Abstract
In plants, chloroplasts are the sites at which photosynthesis occurs, and an increased abundance of chloroplasts increases the nutritional quality of plants and the resultant color of fruits. However, the molecular mechanisms underlying chlorophyll synthesis and chloroplast development in tomato fruits remain unknown. In this study, we isolated a chlorophyll-deficient mutant, reduced chlorophyll mutant 1 (rcm1), by ethylmethanesulfonate mutagenesis; this mutant produced yellowish fruits with altered chloroplast development. MutMap revealed that Solyc08g005010 is the causal gene underlying the rcm1 mutant phenotype. A single-nucleotide base substitution in the second exon of SlRCM1 results in premature termination of its translated protein. SlRCM1 encodes a chloroplast-targeted metalloendopeptidase that is orthologous to the BCM1 protein of Arabidopsis and the stay-green G protein of soybean (Glycine max L. Merr.). Notably, the yellowish phenotype of the lutescent1 mutant can be restored with the allele of SlRCM1 from wild-type tomato. In contrast, knockout of SlRCM1 by the CRISPR/Cas9 system in Alisa Craig yielded yellowish fruits at the mature green stage, as was the case for lutescent1. Amino acid sequence alignment and functional complementation assays showed that SlRCM1 is indeed Lutescent1. These findings provide new insights into the regulation of chloroplast development in tomato fruits.
Similar content being viewed by others
Introduction
Fruit development is a complex and highly coordinated process that involves a series of specific physiological and biochemical changes1,2. During tomato fruit development, chloroplasts serve as sites of photosynthesis and carbohydrate accumulation and can be transformed into chromoplasts for carotenoid formation3. Chloroplast development in tomato plants is directly proportional to fruit development and nutrient accumulation. Both the synthesis and degradation of chloroplasts in plants are in a stable state of dynamic equilibrium. Therefore, it is of enormous importance to study the molecular mechanism underlying chloroplast development in the process of fruit ripening and development.
Chloroplast development has been demonstrated to be regulated by multiple transcription factors, among which GLK2 is an influential transcription factor that regulates this process. Tomato fruits with full-length transcripts of GLK2 mRNA exhibit a dark green shoulder that can promote photosynthesis and accumulate more nutrients, while glk2 mutation eliminates this green shoulder4. GLK2 overexpression resulted in dramatic upregulated expression of the Solyc08g005010 gene, indicating a regulatory network involving chloroplast development5. The TKN2 and TKN4 genes in tomato modulate the gradient of chloroplast development in fruits by regulating GLK2 expression6. Overexpression of the APRR2-like gene in tomato increased the content of chlorophyll in fruits7. SlBBX20 in tomato regulates chloroplast development by modulating the expression of the SlCAB1B,SlCAB6A and SlCHL27 genes8. SlBEL11 encodes a transcription factor that negatively regulates chloroplast development and chlorophyll synthesis in tomato fruits. The SlBEL11 protein can directly bind to the promoters of genes involved in chloroplast development and chlorophyll synthesis,such as TKN2,CAB and POR, and downregulate the expression of these types of genes9. In plant growth and development, auxin signaling affects chlorophyll synthesis and chloroplast development. The auxin response factor ARF10 directly binds to the promoter of GLK1 and activates its expression, thereby promoting chloroplast development and sugar accumulation10. SlARF6A directly binds to the promoters of the GLK1, CAB1, CAB2 and RbcS genes and promotes their expression to modulate tomato fruit chloroplast development11. ARF2A, an auxin signal component, participates in fruit ripening and chloroplast development. In Arabidopsis, the ARF2 gene is involved in the degradation of chlorophyll in leaves12. The chlorophyll content of ARF2A-OX transgenic fruit was significantly lower than that of wild-type fruit at 42 days post-anthesis13.
Chlorophyll synthesis in plants is a complex process involving 15 enzymes encoded by 27 genes14. Mg-chelatase catalyzes the binding of Mg2+ to protoporphyrin IX, which represents the first step in chlorophyll synthesis15, and Mg-protoporphyrin IX is further methylated by Mg-protoporphyrin IX methyltransferase, followed by four subsequent catalytic reactions to produce chlorophyll16. Chlorophyllide a oxygenase (CAO) catalyzes the conversion of chlorophyll a to chlorophyll b and plays an important role in the balance of chlorophyll a and chlorophyll b. Overexpression of CAO in tobacco promotes the synthesis of chlorophyll b and decreases the ratio of chlorophyll a and chlorophyll b17. GUN4, a porphyrin-binding protein, enhances the activity of Mg-chelatase by binding to protoporphyrin, which is the substrate of the Mg-chelatase reaction18. Overexpression of GUN4 can significantly increase the chlorophyll content in tobacco leaves19. SGR1, encoding Mg dechelatase, promotes chloroplast degradation during plant maturation20. Mutation of SGR1 in Chinese cabbage leads to increased chlorophyll concentrations and a stay-green phenotype21. In Arabidopsis, the bcm1 mutant exhibits a pale-green leaf phenotype due to reduced chlorophyll contents. BCM1 interacts with GUN4 to enhance Mg-chelatase activity. BCM1 can also interact with SGR1 to destabilize the SGR1 protein22. GluTR is a glutamyl tRNA reductase involved in porphyrin and chlorophyll biosynthesis23. Interactions between BCM1 and GluTR affect the synthesis of 5-aminolevulinic acid, a key precursor in the biosynthesis of porphyrin during chlorophyll synthesis22. The product of CHLM is magnesium protoporphyrin IX methyltransferase that converts Mg-protoporphyrin IX to Mg-protoporphyrin IX methylester during chlorophyll synthesis24. BCM1 interacts with CHLM to promote the formation of a MgCH-GUN4-CHLM enzyme complex22. The stay-green G gene is a homologous gene of BCM1 in soybean; this gene positively regulates chlorophyll synthesis in the soybean seed coat25. Therefore, BCM1 plays an important and conserved role in chlorophyll synthesis in different crop species.
In tomato, the Lutescent2 (L2) gene, which encodes a zinc metalloprotease, has been characterized, and this gene has been shown to regulate chloroplast development and fruit maturity26. Similarly, the number of chloroplasts per cell in immature green fruit pericarp tissue of lutescent1 (l1) mutant was reduced significantly compared with that of the wild type. In addition, fully expanded leaflets of the l1 mutant exhibited a more dramatic yellowish phenotype26. It was previously reported that the chlorophyll content of the fruits of the l1 mutant was reduced, leading to a senescent phenotype for l127. Chloroplast development is hindered in l128, although the causative gene underlying chloroplast defects and chlorophyll reduction in l1 has not yet been identified.
In this study, we obtained a reduced chlorophyll mutant (rcm1) of tomato; this mutant is an l1 allelic mutant with altered chloroplast development and was generated via ethylmethanesulfonate (EMS) mutagenesis. We discovered via BSA + DNA-Seq and MutMap that the rcm1 mutant carries a single-nucleotide polymorphism (T → A), resulting in premature termination of the SlRCM1 protein. Sequence analysis and functional characterization showed that the SlRCM1 gene, located at the Lutescent1 locus, encodes an ortholog of the BCM1 protein of Arabidopsis and the stay-green G protein of soybean (Glycine max L. Merr.). SlRCM1 regulates chlorophyll synthesis and chloroplast development in fruits at the mature green and red ripe stages, while its ortholog regulates chlorophyll synthesis in the leaves of Arabidopsis and seeds of soybean22,25. These findings highlight the molecular mechanisms underlying chlorophyll synthesis and chloroplast development in tomato fruits.
Results
The tomato rcm1 mutant exhibits altered chloroplast development
Seeds of the tomato cultivar Ligeer 87-5 were treated with 1% EMS to obtain novel mutants. A reduced chlorophyll mutant (rcm1) was isolated from the EMS-mutagenized population (Fig. 1a). The fruits of the rcm1 mutant exhibited reduced amounts of chlorophyll relative to those of the wild type. The fruits at the mature green (MG) and breaker (BR) stages of the rcm1 mutant showed a sharp decrease in chlorophyll relative to those of the wild type. However, the fruits of the rcm1 mutant could still turn red at the red ripe (RR) stage (Fig. 1b). Furthermore, the thylakoid membranes were collapsed in the chloroplasts of fruits of the rcm1 mutant at the mature green stage (Fig. 1c). The chloroplasts in the rcm1 fruits were smaller than those of the wild type (Fig. 1c). Subsequently, the chlorophyll content in the rcm1 mutant fruits was significantly lower than that in the wild-type fruits at the MG, BR and yellow red (YR) stages, and no chlorophyll was detected at the RR stage (Fig. 1d). The carotenoid content in the rcm1 mutant fruits was also significantly lower than that in the wild-type fruits at all four developmental stages (Fig. 1e). The a*/b* colorimetric values (Fig. S1a), which are proportional to the lycopene, and total soluble solids contents (Fig. S1b) of red ripe fruits of the rcm1 mutant decreased relative to those of the wild type. Fruit ripening of the rcm1 mutant was delayed by approximately 6 days compared to that of the wild-type (Fig. S2a), and the ethylene production rate of the rcm1 mutant fruits was lower than that of the wild-type fruits (Fig. S2b).
a Whole-plant morphology of the reduced-chlorophyll mutant (rcm1) and the wild type (Ligeer 87-5). Scale bars, 15 cm. b Phenotypes of Wild Type (WT) and rcm1 tomato fruits at four developmental stages. MG, mature green stage; BR, breaker stage; YR, yellow ripe stage; RR, red ripe stage. Scale bars, 2 cm. c Chloroplast ultrastructure of mature green fruits of wild-type and rcm1 mutant plants via transmission electron microscopy. P, plastoglobuli; TGS, thylakoid granum stacks. Scale bars, 10 μm (left) and 0.5 μm (right). d Chlorophyll content of fruit pericarps from wild type and rcm1 at the MG, BR, YR and RR stages. The data are presented as the means ± SDs (n = 6). e Carotenoid contents of fruit pericarps from wild type and rcm1 at the MG, BR, YR and RR stages. The data are presented as the means ± SDs (n = 6). f Phenotypes of the pistils of the wild type and rcm1 mutant. Scale bars, 2 mm. g Morphology of the leaves of the rcm1 mutant and the wild type. Scale bars, 2 cm. h Chloroplast ultrastructure of leaves of wild-type and rcm1 plants using transmission electron microscopy. SG, starch grains; TGS, thylakoid granum stacks. Scale bars, 0.5 μm. i Chlorophyll content of leaves from wild-type and rcm1 plants. The data are presented as the means ± SDs (n = 6). The asterisks indicate statistically significant differences according to t-tests: **, P-value < 0.01. nd, not detected
To fully evaluate the phenotype of the rcm1 mutant, the leaves and pistils were compared between the rcm1 mutant and wild type. A lack of chlorophyll accumulation was observed in the developing pistils and leaves of rcm1 relative to the wild type (Fig. 1f, g). Transmission electron microscopy revealed impaired development of thylakoid membranes in the chloroplasts of the rcm1 mutant compared to those of the wild type (Fig. 1h). Accordingly, the chlorophyll content in the rcm1 mutant leaves was significantly lower than that in the wild-type leaves (Fig. 1i). Compared to that of the wild type, the maximum photochemical efficiency (Fv/Fm) of leaves of the rcm1 mutant decreased significantly (Fig. S3a, b). Additionally, the quantum efficiency of PSII (Y(II)) of the leaves of the rcm1 mutant was significantly impaired compared with that of the wild type (Fig. S3c). Additionally, pollen vitality in rcm1 was lower than that in Ligeer 87-5 (WT); the percentage of malformed pollen in rcm1 was greater than that in the wild type (Fig. S4).
Cloning of SlRCM1
To genetically characterize the yellowish fruit phenotype of the rcm1 mutant, we generated an F2 population consisting of 307 individuals by crossing Ligeer 87-5 with the rcm1 mutant. The F1 generation of the cross between Ligeer 87-5 and rcm1 displayed a normal phenotype similar to that of Ligeer 87-5, which suggested that the gene underlying the reduced chlorophyll phenotype of the rcm1 mutant is a recessive gene. In the F2 population, the ratio between the number of individuals with a normal phenotype (237) and the number of individuals with a chlorophyll-deficient phenotype (70) was approximately 3:1 (χ2 = 0.68, χ20.05 = 3.84), indicating that the chlorophyll-deficient phenotype was controlled by a single gene (Table S1).
Next, we performed BSA and MutMap analyses to isolate the candidate genes. We sequenced two bulk populations comprising 25 individuals with green fruit or yellowish fruit at the mature green stage. Each library was sequenced at a depth of a ×25 genome equivalent. The generated reads were mapped to the tomato reference genome (M82), and allele frequency differences of 40,124 SNPs from the two pools were calculated and mapped across the 12 chromosomes in tomato to form a Manhattan plot (Fig. 2c). The confidence threshold exceeded 95% only at the beginning of chromosome 8, between SL2.50ch08_1 and SL2.50ch08_1010000 (Table S2). Generally, single-base mutations are frequently generated by EMS mutagenesis29,30. There were six SNPs between the two pools between SL2.50ch08_1 and SL2.50ch08_1010000 (Table S3). Genetic analysis of the segregating populations indicated that the phenotype of the rcm1 mutant is likely controlled by a single locus. Since M82 (the genotype of the reference genome) develops normal chloroplasts in its fruits, SNPs in the recessive pool with an allele frequency of 1 and different from the reference genome (M82) were scored, and only one SNP (A → T) at SL2.50ch08_16268 out of all SNPs in the coding regions was identified.
a Phenotypes of leaves and fruits from the wild-type and rcm1 mutant plants and their F1 progeny. Scale bars, 2 cm. b Chlorophyll content of mature leaves and MG-stage fruits from the wild type and rcm1 mutant and their F1 progeny. The data are presented as the means ± SDs (n = 3). The means followed by different letters indicate statistical significance at P = 0.05. c Differences in allele frequencies between 25 lines with green fruits and 25 lines with yellowish fruits in the F2 population. The X axis represents the 12 chromosomes of tomato. The Y axis represents the difference in allele frequencies (SNP values) between the two pools. The red line represents the 95% confidence interval. The SNPs associated with the 95% confidence interval are located between SL2.50ch08_1 and SL2.50ch08_1010000. The blue line represents the average value of the ΔSNP index per 1 Mb window. d The fragment containing SNP (SL2.50ch08_16268) was subjected to PCR amplification and Sanger sequencing. The red letters indicate the bases of M82 (T), Ligeer 87-5 (T), rcm1 (A) and members of the hybrid F2 generation (A) with reduced chlorophyll at SL2.50ch08_16268. “-“ indicates the antisense strand of the genome. e Structure of the SlRCM1 gene. The black lines represent introns and untranslated regions. The white boxes represent exons. The T (WT)-to-A (rcm1) substitution in the second exon of SlRCM1 caused a conversion from Tyr (WT) to a stop codon (rcm1)
Furthermore, no InDels or other structural variations were detected via 25 equivalent genome sequencing events of the WT and rcm1 mutants. We analyzed the allele frequency of the SNP at SL2.50ch08_16268 in both pools with wild-type and mutant phenotypes. The SNP at SL2.50ch08_16268 in M82 was A, whereas in the mutant pool, it was T (100%) (Table 1). Furthermore, the SNP at SL2.50ch08_16268 in 30 F2 progeny with yellowish fruit was T (100%), which was confirmed via PCR and Sanger sequencing. This SNP (SL2.50ch08_16268) occurs in the second exon of Solyc08g005010 and was further verified by Sanger sequencing. Because the transcriptional direction of Solyc08g005010 is opposite to that of the genome, the nucleotide of SL2.50ch08_16268 in the coding strand of Solyc08g005010 in M82 and Ligeer 87-5 was T, but it was A in the rcm1 mutant (Fig. 2d, e). The substitution of T (wild type) to A (rcm1 mutant) resulted in the conversion of Tyr (wild type) to a stop codon (rcm1 mutant) (Fig. 2e), indicating that the translation of the SlRCM1 protein was terminated prematurely in the rcm1 mutant. The predicted protein encoded by the SlRCM1 allele from the rcm1 mutant did not include any functional domain (Fig. S5a). Therefore, we inferred that the SlRCM1 allele in the rcm1 mutant is null. Furthermore, the expression level of the SlRCM1 allele in the rcm1 mutant was equivalent to that in the wild type (Fig. S5b). Therefore, we considered Solyc08g005010 to be the candidate SlRCM1 gene, which encodes a CAAX-type endopeptidase.
The yellowish phenotype of fruits of rcm1 was complemented by SlRCM1 from Ligeer 87-5
To verify whether SlRCM1 is the causal gene for the chlorophyll-deficient phenotype of the rcm1 mutant, SlRCM1 was overexpressed (OE) under the control of the CaMV35S promoter in the rcm1 mutant (Fig. 3a). Nineteen independent transgenic lines were obtained. The expression levels of the SlRCM1 gene in the OE-11 (#11) and OE-18 (#18) lines in the T1 generation significantly increased relative to those of the rcm1 mutant. The normal-green phenotypes of the fruits of the OE-11 and OE-18 lines were restored at the mature green stage (Fig. 3b). To further verify the biological function of SlRCM1, a transformation construct (Pro SlRCM1 (Ligeer 87-5)::CDSSlRCM1 (Ligeer 87-5)) was prepared by inserting the SlRCM1 CDS from Ligeer 87-5 into a pHELLSGATE8 vector under the native promoter of the transgene (Fig. 3c). We introduced this construct into the rcm1 mutant by Agrobacterium-mediated transformation. Eleven independent transgenic plants were identified. The phenotype of the rcm1 transgenic lines was restored to that of the wild type (normal green fruit) (Fig. 3d). Overall, these results confirmed that SlRCM1 is the correct candidate gene.
a Schematic structure of the expression vector with SlRCM1 driven by the CaMV35S promoter. b Overexpression of SlRCM1 driven by the CaMV35S promoter in rcm1. Overexpressing SlRCM1 in rcm1 rescued the phenotype, as was the case for Ligeer 87-5. The relative expression of SlRCM1 in mature green fruits of SlRCM1-overexpressing lines and untransformed control (rcm1) plants was quantified. Scale bars, 2 cm. c Schematic structure of the expression vector with SlRCM1 driven by its native 2944-bp promoter from Ligeer 87-5. The two arrows represent the primers used to analyze the transcript level of SlRCM1. Fw: Forward primer; Rv: Reverse primer. d The phenotype of the fruits of the transgenic lines could be restored to that of Ligeer 87-5. The relative expression of SlRCM1 in mature green fruits was quantified in the transgenic lines and rcm1. Scale bars, 2 cm. The ACTIN gene (Solyc11g005330) was used as the internal control. The data are presented as the means ± SDs (n = 3)
SlRCM1 regulates chloroplast development in tomato
To further verify that SlRCM1 regulates chloroplast development in tomato fruits, we overexpressed the SlRCM1 gene in S. lycopersicum (L.) cv. Alisa Craig (AC) by Agrobacterium-mediated transformation. Compared to those of the wild type, the fruits of the OE-1 and OE-2 lines appeared dark green at the mature green stage (Fig. 4a). The number of thylakoids and thylakoid granum stacks in mature green fruits of the OE lines was higher than that of the wild type (Fig. 4b). Interestingly, the locular material surrounding the seeds remained green, and chlorophyll could still be detected at the red ripe stage in the overexpression lines (Fig. 4a). Indeed, the expression levels of the SlRCM1 gene in the OE-1 and OE-2 lines were significantly higher than those in the wild-type (AC) line (Fig. 4c). The chlorophyll content in the MG- and RR-stage fruits of OE-1 and OE-2 lines increased significantly relative to that in the wild type (Fig. 4e, f). In addition, the carotenoid content in the MG- and RR-stage fruits of the OE-1 and OE-2 lines increased significantly relative to that of the wild type (Fig. 4g, h). Chlorophyll in the RR-stage fruit of AC was not detected. Furthermore, we used CRISPR/Cas9 (CR) to edit the first exon of SlRCM1 in the AC background (Fig. 4a, d). We determined the type of mutation in the T1 generation using PCR and Sanger sequencing. The SlRCM1 knockout line CR-1 contained a 2-bp deletion in SlRCM1, and the knockout line CR-2 contained a 1-bp insertion in SlRCM1 (Fig. 4d). Both the CR-1 and CR-2 lines exhibited a reduced chlorophyll phenotype at and after the mature green stage (Fig. 4a). Furthermore, the development of thylakoid membranes was impaired in the mature green fruits of the CR lines compared to that of the wild type (Fig. 4b). The chlorophyll content in the MG-stage fruits of CR-1 and CR-2 plants decreased significantly relative to that in the wild-type plants (Fig. 4e). These results indicate that SlRCM1 is responsible for chlorophyll synthesis and chloroplast development in tomato fruits.
a Fruits of AC, SlRCM1 overexpression and knockout lines at the mature green (MG) and red ripe (RR) stages. Scale bars, 3 cm. b The structures of chloroplasts in mature green fruits of AC, SlRCM1-OE (OE) and SlRCM1-CRISPR (CR) were observed via transmission electron microscopy. C, chloroplast; P, plastoglobulus; TGS, thylakoid granum stack. Scale bars, 0.5 μm. c The relative expression of SlRCM1 in mature green fruits of SlRCM1 overexpression lines and the control (AC). The ACTIN gene (Solyc11g005330) was used as the internal control. The data are presented as the means ± SDs (n = 3). d Mutation types of SlRCM1 knockout lines in the T1 generation were identified. e Chlorophyll content in the fruit pericarps of AC, SlRCM1-OE (OE) and SlRCM1-CRISPR (CR) lines at the mature green (MG) stage. f Chlorophyll content in the fruit pericarps of AC, SlRCM1-OE (OE) and SlRCM1-CRISPR (CR) lines at the red ripe (RR) stage. g Carotenoid content in the fruit pericarps of AC, SlRCM1-OE (OE) and SlRCM1-CRISPR (CR) lines at the MG stage. h Carotenoid content in the fruit pericarps of AC, SlRCM1-OE (OE) and SlRCM1-CRISPR (CR) lines at the RR stage. The data are presented as the means ± SDs (n = 6). The asterisks indicate statistically significant differences according to t-tests: **, P-value < 0.01. nd, not detected
SlRCM1 is Lutescent1
A spontaneous lutescent1 mutant (l1), LA3717, was derived from Ailsa Craig (AC). The l1 mutation has been shown to dramatically affect tomato fruit development. l1 mutants have a low chlorophyll content in their fruits, especially under high-light and dark conditions, which enhances the rate of chlorophyll loss26. To date, the gene underlying the l1 mutant has not yet been cloned. The fruits of SlRCM1 knockout lines (CR) generated using the CRISPR/Cas9 system exhibited the same yellowish phenotype as did the fruits of the l1 mutant in the Alisa Craig (AC) background described in the Tomato Genetics Resource Center (TGRC; https://tgrc.ucdavis.edu) (Fig. 5a). The yellowish phenotype of the fruits of the l1 mutant is purportedly controlled by a single gene located at the beginning of chromosome 826,31. We used SlRCM1-specific primers to amplify full-length gDNA and the 5-kb promoter of SlRCM1 from AC, Ligeer 87-5 and l1 (Table S4). A base deletion led to premature termination of the SlRCM1 protein in l1 (Fig. 5b, c). Furthermore, the expression level of the SlRCM1 allele in the l1 mutant was equivalent to that in AC (Fig. S5c). SlRCM1 was subsequently overexpressed under the control of the CaMV35S promoter in the l1 mutant. Nine independent transgenic lines were identified. The expression level of the SlRCM1 gene in the 2nd (#2) and 6th (#6) transgenic lines was significantly higher than that in the control (l1) (Fig. 5d). Functional complementation of SlRCM1 in l1 restored normal plant growth and development (Fig. 5e). Taken together, these results indicate that SlRCM1 may be a causal gene underlying the yellowish phenotype of the l1 mutant.
a Phenotypes of fruits from the CR-1 line, lutescent1 mutant (l1) and AC at different developmental stages. IMG, immature green stage; MG, mature green stage; BR, breaker stage; YR, yellow ripe stage; RR, red ripe stage. The l1 is a mutant in the AC background. Scale bars, 3 cm. b Nucleotide sequence alignment of SlRCM1 fragments in Ligeer 87-5, Ailsa Craig (AC) and l1. The other bases are consistent among Ligeer 87-5, AC and l1. The black star indicates the base at SL2.50ch08_16268 in the first exon of the SlRCM1 gene. c Amino acid sequence alignment of the SlRCM1 protein of Ligeer 87-5, AC and l1. d Relative expression of SlRCM1 in mature green fruits of the SlRCM1-overexpressing lines and the control (l1). The ACTIN gene (Solyc11g005330) was used as an internal control. The data are presented as the means ± SDs (n = 3). e Complementation of the l1 mutant using SlRCM1. Phenotypes of mature green fruits from l1- and SlRCM1-overexpressing lines were imaged and are shown. Scale bars, 3 cm
SlRCM1 expression and subcellular localization
The gDNA of the SlRCM1 gene is 6,299 bp in length and consists of nine exons and eight introns. The SlRCM1 gene encodes a protein comprising 376 amino acids. Alignment of amino acid sequences (Fig. S6) and phylogenetic analysis (Fig. 6a) showed that the SlRCM1 protein is highly conserved among soybean, potato, pepper, tobacco, Arabidopsis, rice, maize, jute and Cephalotus follicularis, suggesting that the SlRCM1 protein may play an essential role in plants. Tomato SlRCM1 may function in chloroplast development and chlorophyll synthesis through a conserved biological process, similar to that in soybean and Arabidopsis thaliana.
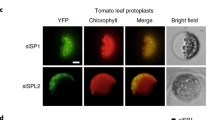
a Phylogenetic analysis of SlRCM1. The tomato SlRCM1 protein sequence was used to query (via BLAST) the homologous proteins of different species such as soybean (G), potato (Sotub08g006950), pepper (CA00g50240), tobacco (Niben101Scf06822g03002), Arabidopsis thaliana (BCM1 and BCM4), rice (OsG), maize (GRMZM2G005859), jute (OMO77278), and Cephalotus follicularis (GAV69089) (http://www.ncbi.nlm.nih.gov/). b Relative expression of SlRCM1 in different tissues of Alisa Craig (AC) by qRT-PCR. RNA was extracted from the roots, stems, leaves, flowers, immature green fruits (IMG), mature green fruits (MG), breaker-stage fruits (BR) and red ripe fruits (RR). The ACTIN gene (Solyc11g005330) was used as the internal control. The data are presented as the means ± SDs (n = 3). c The SlRCM1 promoter drives GUS gene expression in AC. Tissues of transgenic tomato plants were stained with X-Gluc. d Subcellular localization of the SlRCM1 protein. SlRCM1::GFP fusion proteins were transiently expressed in tobacco leaves. Chloroplasts exhibit red fluorescence under a confocal microscope. The arrows highlight the positions of the red and green fluorescence. Scale bars, 25 μm
To investigate the expression pattern of SlRCM1, quantitative reverse transcription polymerase chain reaction (qRT-PCR) was performed as described previously32. SlRCM1 expression was detected in all tissues, and relatively high expression levels were detected in the leaves, flowers and immature green fruits (Fig. 6b). Furthermore, we used GUS staining to study the expression of SlRCM1 in several tissues. The GUS staining revealed that SlRCM1 was expressed in the stems, leaves, flowers and fruits (Fig. 6c).
To determine the SlRCM1 subcellular localization, we constructed a SlRCM1::GFP fusion protein. Through Agrobacterium-mediated infiltration, the SlRCM1::GFP fusion protein was transiently expressed in tobacco. Under a Nikon e600 fluorescence microscope (Nikon, Tokyo, Japan), the autofluorescence of the chloroplast appeared red33. The SlRCM1::GFP fluorescence overlapped with the red fluorescence, indicating that the SlRCM1 protein is localized in the chloroplast (Fig. 6d).
The expression level of SlRCM1 is directly regulated by the transcription factor SlARF2A
It was previously reported that ARF proteins regulate gene expression by binding to the TGTCTC cis-element of the target gene promoter34. Yeast one-hybrid (Y1H) and dual luciferase experiments indicated that SlARF2A binds to cis-elements of the SlRCM1 promoter. The TGTCTC cis-element is located -245 to -239 bp upstream of the start codon of SlRCM1 (Fig. 7a). Yeast cells that were cotransformed with pGADT7-SlARF2A and pAbAi-SlRCM1-Pro were able to grow on 20 ng/mL aureobasidin A (ABA) SD/-Leu media, but the negative control cotransformed with pGADT7 and pAbAi-SlRCM1-Pro did not grow (Fig. 7b), indicating that the ARF2A protein could directly bind to the promoter of SlRCM1.
a Schematic representation of the SlRCM1 promoter and its cis-elements. The sequence in the black box refers to the TGTCTC cis-element. b Yeast one-hybrid experiment of SlARF2A protein binding to the promoter of SlRCM1. The complete open reading frame (ORF) of SlARF2A was cloned into pGADT7 (AD) to generate a pGADT7-SlARF2A effector construct. The reporter vector was introduced into yeast strain Y1Hgold together with the effector vector. A combination of the reporter vector and the empty vector pGADT7 was used as a negative control. The yeast cells were cultured on SD-Leu media supplemented with various concentrations of ABA (0, 10, and 20 ng/ml). c Schematic representation of constructs used for the dual luciferase assay. The SlARF2A ORF was cloned into pGreenII 62-SK to generate a pGreenII 62-SK-SlARF2A effector construct. The SlRCM1 promoter was fused to pGreenII 0800-LUC to create a pGreen II 0800-SlRCM1-Pro reporter construct. d Relative LUC/RLU ratio. The vectors of both the effectors and reporters were transformed into Agrobacterium cells, which were then used to infiltrate tobacco leaves. LUC, firefly luciferase activity; RLU, Renilla luciferase activity. The data are presented as the means ± SDs (n = 6). The asterisks indicate statistically significant differences according to t-tests: **, P < 0.01
To further indicate that the SlARF2A protein interacted with the promoter of SlRCM1 in plants, we constructed a luciferase (LUC) reporter harboring a promoter sequence from -1,478 bp to the start codon (ATG) derived from SlRCM1 (Fig. 7c). Tobacco leaves were coinfiltrated with Agrobacterium tumefaciens (GV3101) strains containing the indicated effector constructs containing SlARF2A and the reporter constructs containing the SlRCM1 promoter (Fig. 7d). Taken together, these results indicate that SlARF2A physically binds to the SlRCM1 promoter to downregulate its expression.
Discussion
Chlorophyll is essential for plants to absorb, transfer and convert light energy to bioenergy and plays a vital role in photosynthesis to promote plant growth and development35. Here, we cloned the SlRCM1 gene, which controls chlorophyll synthesis and chloroplast development in tomato fruits. A single-nucleotide replacement resulting in a premature termination mutation in SlRCM1 impaired chloroplast development in the rcm1 mutant. Increasing the chlorophyll content in tomato fruits contributes to improved nutrition36. Consistently, the soluble solids content of red ripe fruits of SlRCM1-overexpressing (OE) lines was significantly higher than that of the wild type, whereas the knockout lines (CR) showed the opposite effect (Fig. S7). In addition, the a*/b* colorimetric value of the ripe red fruits of the SlRCM1-overexpressing lines (OE) was significantly higher than that of the wild type, while the knockout lines (CR) showed the opposite effect (Fig. S8). Therefore, characterization of the SlRCM1 gene may provide insights into chlorophyll synthesis and chloroplast development in tomato fruits.
The rcm1 mutant was obtained from an EMS-mutagenized population. EMS mutagenesis is widely used in the construction of mutant libraries and in plant functional genomics37,38. For instance, the hst1 mutant generated from EMS mutagenesis was crossed with the wild type to map the OsRR22 gene39. TILLING analysis40 of an EMS-mutagenized population of Arabidopsis showed that EMS mutagenesis produces a large number of single-base substitutions within hotspot segments41. In this study, a reduced-chlorophyll mutant, rcm1, was acquired by EMS mutagenesis and crossed with wild type to map the SlRCM1 gene responsible for the yellowish-fruit–producing mutant. Due to the close genetic background between the EMS-induced mutant and its wild type, crossing a mutant with its wild type is a fast and effective approach to isolate a gene42,43. Two SNPs in the recessive pool with an allele frequency of 1 were found between SL2.50ch08_1 and SL2.50ch08_1010000 (Table S2). Since the SNP at SL2.50ch08_ 394612 was located in an intergenic region, we speculated that it is not a causal SNP for the reduced chlorophyll phenotype of rcm1. The SNP (A → T) in the second exon of Solyc08g005010 resulted in a premature stop codon (Fig. 2e). Furthermore, the rcm1 mutant phenotype was rescued by overexpression of the SlRCM1 gene in the rcm1 mutant, which confirmed that SlRCM1 was responsible for the mutant phenotype (Fig. 3). MutMap analysis could therefore be a feasible approach to genetically identify genes in EMS-mutagenized mutants.
SlRCM1 has highly conserved functions in both chloroplast development and chlorophyll synthesis. There are two homologous genes of SlRCM1 in Arabidopsis: BCM1 and BCM222. SlRCM1 was found to share 78% amino acid sequence identity with BCM1 and 84% identity with BCM2 (Fig. S1). Previous studies have shown that BCM1 and BCM2 regulate chlorophyll synthesis and chloroplast development in Arabidopsis22. Yeast two-hybrid (Y2H) assays, bimolecular fluorescence complementation (BiFC) assays, coimmunoprecipitation (Co-IP) assays and enzyme activity-measuring experiments in Arabidopsis demonstrated that BCM1 interacts with GUN4 to stimulate Mg-chelatase activity and optimize chlorophyll synthesis22. BCM1 also interacts with SGR to prevent chlorophyll degradation22. SlRCM1’s ortholog in soybean is the stay-green G gene and regulates chlorophyll synthesis and chloroplast development in the seed coat of soybean25. However, biological divergence has shown that SlRCM1 regulates chlorophyll synthesis and chloroplast development in fruits, while its ortholog regulates chlorophyll synthesis in the seed coat of soybean25 and in the leaves of Arabidopsis22.
It has been reported that SlRCM1 orthologs modulate chlorophyll synthesis in the leaves of Arabidopsis and in the seed coat of soybean22,25. BCM1 overexpression did not alter the chlorophyll content in Arabidopsis thaliana, but chlorophyll accumulation was inhibited only in the leaves of the bcm1 mutant22. In the present study, fruits of the tomato SlRCM1 knockout lines showed a yellowish phenotype due to impaired chloroplasts. The chlorophyll content in the MG-stage fruits of the SlRCM1-overexpressing lines increased significantly. Moreover, chlorophyll was also detected in the RR-stage fruits of the SlRCM1 overexpression lines (Fig. 4). Taken together, these results showed that SlRCM1 has a diversified and strong effect on chlorophyll synthesis and chloroplast development in fruits.
In summary, we have demonstrated that SlRCM1, a chloroplast-targeted ortholog of the BCM1 protein in Arabidopsis and stay-green G protein of soybean, participates in chlorophyll synthesis and chloroplast development in tomato fruits. SlRCM1 modulates the number of thylakoids and the structure of thylakoid membranes in chloroplasts. SlRCM1 was identified as the causal gene at the Lutescent1 locus. An understanding of SlRCM1 provides insights into the molecular mechanism underlying fruit development and target genes for genetic improvement in horticultural crop species.
Materials and methods
Plant materials and mutant screening
A chlorophyll-deficient mutant (rcm1) was derived from EMS mutagenesis of the processed tomato inbred line Ligeer 87-5. For EMS mutagenesis, Ligeer 87-5 seeds were immersed in 1% EMS solution and shaken in a shaker for 12 h. The seeds were then rinsed with running water for 10 min and germinated in an artificial incubator at 30 °C. Mutants were screened in the M2 generation44. We constructed an F2 mapping population by crossing Ligeer 87-5 with rcm1. The l1 mutant (LA3717) in the AC background was obtained from the TGRC (https://tgrc.ucdavis.edu). For the overexpression constructs, SlRCM1 was amplified and cloned into a pHELLSGATE8 vector driven by the CaMV35S promoter and its native promoter45. The overexpression vector was introduced into Solanum lycopersicum cv. AC and the rcm1 and l1 mutants through Agrobacterium-mediated transformation. SlRCM1 knockout mutants in the AC background were generated using the CRISPR/Cas9 system46. The 2934-bp promoter of the SlRCM1 gene was amplified from AC and cloned into a pV3P vector driving β-glucosidase (GUS), yielding a ProSlRCM1::GUS vector, which was subsequently transformed into AC. The tomato tissues were quickly frozen in liquid nitrogen and stored at −80 °C. The tomato plants used in this study were grown in a greenhouse, and the primers used in this experiment are listed in Table S4.
Transmission electron microscopy
The fruits of Ligeer 87-5, the rcm1 mutant, AC and the transgenic lines were isolated and fixed in 0.05 M cacodylate buffer consisting of 2% glutaraldehyde and dehydrated in ethanol. After embedding in Spurr resin, ultrathin sections of the samples were obtained using a Leica EMUC6 ultramicrotome. A Hitachi H-7650 transmission electron microscope was used to observe the ultrastructure of the plastids.
Determination of the chlorophyll content
For chlorophyll extraction, fruits of Ligeer 87-5, the rcm1 mutant, AC and the transgenic lines at different developmental stages were placed in 10 mL of 80% (v/v) acetone in the dark until the tissues became white. Their absorbance was subsequently measured at 646, 663 and 470 nm, and the chlorophyll content was measured and calculated as previously reported47.
BSA and DNA-seq
An F2 population comprising 307 individuals was derived from a cross between Ligeer 87-5 and the rcm1 mutant. In this population, equal amounts of DNA were pooled from 25 plants with a wild-type phenotype and 25 plants with a mutant phenotype. Approximately 25× genome sequences for each pool were generated using the Illumina HiSeq X Ten platform48. Due to the close genetic background of the M82 and Ligeer 87-5 processed tomato genotypes, the M82 genome version SL 2.50 (http://solgenomics.net) was used as the reference genome to facilitate mining of the causal SNPs.
Subcellular localization
The SlRCM1 coding sequence without the stop codon was amplified from the cDNA of AC and then cloned into a pHBT vector driven by the CaMV35S promoter, yielding CaMV35S::SlRCM1-GFP. This vector was subsequently transformed into Agrobacterium tumefaciens strain GV3101, which was then injected into the leaves of Nicotiana benthamiana (N. benthamiana) as previously described49. After 48 h of incubation at 25 °C, the fluorescence of GFP and RFP in the tobacco leaves was observed using Leica Confocal software. Chloroplasts that exhibit red fluorescence were used as positive controls. The primers used in this experiment are listed in Table S4.
Quantitative reverse transcription PCR
Total RNA was extracted from frozen tissues using TRIzol reagent (Vazyme, Nanjing, China). RNA was then reverse transcribed into cDNA using a first-strand cDNA Synthesis Kit (Vazyme). The product length ranged from 80 to 200 bp. The qRT-PCR was used to determine the transcript levels of genes in 96-well plates with a Roche LightCycler® 480 system according to the manufacturer’s protocol32. The expression of the ACTIN gene (Solyc11g005330) was used as an internal control. The primers used were designed using Primer Premier 5 (Table S4).
Colorimeter-based evaluation of red ripe fruits
The a* and b* values of ripe red fruits were measured using a CM-5 colorimeter. Three independent points were determined for each fruit assay, and six fruits were measured per line.
Measurement of total soluble solids
The total soluble solids (Brix) of red ripe fruits were measured using a digital refractometer (PR100, Atago Co., Ltd.). All sample assays were performed for three technical replicates and six biological replicates.
Ethylene assays
Tomato fruits were collected at 37, 43 and 49 days after flowering. The determination of ethylene content in the tomato fruits was based on previous methods50. All the samples consisted of three technical replicates and three biological replicates.
Pollen viability assays
The pollen of blooming flowers was soaked in I-KI solution (1% KI and 0.5% I2) for 2 min. The stained pollen was then examined under a low-magnification microscope. The viable pollen was stained dark blue by I-KI.
Chlorophyll fluorescence measurements
After the leaves of Ligeer 87-5, the rcm1 mutant, AC and L1-CR were conditioned in the dark for 30 min, an imaging pulse amplitude modulated chlorophyll fluorimeter (IMAG-MAXI; Heinz Walz, Effeltrich, Germany) was used to determine chlorophyll fluorescence. The maximal photochemical efficiency of PSII (Fv/Fm) and the quantum efficiency of PSII (Y(II)) were calculated and determined according to previously reported methods51.
Yeast one-hybrid assays
The SlARF2A ORF was cloned into a pGADT7 vector to yield a prey construct. The SlRCM1 promoter was inserted into a pAbAi vector to yield a bait construct. The pAbAi bait vector was then used to transform yeast strain Y1HGold, which was integrated into the yeast genome to generate reporter strains. The prey vector was introduced into the reporter strains and grown for three days on SD/-Leu-Ura media. The positive yeast strains were selected and diluted in double-distilled water to an OD600 of 0.1, and 2 μL of the suspension was spotted onto SD/–Leu media with or without ABA (0, 10 and 20 ng/mL), followed by 3 days of incubation at 30 °C. pGADT7 and pAbAi-PSY1-Pro served as negative controls. The primers used in this experiment are listed in Table S4.
Dual luciferase transactivation assays
To generate an effector construct, the full-length ARF2A ORF was inserted into a pGreenII 62-SK vector. Similarly, the promoter from SlRCM1 was inserted into pGreenII 0800-LUC to yield a reporter construct. The constructs and the pSoup helper plasmid were simultaneously introduced into Agrobacterium tumefaciens (GV3101). Tobacco leaves were infiltrated with the Agrobacterium strains and harvested three days later. The firefly LUC and Renilla luciferase (RLU) activities were quantified using a dual-luciferase reporter assay system. The transactivation activities were expressed as the ratio of LUC to RLU activity. The primers used in this experiment are listed in Table S4.
References
Almeida, J., Perez-Fons, L. & Fraser, P. D. A transcriptomic, metabolomic and cellular approach to the physiological adaptation of tomato fruit to high temperature. Plant Cell Environ. 13854, 1–9 (2020).
Wang, A. Q. et al. The tomato HIGH PIGMENT1/DAMAGED DNA BINDING PROTEIN 1 gene contributes to regulation of fruit ripening. Hortic. Res. 6, 15 (2019).
Bruno, A. K. & Wetzel, C. M. The early light-inducible protein (ELIP) gene is expressed during the chloroplast-to-chromoplast transition in ripening tomato fruit. J. Exp. Bot. 55, 2541–2548 (2004).
Powell, A. L. et al. Uniform ripening encodes a Golden 2-like transcription factor regulating tomato fruit chloroplast development. Science 336, 1711–1715 (2012).
Nguyen, C. V. et al. Tomato GOLDEN2-LIKE transcription factors reveal molecular gradients that function during fruit development and ripening. Plant Cell 26, 585–601 (2014).
Nadakuduti, S. S., Holdsworth, W. L., Klein, C. L. & Barry, C. S. KNOX genes influence a gradient of fruit chloroplast development through regulation of GOLDEN2-LIKE expression in tomato. Plant J. 78, 1022–1033 (2014).
Pan, Y. et al. Network inference analysis identifies an APRR2-like gene linked to pigment accumulation in tomato and pepper fruits. Plant Physiol. 161, 1476–1485 (2013).
Xiong, C. et al. A tomato B-box protein SlBBX20 modulates carotenoid biosynthesis by directly activating PHYTOENE SYNTHASE 1, and is targeted for 26S proteasome-mediated degradation. N. Phytol. 221, 279–294 (2019).
Meng, L. et al. BEL1-LIKE HOMEODOMAIN 11 regulates chloroplast development and chlorophyll synthesis in tomato fruit. Plant J. 94, 1126–1140 (2018).
Yuan, Y. et al. SlARF10, an auxin response factor, is involved in chlorophyll and sugar accumulation during tomato fruit development. J. Exp. Bot. 69, 5507–5518 (2018).
Yuan, Y. et al. Auxin response factor 6A regulates photosynthesis, sugar accumulation, and fruit development in tomato. Hortic. Res. 6, 85 (2019).
Lim, P. O. et al. Auxin response factor 2 (ARF2) plays a major role in regulating auxin-mediated leaf longevity. J. Exp. Bot. 61, 1419–1430 (2010).
Breitel, D. A. et al. AUXIN RESPONSE FACTOR 2 intersects hormonal signals in the regulation of tomato fruit ripening. Plos Genet. 12, e1005903 (2016).
Beale, S. I. Green genes gleaned. Trends Plant Sci. 10, 309–312 (2005).
Axelsson, E. et al. Recessiveness and dominance in barley mutants deficient in Mg-chelatase subunit D, an AAA protein involved in chlorophyll biosynthesis. Plant Cell 18, 3606–3616 (2006).
Alawady, A., Reski, R., Yaronskaya, E. & Grimm, B. Cloning and expression of the tobacco CHLM sequence encoding Mg protoporphyrin IX methyltransferase and its interaction with Mg chelatase. Plant Mol. Biol. 57, 679–691 (2005).
Biswal, A. K. et al. Light intensity-dependent modulation of chlorophyll b biosynthesis and photosynthesis by overexpression of chlorophyllide a oxygenase in tobacco. Plant Physiol. 159, 433–449 (2012).
Richter, A. S. et al. Phosphorylation of GENOMES UNCOUPLED 4 alters stimulation of Mg chelatase activity in angiosperms. Plant Physiol. 172, 1578–1595 (2016).
Peter, E. & Grimm, B. GUN4 is required for posttranslational control of plant tetrapyrrole biosynthesis. Mol. Plant 2, 1198–1210 (2009).
Hao, N. et al. CsMYB36 is involved in the formation of yellow green peel in cucumber (Cucumis sativus L.). Theor. Appl. Genet. 131, 1659–1669 (2018).
Wang, N. et al. Defect in Brnym1, a magnesium-dechelatase protein, causes a stay-green phenotype in an EMS-mutagenized Chinese cabbage (Brassica campestris L. ssp. pekinensis) line. Hortic. Res. 7, 8 (2020).
Wang, P., Richter, A. S., Kleeberg, J. R. W., Geimer, S. & Grimm, B. Post-translational coordination of chlorophyll biosynthesis and breakdown by BCMs maintains chlorophyll homeostasis during leaf development. Nat. Commun. 11, 1254 (2020).
Czarnecki, O. et al. An Arabidopsis GluTR binding protein mediates spatial separation of 5-aminolevulinic acid synthesis in chloroplasts. Plant Cell 23, 4476–4491 (2011).
Pontier, D., Albrieux, C., Joyard, J., Lagrange, T. & Block, M. A. Knock-out of the magnesium protoporphyrin IX methyltransferase gene in Arabidopsis effects on chloroplast development and on chloroplast-to-nucleus signaling. J. Biol. Chem. 282, 2297–2304 (2007).
Wang, M. et al. Parallel selection on a dormancy gene during domestication of crops from multiple families. Nat. Genet. 50, 1435–1441 (2018).
Barry, C. S. et al. Altered chloroplast development and delayed fruit ripening caused by mutations in a zinc metalloprotease at the lutescent2 locus of tomato. Plant Physiol. 159, 1086–1098 (2012).
Almeida, J. et al. Fruits from ripening impaired, chlorophyll degraded and jasmonate insensitive tomato mutants have altered tocopherol content and composition. Phytochemistry 111, 72–83 (2015).
Mizzotti, C. et al. Time-course transcriptome analysis of arabidopsis siliques discloses genes essential for fruit development and maturation. Plant Physiol. 178, 1249–1268 (2018).
Menda, N., Semel, Y., Peled, D., Eshed, Y. & Zamir, D. In silico screening of a saturated mutation library of tomato. Plant J. 38 (2004).
Saito, T. et al. TOMATOMA: a novel tomato mutant database distributing Micro-Tom mutant collections. Plant Cell Physiol. 52, 283–296 (2011).
Tanksley, S. D. et al. High density molecular linkage maps of the tomato and potato genomes. Genetics 132, 1141–1160 (1992).
Xie, Q. et al. The HD-Zip IV transcription factor SlHDZIV8 controls multicellular trichome morphology by regulating the expression of Hairless-2. J. Exp. Bot. (2020).
Bobik, K. et al. The essential chloroplast ribosomal protein uL15c interacts with the chloroplast RNA helicase ISE2 and affects intercellular trafficking through plasmodesmata. N. Phytol. 221, 850–865 (2019).
Yuan, Y. J. et al. SIARF10, an auxin response factor, is involved in chlorophyll and sugar accumulation during tomato fruit development. J. Exp. Bot. 69, 5507–5518 (2018).
Croce, R. & van Amerongen, H. Natural strategies for photosynthetic light harvesting. Nat. Chem. Biol. 10, 492–501 (2014).
Cocaliadis, M. F., Fernandez-Munoz, R., Pons, C., Orzaez, D. & Granell, A. Increasing tomato fruit quality by enhancing fruit chloroplast function. A double-edged sword? Exp. Bot. 65, 4589–4598 (2014).
Henry, I. M. et al. Efficient genome-wide detection and cataloging of EMS-induced mutations using exome capture and next-generation sequencing. Plant Cell 26, 1382–1397 (2014).
Sashidhar, N., Harloff, H. J. & Jung, C. Identification of phytic acid mutants in oilseed rape (Brassica napus) by large-scale screening of mutant populations through amplicon sequencing. N. Phytol. 225, 2022–2034 (2020).
Takagi, H. et al. MutMap accelerates breeding of a salt-tolerant rice cultivar. Nat. Biotechnol. 33, 445–449 (2015).
Lakhssassi, N. et al. Soybean TILLING-by-Sequencing reveals the role of novel GmSACPD members in the unsaturated fatty acid biosynthesis while maintaining healthy nodules. J. Exp. Bot. 71, 6969–6987 (2020).
Greene, E. A. et al. Spectrum of chemically induced mutations from a large-scale reverse-genetic screen in Arabidopsis. Genetics 164, 731–740 (2003).
Abe, A. et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 30, 174–178 (2012).
Fekih, R. et al. MutMap plus: genetic mapping and mutant identification without crossing in rice. PLoS ONE 8, e68529 (2013).
Saito, T. et al. TOMATOMA: a novel tomato mutant database distributing micro-tom mutant collections. Plant Cell Physiol. 52, 283–296 (2011).
Chang, J. et al. Hair, encoding a single C2H2 zinc-finger protein, regulates multicellular trichome formation in tomato. Plant J. 96, 90–102 (2018).
Cui, L. et al. miR156a-targeted SBP-Box transcription factor SlSPL13 regulates inflorescence morphogenesis by directly activating SFT in tomato. Plant Biotechnol. J. 18, 1670–1682 (2020).
Lichtenthaler, H. K. Chlorophylls and carotenoids-pigments of photosynthetic biomembranes. Method. Enzymol. 148, 350–382 (1987).
Meynert, A. M., Ansari, M., FitzPatrick, D. R. & Taylor, M. S. Variant detection sensitivity and biases in whole genome and exome sequencing. Bmc Bioinformatics 15, 247 (2014).
Tian, Z. et al. The potato ERF transcription factor StERF3 negatively regulates resistance to phytophthora infestans and salt tolerance in potato. Plant Cell Physiol. 56, 992–1005 (2015).
Vrebalov, J. et al. A MADS-box gene necessary for fruit ripening at the tomato ripening-inhibitor (Rin) locus. Science 296, 343–346 (2002).
Li, H. et al. Hydrogen peroxide mediates abscisic acid-induced HSP70 accumulation and heat tolerance in grafted cucumber plants. Plant Cell Environ. 37, 2768–2780 (2014).
Acknowledgements
We thank Dr. John K. Ahiakpa for his careful reading of the manuscript. This work was supported by grants from the National Key Research and Development Program of China (2018YFD1000800), the National Natural Science Foundation of China (31991182; 31972426), the Wuhan Frontier Projects for Applied Foundation (2019020701011492), the Fundamental Research Funds for the Central Universities (2662018PY073), and the Hubei Provincial Natural Science Foundation of China (2019CFA017).
Author information
Authors and Affiliations
Contributions
Y.Z. and Z.Y. planned and designed the study. G.L., H.Y., L.Y., C.L., J.Y., W.C., Y.W. and P.G. performed the experiments, conducted the fieldwork, and analyzed the data. J.Z. and Y.Z. provided suggestions for experiments. G.L. wrote the manuscript. Y.Z. supervised the project and revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, G., Yu, H., Yuan, L. et al. SlRCM1, which encodes tomato Lutescent1, is required for chlorophyll synthesis and chloroplast development in fruits. Hortic Res 8, 128 (2021). https://doi.org/10.1038/s41438-021-00563-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-021-00563-6
This article is cited by
-
Genome-wide identification of GA2ox genes family and analysis of PbrGA2ox1-mediated enhanced chlorophyll accumulation by promoting chloroplast development in pear
BMC Plant Biology (2024)
-
Melon yellow-green plant (Cmygp) encodes a Golden2-like transcription factor regulating chlorophyll synthesis and chloroplast development
Theoretical and Applied Genetics (2023)
-
Introgression of the sesquiterpene biosynthesis from Solanum habrochaites to cultivated tomato offers insights into trichome morphology and arthropod resistance
Planta (2021)