Abstract
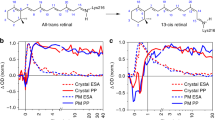
The activation of rhodopsin, the light-sensitive G-protein-coupled receptor responsible for dim-light vision in vertebrates, is driven by an ultrafast excited-state double-bond isomerization with a quantum efficiency of almost 70%. The origin of such light sensitivity is not understood and a key question is whether in-phase nuclear motion controls the quantum efficiency value. In this study we used hundreds of quantum–classical trajectories to show that, 15 fs after light absorption, a degeneracy between the reactive excited state and a neighbouring state causes the splitting of the rhodopsin population into subpopulations. These subpopulations propagate with different velocities and lead to distinct contributions to the quantum efficiency. We also show here that such splitting is modulated by protein electrostatics, thus linking amino acid sequence variations to quantum efficiency modulation. Finally, we discuss how such a linkage that in principle could be exploited to achieve higher quantum efficiencies would simultaneously increase the receptor thermal noise leading to a trade-off that may have played a role in rhodopsin evolution.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the main article and the Supplementary Information. Cartesian coordinates generated along the trajectories can be found at https://doi.org/10.5281/zenodo.5826280. Source data are provided with this paper.
Code availability
The authors declare that the present research has been produced with distributed software available to the public, as also detailed in the Supplementary Information.
References
Ernst, O. P. et al. Microbial and animal rhodopsins: structures, functions, and molecular mechanisms. Chem. Rev. 114, 126–163 (2014).
Khorana, H. G. Rhodopsin, photoreceptor of the rod cell. An emerging pattern for structure and function. J. Biol. Chem. 267, 1–4 (1992).
Peteanu, L. A., Schoenlein, R. W., Wang, Q., Mathies, R. A. & Shank, C. V. The first step in vision occurs in femtoseconds: complete blue and red spectral studies. Proc. Natl Acad. Sci. USA 90, 11762–11766 (1993).
Polli, D. et al. Conical intersection dynamics of the primary photoisomerization event in vision. Nature 467, 440–443 (2010).
Gozem, S., Schapiro, I., Ferre, N. & Olivucci, M. The molecular mechanism of thermal noise in rod photoreceptors. Science 337, 1225–1228 (2012).
Dartnall, H. J. A. The photosensitivities of visual pigments in the presence of hydroxylamine. Vision Res. 8, 339–358 (1968).
Wang, Q., Schoenlein, R., Peteanu, L., Mathies, R. & Shank, C. Vibrationally coherent photochemistry in the femtosecond primary event of vision. Science 266, 422–424 (1994).
Kukura, P. Structural observation of the primary isomerization in vision with femtosecond-stimulated Raman. Science 310, 1006–1009 (2005).
Johnson, P. J. M. et al. Local vibrational coherences drive the primary photochemistry of vision. Nat. Chem. 7, 980–986 (2015).
Mathies, R. A. A coherent picture of vision. Nat. Chem. 7, 945–947 (2015).
Schapiro, I. et al. The ultrafast photoisomerizations of rhodopsin and bathorhodopsin are modulated by bond length alternation and HOOP driven electronic effects. J. Am. Chem. Soc. 133, 3354–3364 (2011).
Gozem, S., Luk, H. L., Schapiro, I. & Olivucci, M. Theory and simulation of the ultrafast double-bond isomerization of biological chromophores. Chem. Rev. 117, 13502–13565 (2017).
Schnedermann, C. et al. Evidence for a vibrational phase-dependent isotope effect on the photochemistry of vision. Nat. Chem. 10, 449–455 (2018).
Weingart, O. et al. Product formation in rhodopsin by fast hydrogen motions. Phys. Chem. Chem. Phys. 13, 3645–3648 (2011).
Sen, S. & Schapiro, I. A comprehensive benchmark of the XMS-CASPT2 method for the photochemistry of a retinal chromophore model. Mol. Phys. 116, 2571–2582 (2018).
Gozem, S. et al. Mapping the excited state potential energy surface of a retinal chromophore model with multireference and equation-of-motion coupled-cluster methods. J. Chem. Theory Comput. 9, 4495–4506 (2013).
Andersson, K., Malmqvist, P. A., Roos, B. O., Sadlej, A. J. & Wolinski, K. Second-order perturbation theory with a CASSCF reference function. J. Phys. Chem. 94, 5483–5488 (1990).
Luk, H. L., Melaccio, F., Rinaldi, S., Gozem, S. & Olivucci, M. Molecular bases for the selection of the chromophore of animal rhodopsins. Proc. Natl Acad. Sci. USA 112, 15297–15302 (2015).
Granovsky, A. A. Firefly, v.8.0.0 (Moscow State University, 2013).
Warshel, A., Chu, Z. T. & Hwang, J. K. The dynamics of the primary event in rhodopsins revisited. Chem. Phys. 158, 303–314 (1991).
Lim, J. S. & Kim, S. K. Experimental probing of conical intersection dynamics in the photodissociation of thioanisole. Nat. Chem. 2, 627–632 (2010).
Warshel, A. & Chu, Z. T. Nature of the surface crossing process in bacteriorhodopsin: computer simulations of the quantum dynamics of the primary photochemical event. J. Phys. Chem. B 105, 9857–9871 (2001).
Toniolo, A., Olsen, S., Manohar, L. & Martínez, T. J. Conical intersection dynamics in solution: the chromophore of green fluorescent protein. Faraday Discuss. 127, 149–163 (2004).
Park, J. W. & Rhee, Y. M. Electric field keeps chromophore planar and produces high yield fluorescence in green fluorescent protein. J. Am. Chem. Soc. 138, 13619–13629 (2016).
Romei, M. G., Lin, C. Y., Mathews, I. I. & Boxer, S. G. Electrostatic control of photoisomerization pathways in proteins. Science 367, 76–79 (2020).
Gozem, S. et al. Excited-state vibronic dynamics of bacteriorhodopsin from two-dimensional electronic photon echo spectroscopy and multiconfigurational quantum chemistry. J. Phys. Chem. Lett. 11, 3889–3896 (2020).
Zener, C. Non-adiabatic crossing of energy levels. Proc. R. Soc. Lond. A 137, 696–702 (1932).
Gai, F., Hasson, K. C., McDonald, J. C. & Anfinrud, P. A. Chemical dynamics in proteins: the photoisomerization of retinal in bacteriorhodopsin. Science 279, 1886–1891 (1998).
Albert, J., Hader, K. & Engel, V. Coupled electron-nuclear quantum dynamics through and around a conical intersection. J. Chem. Phys. 147, 064302 (2017).
Olivucci, M., Tran, T., Worth, G. A. & Robb, M. A. Unlocking the double bond in protonated Schiff bases by coherent superposition of S1 and S2. J. Phys. Chem. Lett. 12, 5639–5643 (2021).
Xie, C., Malbon, C. L., Guo, H. & Yarkony, D. R. Up to a sign. The insidious effects of energetically inaccessible conical intersections on unimolecular reactions. Acc. Chem. Res. https://doi.org/10.1021/acs.accounts.8b00571 (2019).
Frutos, L. M., Andruniow, T., Santoro, F., Ferre, N. & Olivucci, M. Tracking the excited-state time evolution of the visual pigment with multiconfigurational quantum chemistry. Proc. Natl Acad. Sci. USA 104, 7764–7769 (2007).
Warshel, A. Bicycle-pedal model for the first step in the vision process. Nature 260, 679–683 (1976).
Miller, W. H. Perspective: quantum or classical coherence. J. Chem. Phys. 136, 210901 (2012).
Lin, S. W. et al. Vibrational assignment of torsional normal modes of rhodopsin: probing excited-state isomerization dynamics along the reactive C11=C12 torsion coordinate. J. Phys. Chem. B 102, 2787–2806 (1998).
Jarzȩcki, A. A. & Spiro, T. G. Porphyrin distortion from resonance Raman intensities of out-of-plane modes: computation and modeling of N-methylmesoporphyrin, a ferrochelatase transition state analog. J. Phys. Chem. A 109, 421–430 (2005).
Okada, T. et al. The retinal conformation and its environment in rhodopsin in light of a new 2.2 Å crystal structure. J. Mol. Biol. 342, 571–583 (2004).
Cornell, W. D. et al. A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 118, 2309 (1996).
Ferré, N., Cembran, A., Garavelli, M. & Olivucci, M. Complete-active-space self-consistent-field/Amber parameterization of the Lys296–retinal–Glu113 rhodopsin chromophore-counterion system. Theor. Chem. Acc. 112, 335–341 (2004).
Aquilante, F. et al. Molcas 8: new capabilities for multiconfigurational quantum chemical calculations across the periodic table. J. Comput. Chem. 37, 506–541 (2016).
Ponder, J. W. & Richards, F. M. Tinker molecular modeling package. J. Comput. Chem. 8, 1016–1024 (1987).
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013).
Granucci, G. & Persico, M. Critical appraisal of the fewest switches algorithm for surface hopping. J. Chem. Phys. 126, 134114 (2007).
Acknowledgements
We thank R. A. Mathies for directing our attention towards the Rh resonance Raman spectrum, B. Chang for suggesting the idea of a trade-off and S. Haacke for helpful discussions. We also thank N. Ferrè for the frequency calculation code in Molcas/Tinker and L. Barneschi for help in generating the plots in Fig. 5. The research was partially supported by the grants NSF CHE-CLP-1710191, NIH GM126627-01 and USIAS 2015, the Ohio Supercomputer Center, the Ministero dell’Istruzione, dell’Università e della Ricerca (MIUR) for a ‘Dipartimento di Eccellenza 2018–2022’, the Fondazione Banca d’Italia (to M.O.), the Interdisciplinary Thematic Institute QMat (as part of the ITI 2021–2028 program of the University of Strasbourg), the CNRS and Inserm via the IdEx Unistra (ANR 10 IDEX 0002), SFRI STRAT’US (ANR 20 SFRI 0012) and Labex NIE (ANR-11-LABX-0058_NIE) projects of the French Investments for the Future Program (to J.L.).
Author information
Authors and Affiliations
Contributions
M.O. and X.Y. designed the study. X.Y. carried out the molecular dynamics simulations. T.A. contributed to the resonance Raman spectra simulation. X.Y., M.M., J.L. and S.G. analysed the data. M.O. and X.Y. wrote the paper. All authors discussed and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Peer review
Peer review information
Nature Chemistry thanks Todd Martinez, Young Min Rhee and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Non-adiabatic coupling evolution.
Non-adiabatic coupling evolution. Time evolution of the magnitude of the S2/S1 NADC modulus (see color legend) along a 3D cut of the S1 PES. The cut is represented by four 2D cross-sections corresponding to different α values and spanning the δop and BLA coordinates.
Supplementary information
Supplementary information
Supplementary Discussion (Sections 1–17), Tables 1 and 2, Figs. 1–22 and input for GROMACS and Molcas/Tinker molecular dynamics.
Supplementary Data 1
Equilibrium structure of the rhodopsin QM/MM model in Cartesian coordinates.
Source data
Source Data Fig. 2
Source data for Fig. 2a–d.
Source Data Fig. 3
Source data for Fig. 3a–f.
Source Data Fig. 4
Source data for Fig. 4a–e.
Source Data Fig. 5
Source data for Fig. 5a–b.
Source Data Extended Data Fig. 1
Statistical data for Extended Data Fig. 1.
Rights and permissions
About this article
Cite this article
Yang, X., Manathunga, M., Gozem, S. et al. Quantum–classical simulations of rhodopsin reveal excited-state population splitting and its effects on quantum efficiency. Nat. Chem. 14, 441–449 (2022). https://doi.org/10.1038/s41557-022-00892-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-022-00892-6