays an important role in the pathogenesis of collagen-induced arthritis. Mediators Inflamm. 2016, 9656270 (2016).
Guo, C. et al. NLRP3 inflammasome activation contributes to the pathogenesis of rheumatoid arthritis. Clin. Exp. Immunol. 194, 231–243 (2018).
Xie, Q. et al. MicroRNA33 regulates the NLRP3 inflammasome signaling pathway in macrophages. Mol. Med. Rep. 17, 3318–3327 (2018).
Yang, Q. et al. RelA/MicroRNA-30a/NLRP3 signal axis is involved in rheumatoid arthritis via regulating NLRP3 inflammasome in macrophages. Cell Death Dis. 12, 1060 (2021).
Lai, N. S. et al. Anti-citrullinated protein antibodies suppress let-7a expression in monocytes from patients with rheumatoid arthritis and facilitate the inflammatory responses in rheumatoid arthritis. Immunobiology 220, 1351–1358 (2015).
Zhu, W. et al. MiR-let-7a regulates anti-citrullinated protein antibody-induced macrophage activation and correlates with the development of experimental rheumatoid arthritis. Int. Immunopharmacol. 51, 40–46 (2017).
Kim, S. J. et al. Identification of a novel toll-like receptor 7 endogenous ligand in rheumatoid arthritis synovial fluid that can provoke arthritic joint inflammation. Arthritis Rheumatol. 68, 1099–1110 (2016).
Banerjee, A. et al. Micro-RNA-155 inhibits IFN-gamma signaling in CD4+ T cells. Eur. J. Immunol. 40, 225–231 (2010).
Jiang, S. et al. Molecular dissection of the miR-17-92 cluster’s critical dual roles in promoting Th1 responses and preventing inducible Treg differentiation. Blood 118, 5487–5497 (2011).
Steiner, D. F. et al. MicroRNA-29 regulates T-box transcription factors and interferon-gamma production in helper T cells. Immunity 35, 169–181 (2011).
Dong, L. et al. Decreased expression of microRNA-21 correlates with the imbalance of Th17 and Treg cells in patients with rheumatoid arthritis. J. Cell Mol. Med. 18, 2213–2224 (2014).
Escobar, T. M. et al. MiR-155 activates cytokine gene expression in Th17 cells by regulating the DNA-binding protein Jarid2 to relieve polycomb-mediated repression. Immunity 40, 865–879 (2014).
Lu, L. F. et al. Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells by targeting SOCS1 protein. Immunity 30, 80–91 (2009).
Hippen, K. L. et al. Effects of MicroRNA on Regulatory T Cells and Implications for Adoptive Cellular Therapy to Ameliorate Graft-versus-Host Disease. Front. Immunol. 9, 57 (2018).
Kmiolek, T. et al. The Interplay between Transcriptional Factors and MicroRNAs as an Important Factor for Th17/Treg Balance in RA Patients. Int. J. Mol. Sci. 21, 7169 (2020).
Zhou, Q. et al. Decreased expression of miR-146a and miR-155 contributes to an abnormal Treg phenotype in patients with rheumatoid arthritis. Ann. Rheum. Dis. 74, 1265–1274 (2015).
Lu, L. F. et al. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell 142, 914–929 (2010).
Tang, X. et al. Correlation between the expression of microRNA-301a-3p and the proportion of th17 cells in patients with rheumatoid arthritis. Inflammation 39, 759–767 (2016).
Korn, T. et al. IL-17 and Th17 Cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Yang, P. et al. MicroRNA let-7g-5p alleviates murine collagen-induced arthritis by inhibiting Th17 cell differentiation. Biochem. Pharm. 174, 113822 (2020).
Di, Y. et al. Catalpol inhibits tregs-to-Th17 cell transdifferentiation by up-regulating let-7g-5p to reduce STAT3 protein levels. Yonsei Med. J. 63, 56–65 (2022).
Zhang, M. F. et al. MicroRNA-26b-5p alleviates murine collagen-induced arthritis by modulating Th17 cell plasticity. Cell Immunol. 365, 104382 (2021).
Xie, M. et al. NF-kappaB-driven miR-34a impairs Treg/Th17 balance via targeting Foxp3. J. Autoimmun. 102, 96–113 (2019).
Jin, S. et al. Protectin DX restores Treg/Th17 cell balance in rheumatoid arthritis by inhibiting NLRP3 inflammasome via miR-20a. Cell Death Dis. 12, 280 (2021).
Jin, S. et al. Maresin 1 improves the Treg/Th17 imbalance in rheumatoid arthritis through miR-21. Ann. Rheum. Dis. 77, 1644–1652 (2018).
Wu, Y. H. et al. Upregulated Expression of microRNA-16 Correlates with Th17/Treg Cell Imbalance in Patients with Rheumatoid Arthritis. DNA Cell Biol. 35, 853–860 (2016).
Lu, J. et al. Follicular helper T cells: potential therapeutic targets in rheumatoid arthritis. Cell Mol. Life Sci. 78, 5095–5106 (2021).
Pratama, A. et al. MicroRNA-146a regulates ICOS-ICOSL signalling to limit accumulation of T follicular helper cells and germinal centres. Nat. Commun. 6, 6436 (2015).
Liu, W. H. et al. A miR-155-Peli1-c-Rel pathway controls the generation and function of T follicular helper cells. J. Exp. Med. 213, 1901–1919 (2016).
Hu, R. et al. miR-155 promotes T follicular helper cell accumulation during chronic, low-grade inflammation. Immunity 41, 605–619 (2014).
Alivernini, S. et al. MicroRNA-155 influences B-cell function through PU.1 in rheumatoid arthritis. Nat. Commun. 7, 12970 (2016).
Heinicke, F. et al. MicroRNA Expression Differences in Blood-Derived CD19+ B Cells of Methotrexate Treated Rheumatoid Arthritis Patients. Front. Immunol. 12, 663736 (2021).
Pan, F. et al. Dendritic Cells from Rheumatoid Arthritis Patient Peripheral Blood Induce Th17 Cell Differentiation via miR-363/Integrin alphav/TGF-beta Axis. Scand. J. Immunol. 85, 441–449 (2017).
Kurowska-Stolarska, M. et al. MicroRNA-34a dependent regulation of AXL controls the activation of dendritic cells in inflammatory arthritis. Nat. Commun. 8, 15877 (2017).
De la Rosa, I. A. et al. Impaired microRNA processing in neutrophils from rheumatoid arthritis patients confers their pathogenic profile. Modulation by biological therapies. Haematologica 105, 2250–2261 (2020).
Murata, K. et al. MicroRNA-451 down-regulates neutrophil chemotaxis via p38 MAPK. Arthritis Rheumatol. 66, 549–559 (2014).
Lian, J. B. et al. MicroRNA control of bone formation and homeostasis. Nat. Rev. Endocrinol. 8, 212–227 (2012).
Mizoguchi, F. et al. Osteoclast-specific Dicer gene deficiency suppresses osteoclastic bone resorption. J. Cell Biochem. 109, 866–875 (2010).
Wang, T. et al. MicroRNA-106b inhibits osteoclastogenesis and osteolysis by targeting RANKL in giant cell tumor of bone. Oncotarget 6, 18980–18996 (2015).
Kong, X. H. et al. MicroRNA-20a suppresses RANKL-modulated osteoclastogenesis and prevents bone erosion in mice with rheumatoid arthritis through the TLR4/p38 pathway. J. Biol. Regul. Homeost. Agents 35, 921–931 (2021).
Tao, Y. et al. Downregulation of miR-106b attenuates inflammatory responses and joint damage in collagen-induced arthritis. Rheumatol. (Oxf.) 56, 1804–1813 (2017).
Guo, L. J. et al. MiR-125a TNF receptor-associated factor 6 to inhibit osteoclastogenesis. Exp. Cell Res. 321, 142–152 (2014).
Liu, K. et al. MiR-125 regulates PI3K/Akt/mTOR signaling pathway in rheumatoid arthritis rats via PARP2. Biosci. Rep. 39, BSR20180890 (2019).
Jiang, B. et al. MiR-143-3p inhibits the differentiation of osteoclast induced by synovial fibroblast and monocyte coculture in adjuvant-induced arthritic rats. Biomed. Res. Int. 2021, 5565973 (2021).
de la Rica, L. et al. NF-kappaB-direct activation of microRNAs with repressive effects on monocyte-specific genes is critical for osteoclast differentiation. Genome Biol. 16, 2 (2015).
Donate, P. B. et al. Cigarette smoke induces miR-132 in Th17 cells that enhance osteoclastogenesis in inflammatory arthritis. Proc. Natl. Acad. Sci. USA 118, e2017120118 (2021).
Lee, Y. et al. MicroRNA-124 regulates osteoclast differentiation. Bone 56, 383–389 (2013).
Nakamachi, Y. et al. MicroRNA-124 inhibits the progression of adjuvant-induced arthritis in rats. Ann. Rheum. Dis. 75, 601–608 (2016).
Pauley, K. M. & Cha, S. MiRNA-146a in rheumatoid arthritis: a new therapeutic strategy. Immunotherapy 3, 829–831 (2011).
Chen, Y. et al. MiR-145-5p increases osteoclast numbers in vitro and aggravates bone erosion in collagen-induced arthritis by targeting osteoprotegerin. Med. Sci. Monit. 24, 5292–5300 (2018).
Noack, M. & Miossec, P. Selected cytokine pathways in rheumatoid arthritis. Semin. Immunopathol. 39, 365–383 (2017).
Ohnuma, K. et al. MicroRNA-124 inhibits TNF-alpha- and IL-6-induced osteoclastogenesis. Rheumatol. Int. 39, 689–695 (2019).
Miller, C. H. et al. RBP-J-regulated miR-182 promotes TNF-alpha-induced osteoclastogenesis. J. Immunol. 196, 4977–4986 (2016).
Sun, Y. et al. Recent advances in osteoclast biological behavior. Front. Cell Dev. Biol. 9, 788680 (2021).
Wang, T. et al. Proteus mirabilis Vesicles Induce Mitochondrial Apoptosis by Regulating miR96-5p/Abca1 to Inhibit Osteoclastogenesis and Bone Loss. Front. Immunol. 13, 833040 (2022).
Maeda, Y. et al. Synovium-derived microRNAs regulate bone pathways in rheumatoid arthritis. J. Bone Min. Res. 32, 461–472 (2017).
Hassan, M. Q. et al. MiR-218 directs a Wnt signaling circuit to promote differentiation of osteoblasts and osteomimicry of metastatic cancer cells. J. Biol. Chem. 287, 42084–42092 (2012).
Iwamoto, N. et al. Osteogenic differentiation of fibroblast-like synovial cells in rheumatoid arthritis is induced by microRNA-218 through a ROBO/Slit pathway. Arthritis Res. Ther. 20, 189 (2018).
Yang, W. H. et al. Leptin induces oncostatin M production in osteoblasts by downregulating miR-93 through the Akt signaling pathway. Int. J. Mol. Sci. 15, 15778–15790 (2014).
Tsai, C. H. et al. Osteopontin inhibition of miR-129-3p enhances IL-17 expression and monocyte migration in rheumatoid arthritis. Biochim. Biophys. Acta Gen. Subj. 1861, 15–22 (2017).
Li, T. M. et al. YKL-40-induced inhibition of miR-590-3p promotes interleukin-18 expression and angiogenesis of endothelial progenitor cells. Int. J. Mol. Sci. 18, 920 (2017).
Kusumbe, A. P. et al. Coupling of angiogenesis and osteogenesis by a specific vessel subtype in bone. Nature 507, 323–328 (2014).
Chen, C. Y. et al. CCN1 promotes VEGF production in osteoblasts and induces endothelial progenitor cell angiogenesis by inhibiting miR-126 expression in rheumatoid arthritis. J. Bone Min. Res. 32, 34–45 (2017).
Chen, C. Y. et al. Enhancement of CCL2 expression and monocyte migration by CCN1 in osteoblasts through inhibiting miR-518a-5p: implication of rheumatoid arthritis therapy. Sci. Rep. 7, 421 (2017).
Huang, C. C. et al. S1P Increases VEGF production in osteoblasts and facilitates endothelial progenitor cell angiogenesis by inhibiting miR-16-5p expression via the c-Src/FAK signaling pathway in rheumatoid arthritis. Cells 10, 2168 (2021).
Chen, Y. J. et al. Systematic analysis of differential expression profile in rheumatoid arthritis chondrocytes using next-generation sequencing and bioinformatics approaches. Int. J. Med. Sci. 15, 1129–1142 (2018).
Zhuo, Q. et al. LncRNA ZNF667-AS1 alleviates rheumatoid arthritis by sponging miR-523-3p and inactivating the JAK/STAT signalling pathway. Autoimmunity 54, 406–414 (2021).
Zhang, H. J. et al. LncRNA HOTAIR alleviates rheumatoid arthritis by targeting miR-138 and inactivating NF-kappaB pathway. Int. Immunopharmacol. 50, 283–290 (2017).
Li, G. et al. LncRNA MEG3 inhibits rheumatoid arthritis through miR-141 and inactivation of AKT/mTOR signalling pathway. J. Cell Mol. Med. 23, 7116–7120 (2019).
Hu, J. et al. MiR-23a inhibited IL-17-mediated proinflammatory mediators expression via targeting IKKalpha in articular chondrocytes. Int. Immunopharmacol. 43, 1–6 (2017).
Li, G. et al. circFADS2 protects LPS-treated chondrocytes from apoptosis acting as an interceptor of miR-498/mTOR cross-talking. Aging (Albany NY) 11, 3348–3361 (2019).
Zhou, Y. et al. MicroRNA-27b-3p inhibits apoptosis of chondrocyte in rheumatoid arthritis by targeting HIPK2. Artif. Cells Nanomed. Biotechnol. 47, 1766–1771 (2019).
Jiang, L. & Cao, S. Role of microRNA-26a in cartilage injury and chondrocyte proliferation and apoptosis in rheumatoid arthritis rats by regulating expression of CTGF. J. Cell Physiol. 235, 979–992 (2020).
Miyaki, S. et al. MicroRNA-140 plays dual roles in both cartilage development and homeostasis. Genes Dev. 24, 1173–1185 (2010).
Si, H. B. et al. MiR-140 attenuates the progression of early-stage osteoarthritis by retarding chondrocyte senescence. Mol. Ther. Nucleic Acids 19, 15–30 (2020).
Wang, Z. et al. MiR-140-5p/miR-149 affects chondrocyte proliferation, apoptosis, and autophagy by targeting FUT1 in osteoarthritis. Inflammation 41, 959–971 (2018).
Li, X. et al. MiR-29a and MiR-140 Protect Chondrocytes against the Anti-Proliferation and Cell Matrix Signaling Changes by IL-1beta. Mol. Cells 39, 103–110 (2016).
Li, H. et al. MiR-140-5p inhibits synovial fibroblasts proliferation and inflammatory cytokines secretion through targeting TLR4. Biomed. Pharmacother. 96, 208–214 (2017).
Chen, J. et al. Exosomal miRNA-486-5p derived from rheumatoid arthritis fibroblast-like synoviocytes induces osteoblast differentiation through the Tob1/BMP/Smad pathway. Biomater. Sci. 8, 3430–3442 (2020).
Liu, D. et al. Synovial fibroblast-derived exosomal microRNA-106b suppresses chondrocyte proliferation and migration in rheumatoid arthritis via down-regulation of PDK4. J. Mol. Med. (Berl.) 98, 409–423 (2020).
Ding, Y. et al. Exosomes derived from synovial fibroblasts under hypoxia aggravate rheumatoid arthritis by regulating Treg/Th17 balance. Exp. Biol. Med. (Maywood) 245, 1177–1186 (2020).
Meng, Q. & Qiu, B. Exosomal microRNA-320a derived from mesenchymal stem cells regulates rheumatoid arthritis fibroblast-like synoviocyte activation by suppressing CXCL9 expression. Front. Physiol. 11, 441 (2020).
Chen, Z. et al. Therapeutic potential of mesenchymal cell-derived miRNA-150-5p-expressing exosomes in rheumatoid arthritis mediated by the modulation of MMP14 and VEGF. J. Immunol. 201, 2472–2482 (2018).
Meng, H. Y. et al. The inhibition by human MSCs-derived miRNA-124a overexpression exosomes in the proliferation and migration of rheumatoid arthritis-related fibroblast-like synoviocyte cell. BMC Musculoskelet. Disord. 21, 150 (2020).
Huang, Y. et al. Exosomal microRNA-140-3p from human umbilical cord mesenchymal stem cells attenuates joint injury of rats with rheumatoid arthritis by silencing SGK1. Mol. Med. 28, 36 (2022).
Tavasolian, F. et al. MiRNA-146a improves immunomodulatory effects of MSC-derived exosomes in rheumatoid arthritis. Curr. Gene Ther. 20, 297–312 (2020).
Wu, L. F. et al. Identification of novel rheumatoid arthritis-associated MiRNA-204-5p from plasma exosomes. Exp. Mol. Med. 54, 334–345 (2022).
Vickers, K. C. et al. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 13, 423–433 (2011).
Arroyo, J. D. et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 108, 5003–5008 (2011).
Lindner, K. et al. Circulating microRNAs: emerging biomarkers for diagnosis and prognosis in patients with gastrointestinal cancers. Clin. Sci. (Lond.) 128, 1–15 (2015).
Luque-Tevar, M. et al. Integrative clinical, molecular, and computational analysis identify novel biomarkers and differential profiles of anti-TNF response in rheumatoid arthritis. Front. Immunol. 12, 631662 (2021).
Taha, M. et al. Serum a proliferation-inducing ligand and MicroRNA-223 are associated with rheumatoid arthritis: diagnostic and prognostic implications. Mol. Med. 26, 92 (2020).
Filkova, M. et al. Association of circulating miR-223 and miR-16 with disease activity in patients with early rheumatoid arthritis. Ann. Rheum. Dis. 73, 1898–1904 (2014).
Ouboussad, L. et al. Profiling microRNAs in individuals at risk of progression to rheumatoid arthritis. Arthritis Res. Ther. 19, 288 (2017).
Cunningham, C. C. et al. Serum miRNA Signature in Rheumatoid Arthritis and “At-Risk Individuals”. Front. Immunol. 12, 633201 (2021).
Anaparti, V. et al. Whole blood microRNA expression pattern differentiates patients with rheumatoid arthritis, their seropositive first-degree relatives, and healthy unrelated control subjects. Arthritis Res. Ther. 19, 249 (2017).
Tang, J. et al. Identification of circulating miR-22-3p and let-7a-5p as novel diagnostic biomarkers for rheumatoid arthritis. Clin. Exp. Rheumatol. 40, 69–77 (2022).
Safari, F. et al. Plasma levels of microRNA-146a-5p, microRNA-24-3p, and microRNA-125a-5p as potential diagnostic biomarkers for rheumatoid arthris. Iran. J. Allergy Asthma Immunol. 20, 326–337 (2021).
Ormseth, M. J. et al. Utility of select plasma microRNA for disease and cardiovascular risk assessment in patients with rheumatoid arthritis. J. Rheumatol. 42, 1746–1751 (2015).
Liu, X. et al. Circulating microRNA-23b as a new biomarker for rheumatoid arthritis. Gene 712, 143911 (2019).
Kolarz, B. et al. Hypermethylation of the miR-155 gene in the whole blood and decreased plasma level of miR-155 in rheumatoid arthritis. PLoS ONE 15, e0233897 (2020).
Abdul-Maksoud, R. S. et al. Serum miR-210 and miR-155 expression levels as novel biomarkers for rheumatoid arthritis diagnosis. Br. J. Biomed. Sci. 74, 209–213 (2017).
Wang, Y. et al. MiR-548a-3p regulates inflammatory response via TLR4/NF-kappaB signaling pathway in rheumatoid arthritis. J. Cell Biochem. 120, 1133–1140 (2018).
Xu, D. et al. Exosome-encapsulated miR-6089 regulates inflammatory response via targeting TLR4. J. Cell Physiol. 234, 1502–1511 (2019).
Rodriguez-Muguruza, S. et al. A serum biomarker panel of exomiR-451a, exomiR-25-3p and soluble TWEAK for early diagnosis of rheumatoid arthritis. Front. Immunol. 12, 790880 (2021).
Ormseth, M. J. et al. Development and validation of a microRNA panel to differentiate between patients with rheumatoid arthritis or systemic lupus erythematosus and controls. J. Rheumatol. 47, 188–196 (2020).
Jin, F. et al. Serum microRNA profiles serve as novel biomarkers for autoimmune diseases. Front. Immunol. 9, 2381 (2018).
Zhang, H. et al. B Cell-Related circulating microRNAs with the potential value of biomarkers in the differential diagnosis, and distinguishment between the disease activity and lupus nephritis for systemic lupus erythematosus. Front. Immunol. 9, 1473 (2018).
Cheleschi, S. et al. Circulating mir-140 and leptin improve the accuracy of the differential diagnosis between psoriatic arthritis and rheumatoid arthritis: a case-control study. Transl. Res. 239, 18–34 (2022).
Murata, K. et al. Plasma and synovial fluid microRNAs as potential biomarkers of rheumatoid arthritis and osteoarthritis. Arthritis Res. Ther. 12, R86 (2010).
Duroux-Richard, I. et al. Circulating miRNA-125b is a potential biomarker predicting response to rituximab in rheumatoid arthritis. Mediators Inflamm. 2014, 342524 (2014).
Ciechomska, M. et al. Comprehensive microRNA and transcriptomic profiling of rheumatoid arthritis monocytes: role of microRNA-146b in pro-inflammatory progression. Rheumatol. (Oxf.) 60, 5424–5435 (2021).
Bae, S. C. & Lee, Y. H. MiR-146a levels in rheumatoid arthritis and their correlation with disease activity: a meta-analysis. Int. J. Rheum. Dis. 21, 1335–1342 (2018).
Li, Z. et al. Role of miR-9-5p in preventing peripheral neuropathy in patients with rheumatoid arthritis by targeting REST/miR-132 pathway. Vitr. Cell Dev. Biol. Anim. 55, 52–61 (2019).
Oka, S. et al. Plasma miRNA expression profiles in rheumatoid arthritis associated interstitial lung disease. BMC Musculoskelet. Disord. 18, 21 (2017).
Ormseth, M. J. et al. Plasma miRNAs improve the prediction of coronary atherosclerosis in patients with rheumatoid arthritis. Clin. Rheumatol. 40, 2211–2219 (2021).
Liao, T. L. et al. Rituximab May Cause Increased Hepatitis C Virus Viremia in Rheumatoid Arthritis Patients Through Declining Exosomal MicroRNA-155. Arthritis Rheumatol. 70, 1209–1219 (2018).
Hruskova, V. et al. MicroRNA-125b: association with disease activity and the treatment response of patients with early rheumatoid arthritis. Arthritis Res. Ther. 18, 124 (2016).
Singh, A. et al. MicroRNA-132, miR-146a, and miR-155 as potential biomarkers of methotrexate response in patients with rheumatoid arthritis. Clin. Rheumatol. 38, 877–884 (2019).
Castro-Villegas, C. et al. Circulating miRNAs as potential biomarkers of therapy effectiveness in rheumatoid arthritis patients treated with anti-TNFalpha. Arthritis Res. Ther. 17, 49 (2015).
Krintel, S. B. et al. Prediction of treatment response to adalimumab: a double-blind placebo-controlled study of circulating microRNA in patients with early rheumatoid arthritis. Pharmacogenom. J. 16, 141–146 (2016).
Sode, J. et al. Plasma microRNA profiles in patients with early rheumatoid arthritis responding to adalimumab plus methotrexate vs methotrexate alone: a placebo-controlled clinical trial. J. Rheumatol. 45, 53–61 (2018).
Ciechomska, M. et al. Changes in miRNA-5196 expression as a potential biomarker of anti-TNF-alpha therapy in rheumatoid arthritis and ankylosing spondylitis patients. Arch. Immunol. Ther. Exp. (Warsz.) 66, 389–397 (2018).
Liu, Y. et al. Correlation of microRNA expression profile with clinical response to tumor necrosis factor inhibitor in treating rheumatoid arthritis patients: A prospective cohort study. J. Clin. Lab. Anal. 33, e22953 (2019).
Fernandez-Ruiz, J. C. et al. Analysis of miRNA expression in patients with rheumatoid arthritis during remission and relapse after a 5-year trial of tofacitinib treatment. Int Immunopharmacol. 63, 35–42 (2018).
Nasonov, E. et al. Olokizumab, a monoclonal antibody against interleukin 6, in combination with methotrexate in patients with rheumatoid arthritis inadequately controlled by methotrexate: efficacy and safety results of a randomised controlled phase III study. Ann. Rheum. Dis. 81, 469–479 (2022).
Bure, I. V. et al. Analysis of miRNA Expression in Patients with Rheumatoid Arthritis during Olokizumab Treatment. J. Pers. Med. 10, 205 (2020).
Burmester, G. R. & Pope, J. E. Novel treatment strategies in rheumatoid arthritis. Lancet 389, 2338–2348 (2017).
Guo, Q. et al. Rheumatoid arthritis: pathological mechanisms and modern pharmacologic therapies. Bone Res. 6, 15 (2018).
Iwamoto, N. et al. Methotrexate alters the expression of microRNA in fibroblast-like synovial cells in rheumatoid arthritis. Int. J. Mol. Sci. 22, 11561 (2021).
Ren, B. et al. TNF-alpha-elicited miR-29b potentiates resistance to apoptosis in peripheral blood monocytes from patients with rheumatoid arthritis. Apoptosis 24, 892–904 (2019).
Zisman, D. et al. Tocilizumab (TCZ) decreases angiogenesis in rheumatoid arthritis through its regulatory effect on miR-146a-5p and EMMPRIN/CD147. Front. Immunol. 12, 739592 (2021).
Hayakawa, K. et al. MicroRNA-766-3p Contributes to Anti-Inflammatory Responses through the Indirect Inhibition of NF-kappaB Signaling. Int. J. Mol. Sci. 20, 809 (2019).
Rupaimoole, R. & Slack, F. J. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Disco. 16, 203–222 (2017).
Ong, S. M. et al. MicroRNA-mediated immune modulation as a therapeutic strategy in host-implant integration. Adv. Drug Deliv. Rev. 88, 92–107 (2015).
Wu, J. et al. miR-708-5p promotes fibroblast-like synoviocytes’ cell apoptosis and ameliorates rheumatoid arthritis by the inhibition of Wnt3a/beta-catenin pathway. Drug Des. Devel. Ther. 12, 3439–3447 (2018).
Peng, J. S. et al. Amelioration of experimental autoimmune arthritis through targeting of synovial fibroblasts by intraarticular delivery of microRNAs 140-3p and 140-5p. Arthritis Rheumatol. 68, 370–381 (2016).
Liu, H. et al. Therapeutic prospects of microRNAs carried by mesenchymal stem cells-derived extracellular vesicles in autoimmune diseases. Life Sci. 277, 119458 (2021).
Cosenza, S. et al. Mesenchymal stem cells-derived exosomes are more immunosuppressive than microparticles in inflammatory arthritis. Theranostics 8, 1399–1410 (2018).
Zheng, J. et al. Bone marrow-derived mesenchymal stem cells-secreted exosomal microRNA-192-5p delays inflammatory response in rheumatoid arthritis. Int. Immunopharmacol. 78, 105985 (2020).
Li, G. Q. et al. MicroRNA-21 from bone marrow mesenchymal stem cell-derived extracellular vesicles targets TET1 to suppress KLF4 and alleviate rheumatoid arthritis. Ther. Adv. Chronic Dis. 12, 20406223211007369 (2021).
Huang, Y. et al. miR-223 in exosomes from bone marrow mesenchymal stem cells ameliorates rheumatoid arthritis via downregulation of NLRP3 expression in macrophages. Mol. Immunol. 143, 68–76 (2022).
You, D. G. et al. Metabolically engineered stem cell-derived exosomes to regulate macrophage heterogeneity in rheumatoid arthritis. Sci. Adv. 7, eabe0083 (2021).
Wiklander, O. P. et al. Extracellular vesicle in vivo biodistribution is determined by cell source, route of administration and targeting. J. Extracell. Vesic. 4, 26316 (2015).
Lv, H. et al. Toxicity of cationic lipids and cationic polymers in gene delivery. J. Control Release 114, 100–109 (2006).
Sujitha, S. et al. Berberine encapsulated PEG-coated liposomes attenuate Wnt1/beta-catenin signaling in rheumatoid arthritis via miR-23a activation. Eur. J. Pharm. Biopharm. 149, 170–191 (2020).
Hao, F. et al. Hybrid micelles containing methotrexate-conjugated polymer and co-loaded with microRNA-124 for rheumatoid arthritis therapy. Theranostics 9, 5282–5297 (2019).
Yu, C. et al. Ketoprofen and MicroRNA-124 Co-loaded poly (lactic-co-glycolic acid) microspheres inhibit progression of Adjuvant-induced arthritis in rats. Int J. Pharm. 552, 148–153 (2018).
Zhao, M. et al. Polyketal nanoparticles co-loaded with miR-124 and ketoprofen for treatment of rheumatoid arthritis. J. Pharm. Sci. 110, 2233–2240 (2021).
Zhang, X. et al. Efficient delivery of triptolide plus a miR-30-5p inhibitor through the use of near infrared laser responsive or CADY modified mSNs for efficacy in rheumatoid arthritis therapeutics. Front. Bioeng. Biotechnol. 8, 170 (2020).
Deng, Y. et al. Inflammation‐instructed hierarchical delivery of IL‐4/miR‐21 orchestrates osteoimmune microenvironment toward the treatment of rheumatoid arthritis. Adv. Funct. Mater. 31, 2101033 (2021).
Churov, A. V. et al. MicroRNAs in rheumatoid arthritis: altered expression and diagnostic potential. Autoimmun. Rev. 14, 1029–1037 (2015).
Akhtar, N. et al. MicroRNA-17 suppresses TNF-alpha signaling by interfering with TRAF2 and cIAP2 association in rheumatoid arthritis synovial fibroblasts. J. Immunol. 197, 2219–2228 (2016).
Guggino, G. et al. Downregulation of miRNA17-92 cluster marks Vgamma9Vdelta2 T cells from patients with rheumatoid arthritis. Arthritis Res. Ther. 20, 236 (2018).
Acknowledgements
This review work was supported by the National Natural Science Foundation of China (82072425, 82072498, 82272157), the Natural Science Foundation of Jiangsu Province (BE2020666, BK2021650), Jiangsu Medical Research Project (ZD2022021), the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD), Special Project of Diagnosis and Treatment Technology for Key Clinical Diseases in Suzhou (LCZX202003) and Postgraduate Research & Practice Innovation Program of Jiangsu Province (KYCX22_3217).
Author information
Authors and Affiliations
Contributions
All authors provided a substantial contribution to the conception, design and interpretation of the work, drafted the work or revised it critically for important intellectual content, and approved the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Peng, X., Wang, Q., Li, W. et al. Comprehensive overview of microRNA function in rheumatoid arthritis. Bone Res 11, 8 (2023). https://doi.org/10.1038/s41413-023-00244-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41413-023-00244-1
This article is cited by
-
Mechanistic role of quercetin as inhibitor for adenosine deaminase enzyme in rheumatoid arthritis: systematic review
Cellular & Molecular Biology Letters (2024)