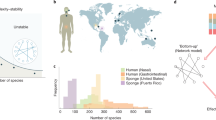
Abstract
Organisms—especially microbes—tend to live together in ecosystems. While some of these ecosystems are very biodiverse, others are not, and while some are very stable over time, others undergo strong temporal fluctuations. Despite a long history of research and a plethora of data, it is not fully understood what determines the biodiversity and stability of ecosystems. Theory and experiments suggest a connection between species interaction, biodiversity and the stability of ecosystems, where an increase in ecosystem stability with biodiversity could be observed in several cases. However, what causes these connections remains unclear. Here, we show in microbial ecosystems in the laboratory that the concentrations of available nutrients can set the strength of interactions between bacteria. High nutrient concentrations allowed the bacteria to strongly alter the chemical environment, causing on average more negative interactions between species. These stronger interactions excluded more species from the community, resulting in a loss of biodiversity. At the same time, the stronger interactions also decreased the stability of the microbial communities, providing a mechanistic link between species interaction, biodiversity and stability in microbial ecosystems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data and sequencing raw data are available at https://doi.org/10.5061/dryad.vdncjsxq9.
Code availability
The code for the simulations is available at https://github.com/cratzke/Interaction-biodiversity-stability.
References
Faust, K. & Raes, J. Microbial interactions: from networks to models. Nat. Rev. Microbiol. 10, 538–550 (2012).
Fuhrman, J. A. Microbial community structure and its functional implications. Nature 459, 193–199 (2009).
Ley, R. E., Peterson, D. A. & Gordon, J. I. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124, 837–848 (2006).
Thompson, L. R. et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Raes, J. & Bork, P. Molecular eco-systems biology: towards an understanding of community function. Nat. Rev. Microbiol. 6, 693–699 (2008).
Strom, S. L. Microbial ecology of ocean biogeochemistry: a community perspective. Science 320, 1043–1045 (2008).
Ghoul, M. & Mitri, S. The ecology and evolution of microbial competition. Trends Microbiol. 24, 833–845 (2016).
Moons, P., Michiels, C. W. & Aertsen, A. Bacterial interactions in biofilms. Crit. Rev. Microbiol. 35, 157–168 (2009).
James, G. A., Beaudette, L. & Costerton, J. W. Interspecies bacterial interactions in biofilms. J. Ind. Microbiol. 15, 257–262 (1995).
Friedman, J., Higgins, L. M. & Gore, J. Community structure follows simple assembly rules in microbial microcosms. Nat. Ecol. Evol. 1, 0109 (2017).
Momeni, B., Xie, L. & Shou, W. Lotka–Volterra pairwise modeling fails to capture diverse pairwise microbial interactions. eLife 6, e25051 (2017).
Hibbing, M. E., Fuqua, C., Parsek, M. R. & Peterson, S. B. Bacterial competition: surviving and thriving in the microbial jungle. Nat. Rev. Micro. 8, 15–25 (2010).
Riley, M. A. & Gordon, D. M. The ecological role of bacteriocins in bacterial competition. Trends Microbiol. 7, 129–133 (1999).
Pande, S. et al. Fitness and stability of obligate cross-feeding interactions that emerge upon gene loss in bacteria. ISME J. 8, 953–962 (2014).
Goldford, J. E. et al. Emergent simplicity in microbial community assembly. Science 361, 469–474 (2018).
Pande, S. & Kost, C. Bacterial unculturability and the formation of intercellular metabolic networks. Trends Microbiol. 25, 349–361 (2017).
Estrela, S. et al. Environmentally mediated social dilemmas. Trends Ecol. Evol. 34, 6–18 (2019).
Ratzke, C. & Gore, J. Modifying and reacting to the environmental pH can drive bacterial interactions. PLoS Biol. 16, e2004248 (2018).
Ratzke, C., Denk, J. & Gore, J. Ecological suicide in microbes. Nat. Ecol. Evol. 2, 867–872 (2018).
Niehaus, L. et al. Microbial coexistence through chemical-mediated interactions. Nat. Commun. 10, 2052 (2019).
Pacheco, A. R., Moel, M. & Segrè, D. Costless metabolic secretions as drivers of interspecies interactions in microbial ecosystems. Nat. Commun. 10, 103 (2019).
Basan, M. et al. Overflow metabolism in Escherichia coli results from efficient proteome allocation. Nature 528, 99–104 (2015).
May, R. M. Will a large complex system be stable? Nature 238, 413–414 (1972).
McCann, K., Hastings, A. & Huxel, G. R. Weak trophic interactions and the balance of nature. Nature 395, 794–798 (1998).
Tilman, D. & Downing, J. A. Biodiversity and stability in grasslands. Nature 367, 363–365 (1994).
Ives, A. R. & Carpenter, S. R. Stability and diversity of ecosystems. Science 317, 58–62 (2007).
Ptacnik, R. et al. Diversity predicts stability and resource use efficiency in natural phytoplankton communities. Proc. Natl Acad. Sci. USA 105, 5134–5138 (2008).
Butchart, S. H. M. et al. Global biodiversity: indicators of recent declines. Science 328, 1164–1168 (2010).
Jones, G. P., McCormick, M. I., Srinivasan, M. & Eagle, J. V. Coral decline threatens fish biodiversity in marine reserves. Proc. Natl Acad. Sci. USA 101, 8251–8253 (2004).
Newbold, T. et al. Has land use pushed terrestrial biodiversity beyond the planetary boundary? A global assessment. Science 353, 288–291 (2016).
Pennisi, E. What determines species diversity? Science 309, 90 (2005).
May, R. M. How many species are there on Earth? Science 241, 1441–1449 (1988).
Wardle, D. A. The influence of biotic interactions on soil biodiversity. Ecol. Lett. 9, 870–886 (2006).
Mougi, A. & Kondoh, M. Diversity of interaction types and ecological community stability. Science 337, 349–351 (2012).
Diamond, J. Factors controlling species diversity: overview and synthesis. Ann. Missouri Bot. Garden 75, 117–129 (1988).
Litchman, E., de Tezanos Pinto, P., Klausmeier, C. A., Thomas, M. K. & Yoshiyama, K. in Fifty Years After the “Homage to Santa Rosalia”: Old and New Paradigms on Biodiversity in Aquatic Ecosystems (eds Naselli-Flores, L. & Rossetti, G.) 15–28 (Springer, 2010).
Volkov, I., Banavar, J. R., Hubbell, S. P. & Maritan, A. Inferring species interactions in tropical forests. Proc. Natl Acad. Sci. USA 106, 13854–13859 (2009).
Carrara, F., Giometto, A., Seymour, M., Rinaldo, A. & Altermatt, F. Inferring species interactions in ecological communities: a comparison of methods at different levels of complexity. Methods Ecol. Evol. 6, 895–906 (2015).
Hautier, Y., Niklaus, P. A. & Hector, A. Competition for light causes plant biodiversity loss after eutrophication. Science 324, 636–638 (2009).
Conley, D. J. et al. Controlling eutrophication: nitrogen and phosphorus. Science 323, 1014–1015 (2009).
Smith, V. H. & Schindler, D. W. Eutrophication science: where do we go from here? Trends Ecol. Evol. 24, 201–207 (2009).
Smith, V. H., Joye, S. B. & Howarth, R. W. Eutrophication of freshwater and marine ecosystems. Limnol. Oceanogr. 51, 351–355 (2006).
Hillebrand, H. & Sommer, U. Diversity of benthic microalgae in response to colonization time and eutrophication. Aquat. Bot. 67, 221–236 (2000).
Moksness, E., Dahl, E. & Støttrup, J. Integrated Coastal Zone Management (John Wiley & Sons, 2009).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Filippo, C. D. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl Acad. Sci. USA 107, 14691–14696 (2010).
Segata, N. Gut microbiome: westernization and the disappearance of intestinal diversity. Curr. Biol. 25, R611–R613 (2015).
Vangay, P. et al. US immigration westernizes the human gut microbiome. Cell 175, 962–972.e10 (2018).
Deehan, E. C. & Walter, J. The fiber gap and the disappearing gut microbiome: implications for human nutrition. Trends Endocrinol. Metab. 27, 239–242 (2016).
Zinöcker, M. K. & Lindseth, I. A. The western diet–microbiome-host interaction and its role in metabolic disease. Nutrients 10, 365 (2018).
Turnbaugh, P. J., Bäckhed, F., Fulton, L. & Gordon, J. I. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 3, 213–223 (2008).
Hector, A. et al. Plant diversity and productivity experiments in European grasslands. Science 286, 1123–1127 (1999).
Pennekamp, F. et al. Biodiversity increases and decreases ecosystem stability. Nature 563, 109–112 (2018).
Bai, Y., Han, X., Wu, J., Chen, Z. & Li, L. Ecosystem stability and compensatory effects in the Inner Mongolia grassland. Nature 431, 181–184 (2004).
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012).
Loreau, M., Naeem, S. & Inchausti, P. Biodiversity and Ecosystem Functioning: Synthesis and Perspectives (Oxford Univ. Press, 2002).
Jessup, C. M. et al. Big questions, small worlds: microbial model systems in ecology. Trends Ecol. Evol. 19, 189–197 (2004).
Prosser, J. I. et al. The role of ecological theory in microbial ecology. Nat. Rev. Microbiol. 5, 384–392 (2007).
Friedman, J. & Gore, J. Ecological systems biology: the dynamics of interacting populations. Curr. Opin. Syst. Biol. 1, 114–121 (2017).
Higgins, L. M., Friedman, J., Shen, H. & Gore, J. Co-occurring soil bacteria exhibit a robust competitive hierarchy and lack of non-transitive interactions. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/175737v1 (2017).
Celiker, H. & Gore, J. Clustering in community structure across replicate ecosystems following a long-term bacterial evolution experiment. Nat. Commun. 5, 4643 (2014).
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J. & Holmes, S. P. Bioconductor workflow for microbiome data analysis: from raw reads to community analyses. F1000Res 5, 1492 (2016).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Acknowledgements
We thank D. Amor for help with analysing the sequencing data, and C. Abreu and the Gore group for reading and commenting on the manuscript. This work was funded by a NIH R01 (GM102311) grant.
Author information
Authors and Affiliations
Contributions
C.R., J.B. and J.G. designed the research. J.B. and C.R. carried out the experiments and performed the mathematical analysis. C.R., J.B. and J.G. discussed and interpreted the results, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Different soil strains have different suitable pH ranges.
We tested the optimal growth pH of 81 isolated soil species. It is a subset of the species shown in Extended Data Fig. 2B. All isolates were pre-cultured in 200µL of 1xNutrient medium for 24h at 25 °C with 1350 rpm shaking speed in 500-µl 96-deepwell plates (Eppendorf, Hauppauge, USA). After 24h of growth the cultures were diluted 1:100 into 500-µl 96-deepwell plates and a final volume of 200µl of Base media with 100mM phosphate with pH values of 3–11. Cultures were incubated for 24h at 25 °C at 1350 rpm on a Heidolph Titramax shaker. Population densities were estimated by CFU counting at the start of the experiment and after 24h, which allows to estimate the fold growth in 24h that is shown in the figure. Several example curves are shown in the upper panel. As can be seen those curves can have several shapes. For simplification, we decided to describe the shape of those curves with a heaviside function in our simulations (see below).
Extended Data Fig. 2 Nutrient concentrations and buffering determine pH change of growth media.
(a) The top and bottom panels show the same data as Fig. 1b. Using intermediate nutrient concentrations also causes intermediate pH shifts (green) compared to high (blue) and low (yellow) nutrient concentrations. Also adding higher concentrations of buffer lowers pH shifts (red) compared to the situation with low buffer (blue). (b) List of soil isolates that were used to measure the data in main text Fig. 1b and Extended Data Fig 1A and 2B. Strains were identified down to genus level by sequencing their 16S rRNA gene and comparing it to the RDP database. The strains belong to a collection of soil strains that we used before for interaction studies1,4. As can be seen many of those strains belong to the genus Bacillus, nevertheless they can change the pH into alkaline or acidic directions. For some cases the sequencing failed which lead to empty entries. PO4 means phosphate.
Extended Data Fig. 3 Bacteria for the pairwise interaction experiments.
The different colony morphologies allowed to distinguish them after plating on agar plates.
Extended Data Fig. 4 High nutrient concentrations lead to stronger negative interactions between bacteria.
The figure shows all the data of main text Fig. 1c for low nutrient concentrations (top) and high nutrient concentrations (middle). The bottom panel shows the difference between the top and middle one. As can be seen in most cases (84%, for spent media without replenishment) increasing nutrient concentrations lead to a stronger inhibition of the interaction partner (values below zero), however in the remaining cases it leads to a relative facilitation (values above zero). Spec_X_Sn_Y means species X was grown in supernatant of species Y.
Extended Data Fig. 5 Growth inhibition caused by high nutrient spent media is partially caused by pH and can be removed by buffering.
The scatter plots show the ratio of final OD in spent and final OD in fresh media for all 64 interaction pairs in buffered media at low (left) and high (right) nutrient concentrations. The solid lines and boxes show the corresponding mean and SEM. This figure is thus equivalent to Fig. 1c in the main text with higher buffer concentrations (100mM phosphate). The black circles show the data of Fig. 1c eg with lower buffer concentrations (10mM phosphate). As can be seen the presence of higher buffer concentrations slightly facilitates growth in spent, but not replenished media, possibly because adding phosphate avoids phosphor to be a limiting resource. However, the strongest effect of buffering can be seen in the replenished supernatant. Whereas there is no effect upon the low nutrient replenished supernatant, bacteria grow much better in high nutrient replenished media with higher buffer concentration compared to lower phosphate (one-sided t-test p-value = 0.006). Since in the replenished media nutrient competition as a mode of interaction does not matter, this shows that the growth hindering and thus toxic effect of replenished high nutrient media can partially be diminished by buffering. Thus, at least a part of the toxic effect of high nutrient supernatant is caused by pH.
Extended Data Fig. 6 Nutrient levels determine interaction strength.
The first three columns correspond to Fig. 1d. The fourth column shows the interaction outcomes for a medium nutrient concentration of 0.4% glucose and 0.32% urea eg. 0.4x the high nutrient condition. As expected the results fall in between the results for the low (no Glucose and Urea) and high (1% Glucose and 0.8% Urea) nutrient outcomes. PO4 means phosphate.
Extended Data Fig. 7 Complex nutrients weakly effect interaction.
Increasing the amount of yeast extract and soytone from 1g/L each to 20g/L leads to a slight decrease in overall diversity (p-value: 0.112). However, the effect of glucose and urea is much stronger. On reason for that may be that yeast extract and soytone also work as buffers, which stabilize pH at high nutrient concentrations. PO4 means phosphate.
Extended Data Fig. 8 Rarefaction curves for data of last day of complex community cultivation in high and low nutrient concentrations and alternative diversity metrics for complex communities.
(a)The curves become flat at the read depth of the samples (= end of curves) which shows that the read depth is sufficient to capture the species richness in the sample. The diversity for q=2 (2D diversity) (b) and richness (c), which puts more emphasis on common species shows the same effect of nutrients and buffering in the diversity as shown for the 1D diversity in Fig. 3.
Extended Data Fig. 9 Initial community compositions.
Shown are the ASVs with more then 0.05 abundance. The corresponding 1D diversity and richness are much higher than at the end of the experiments (Fig. 3), eg those communities collapsed to communities with lower diversity during the experiments. The sequencing of the initial soil community failed.
Extended Data Fig. 10 Community composition over time for different samples sites, replicates and nutrient conditions.
The colors that represent the different species are consistent for a specific sample (compost, flowerpot, soil), but may vary between them. In a few cases different ASVs were identified as the same species, which causes a connection of the same species name with different colors within the same sample site. The white columns indicate days for which the sequencing failed.
Supplementary information
Supplementary Information
Supplementary Figs. 1–13 and discussion.
Rights and permissions
About this article
Cite this article
Ratzke, C., Barrere, J. & Gore, J. Strength of species interactions determines biodiversity and stability in microbial communities. Nat Ecol Evol 4, 376–383 (2020). https://doi.org/10.1038/s41559-020-1099-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-020-1099-4
This article is cited by
-
Manipulating the physical distance between cells during soil colonization reveals the importance of biotic interactions in microbial community assembly
Environmental Microbiome (2024)
-
Changes in soil microbial community and co-occurrence network after long-term no-tillage and mulching in dryland farming
Plant and Soil (2024)
-
Degradation of wild fruit forests created less diverse and diffuse bacterial communities decreased bacterial diversity, enhanced fungal pathogens and altered microbial assembly in the Tianshan Mountain, China
Plant and Soil (2024)
-
Fungal communities are more sensitive to mildew than bacterial communities during tobacco storage processes
Applied Microbiology and Biotechnology (2024)
-
Rhizobial nitrogen fixation efficiency shapes endosphere bacterial communities and Medicago truncatula host growth
Microbiome (2023)