Abstract
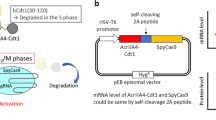
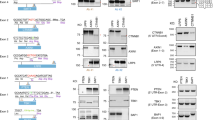
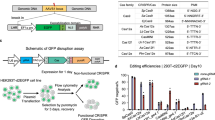
CRISPR/Cas9 has paved the way for the development of therapies that correct genetic mutations. However, constitutive expression of the Cas9 gene can increase off-target mutations and induce an immune response against the Cas9 protein. To limit the time during which the Cas9 nuclease is expressed, we proposed a simple drug inducible system. The approach consists of introducing a premature termination codon (PTC) in the Cas9 gene and subsequently treating with an aminoglycoside drug, which allows readthrough of the complete protein. To validate that system, HEK293T cells were co-transfected with a PX458 plasmid, which was mutated to introduce a PTC in the SpCas9 gene and two sgRNAs targeting the DMD gene (exons 50 and 54). Cells were treated with different doses of geneticin (G418) for 48 h. Western blot confirmed that the Cas9 protein expression, which was shut down by the PTC mutation, can be induced by the drug. The hybrid exon 50-54 formed by the deletion of part of the DMD gene was detected by PCR only in the cells treated with G418. The approach was also used successfully with CjCas9 to edit the FXN gene. Our results show that it is possible to control SpCas9 and CjCas9 expression by CRISPR-SCReT (CRISPR-Stop Codon Read Through) method.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Tremblay JP. The CRISPR system can correct or modify the expression of genes responsible for hereditary diseases. Med Sci (Paris). 2015;31:1014–22.
Doudna JA. The promise and challenge of therapeutic genome editing. Nature. 2020;578:229–36.
Young CS, Pyle AD, Spencer MJ. CRISPR for neuromuscular disorders: gene editing and beyond. Physiology (Bethesda). 2019;34:341–53.
Kim E, Koo T, Park SW, Kim D, Kim K, Cho HY, et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat Commun. 2017;8:14500.
Chang YJ, Bae J, Zhao Y, Lee G, Han J, Lee YH, et al. In vivo multiplex gene targeting with Streptococcus pyogens and Campylobacter jejuni Cas9 for pancreatic cancer modeling in wild-type animal. J Vet Sci. 2020;21:e26.
Iyombe-Engembe JP, Ouellet DL, Barbeau X, Rousseau J, Chapdelaine P, Lague P, et al. Efficient restoration of the dystrophin gene reading frame and protein structure in DMD myoblasts using the CinDel method. Mol Ther Nucleic Acids. 2016;5:e283.
Ouellet DL, Cherif K, Rousseau J, Tremblay JP. Deletion of the GAA repeats from the human frataxin gene using the CRISPR-Cas9 system in YG8R-derived cells and mouse models of Friedreich ataxia. Gene Ther. 2017;24:265–74.
Duchene BL, Cherif K, Iyombe-Engembe JP, Guyon A, Rousseau J, Ouellet DL, et al. CRISPR-induced deletion with SaCas9 restores dystrophin expression in dystrophic models in vitro and in vivo. Mol Ther. 2018;26:2604–16.
Wang D, Zhang F, Gao G. CRISPR-based therapeutic genome editing: strategies and in vivo delivery by AAV vectors. Cell. 2020;181:136–50.
Marino ND, Pinilla-Redondo R, Csorgo B, Bondy-Denomy J. Anti-CRISPR protein applications: natural brakes for CRISPR-Cas technologies. Nat Methods. 2020;17:471–9.
Li A, Tanner MR, Lee CM, Hurley AE, De Giorgi M, Jarrett KE, et al. AAV-CRISPR gene editing is negated by pre-existing immunity to Cas9. Mol Ther. 2020;28:1432–41.
Wagner DL, Amini L, Wendering DJ, Burkhardt LM, Akyuz L, Reinke P, et al. High prevalence of Streptococcus pyogenes Cas9-reactive T cells within the adult human population. Nat Med. 2019;25:242–8.
Charlesworth CT, Deshpande PS, Dever DP, Camarena J, Lemgart VT, Cromer MK, et al. Identification of preexisting adaptive immunity to Cas9 proteins in humans. Nat Med. 2019;25:249–54.
Barkau CL, O’Reilly D, Rohilla KJ, Damha MJ, Gagnon KT. Rationally designed anti-CRISPR nucleic acid inhibitors of CRISPR-Cas9. Nucleic Acid Ther. 2019;29:136–47.
Bondy-Denomy J. Protein inhibitors of CRISPR-Cas9. ACS Chem Biol. 2018;13:417–23.
Senturk S, Shirole NH, Nowak DG, Corbo V, Pal D, Vaughan A, et al. Rapid and tunable method to temporally control gene editing based on conditional Cas9 stabilization. Nat Commun. 2017;8:14370.
Keeling KM, Xue X, Gunn G, Bedwell DM. Therapeutics based on stop codon readthrough. Annu Rev Genomics Hum Genet. 2014;15:371–94.
Keeling KM, Wang D, Conard SE, Bedwell DM. Suppression of premature termination codons as a therapeutic approach. Crit Rev Biochem Mol Biol. 2012;47:444–63.
Malik V, Rodino-Klapac LR, Viollet L, Mendell JR. Aminoglycoside-induced mutation suppression (stop codon readthrough) as a therapeutic strategy for Duchenne muscular dystrophy. Ther Adv Neurol Disord. 2010;3:379–89.
Palmer E, Wilhelm JM, Sherman F. Phenotypic suppression of nonsense mutants in yeast by aminoglycoside antibiotics. Nature. 1979;277:148–50.
Francois B, Russell RJ, Murray JB, Aboul-ela F, Masquida B, Vicens Q, et al. Crystal structures of complexes between aminoglycosides and decoding A site oligonucleotides: role of the number of rings and positive charges in the specific binding leading to miscoding. Nucleic Acids Res. 2005;33:5677–90.
Garreau de Loubresse N, Prokhorova I, Holtkamp W, Rodnina MV, Yusupova G, Yusupov M. Structural basis for the inhibition of the eukaryotic ribosome. Nature. 2014;513:517–22.
Madisen L, Zwingman TA, Sunkin SM, Oh SW, Zariwala HA, Gu H, et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat Neurosci. 2010;13:133–40.
Anjomani Virmouni S, Ezzatizadeh V, Sandi C, Sandi M, Al-Mahdawi S, Chutake Y, et al. A novel GAA-repeat-expansion-based mouse model of Friedreich’s ataxia. Dis Model Mech. 2015;8:225–35.
McHugh DR, Steele MS, Valerio DM, Miron A, Mann RJ, LePage DF, et al. A G542X cystic fibrosis mouse model for examining nonsense mutation directed therapies. PLoS One. 2018;13:e0199573.
Friesen WJ, Johnson B, Sierra J, Zhuo J, Vazirani P, Xue X, et al. The minor gentamicin complex component, X2, is a potent premature stop codon readthrough molecule with therapeutic potential. PLoS One. 2018;13:e0206158.
He X, Urip BA, Zhang Z, Ngan CC, Feng B. Evolving AAV-delivered therapeutics towards ultimate cures. J Mol Med (Berl). 2021;99:593–617.
Pineda M, Lear A, Collins JP, Kiani S. Safe CRISPR: challenges and possible solutions. Trends Biotechnol. 2019;37:389–401.
Wilbie D, Walther J, Mastrobattista E. Delivery aspects of CRISPR/Cas for in vivo genome editing. Acc Chem Res. 2019;52:1555–64.
Marino ND, Pinilla-Redondo R, Csorgo B, Bondy-Denomy J. Anti-CRISPR protein applications: natural brakes for CRISPR-Cas technologies. Nat Methods. 2020;17:417–9.
Maji B, Gangopadhyay SA, Lee M, Shi M, Wu P, Heler R, et al. A high-throughput platform to identify small-molecule inhibitors of CRISPR-Cas9. Cell. 2019;177:1067–79.e19.
Gee P, Lung MSY, Okuzaki Y, Sasakawa N, Iguchi T, Makita Y, et al. Extracellular nanovesicles for packaging of CRISPR-Cas9 protein and sgRNA to induce therapeutic exon skipping. Nat Commun. 2020;11:1334.
Chen R, Huang H, Liu H, Xi J, Ning J, Zeng W, et al. Friend or foe? Evidence indicates endogenous exosomes can deliver functional gRNA and Cas9 protein. Small. 2019;15:e1902686.
Finn JD, Smith AR, Patel MC, Shaw L, Youniss MR, van Heteren J, et al. A single administration of CRISPR/Cas9 lipid nanoparticles achieves robust and persistent in vivo genome editing. Cell Rep. 2018;22:2227–35.
Petris G, Casini A, Montagna C, Lorenzin F, Prandi D, Romanel A, et al. Hit and go CAS9 delivered through a lentiviral based self-limiting circuit. Nat Commun. 2017;8:15334.
Guo T, Feng YL, Xiao JJ, Liu Q, Sun XN, Xiang JF, et al. Harnessing accurate non-homologous end joining for efficient precise deletion in CRISPR/Cas9-mediated genome editing. Genome Biol. 2018;19:170.
Li A, Lee CM, Hurley AE, Jarrett KE, De Giorgi M, Lu W, et al. A self-deleting AAV-CRISPR system for in vivo genome editing. Mol Ther Methods Clin Dev. 2019;12:111–22.
Barton-Davis ER, Cordier L, Shoturma DI, Leland SE, Sweeney HL. Aminoglycoside antibiotics restore dystrophin function to skeletal muscles of mdx mice. J Clin Invest. 1999;104:375–81.
Baradaran-Heravi A, Balgi AD, Zimmerman C, Choi K, Shidmoossavee FS, Tan JS, et al. Novel small molecules potentiate premature termination codon readthrough by aminoglycosides. Nucleic Acids Res. 2016;44:6583–98.
Ferguson MW, Gerak CAN, Chow CCT, Rastelli EJ, Elmore KE, Stahl F, et al. The antimalarial drug mefloquine enhances TP53 premature termination codon readthrough by aminoglycoside G418. PLoS One. 2019;14:e0216423.
Frew J, Baradaran-Heravi A, Balgi AD, Wu X, Yan TD, Arns S, et al. Premature termination codon readthrough upregulates progranulin expression and improves lysosomal function in preclinical models of GRN deficiency. Mol Neurodegener. 2020;15:21.
Borgatti M, Altamura E, Salvatori F, D’Aversa E, Altamura N. Screening readthrough compounds to suppress nonsense mutations: possible application to beta-thalassemia. J Clin Med. 2020;9:289.
Baiazitov RY, Friesen W, Johnson B, Mollin A, Sheedy J, Sierra J, et al. Chemical modifications of G418 (geneticin): Synthesis of novel readthrough aminoglycosides results in an improved in vitro safety window but no improvements in vivo. Carbohydr Res. 2020;495:108058.
Richardson R, Smart M, Tracey-White D, Webster AR, Moosajee M. Mechanism and evidence of nonsense suppression therapy for genetic eye disorders. Exp Eye Res. 2017;155:24–37.
Popp MW, Maquat LE. Leveraging rules of nonsense-mediated mRNA decay for genome engineering and personalized medicine. Cell. 2016;165:1319–22.
Manuvakhova M, Keeling K, Bedwell DM. Aminoglycoside antibiotics mediate context-dependent suppression of termination codons in a mammalian translation system. RNA. 2000;6:1044–55.
Floquet C, Rousset JP, Bidou L. [Allele-specific therapy: suppression of nonsense mutations by readthrough inducers]. Med Sci (Paris). 2012;28:193–9.
Benhabiles H, Gonzalez-Hilarion S, Amand S, Bailly C, Prevotat A, Reix P, et al. Optimized approach for the identification of highly efficient correctors of nonsense mutations in human diseases. PLoS One. 2017;12:e0187930.
Acknowledgements
We would like to thank the Canadian Francophonie Scholarship Program (CFSP) financed by Foreign Affairs for studentship support to PY.
Funding
This work was supported by grants from the Canadian Institutes of Health Research (CIHR), the FRQS ThéCel network, Ataxia Canada, and the Foundation for Cell and Gene Therapies (also named the Jesse’s Journey).
Author information
Authors and Affiliations
Contributions
PY designed the experiments, performed the experiments, and wrote the manuscript. BLD provided technical assistance for the molecular biology. NM assisted with the design of the experiments and corrected the manuscript. JPT conceived the experiments and corrected the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Yaméogo, P., Duchêne, B.L., Majeau, N. et al. CRISPR-SCReT (CRISPR-Stop Codon Read Through) method to control Cas9 expression for gene editing. Gene Ther 29, 171–177 (2022). https://doi.org/10.1038/s41434-021-00297-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41434-021-00297-z