Abstract
Mutations in the rifampicin (Rif)-binding site of RNA polymerase (RNAP) confer antibiotic resistance and often have global effects on transcription that compromise fitness and stress tolerance of resistant mutants. We suggested that the non-essential genome, through its impact on the bacterial transcription cycle, may represent an untapped source of targets for combination antimicrobial therapies. Using transposon sequencing, we carried out a genome-wide analysis of fitness cost in a clinically common rpoB H526Y mutant. We find that genes whose products enable increased transcription elongation rates compound the fitness costs of resistance whereas genes whose products function in cell wall synthesis and division mitigate it. We validate our findings by showing that the cell wall synthesis and division defects of rpoB H526Y result from an increased transcription elongation rate that is further exacerbated by the activity of the uracil salvage pathway and unresponsiveness of the mutant RNAP to the alarmone ppGpp. We applied our findings to identify drugs that inhibit more readily rpoB H526Y and other RifR alleles from the same phenotypic class. Thus, genome-wide analysis of fitness cost of antibiotic-resistant mutants should expedite the discovery of new combination therapies and delineate cellular pathways that underlie the molecular mechanisms of cost.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All sequencing data generated in the Tn-seq and NET-seq experiments are publicly available through the sequence read archive (SRA) database of NCBI. Tn-seq data (Fig. 1, Extended Data Fig. 1 and Supplementary Table 1) are available through the SRA with the accession numbers SRX7174171 and SRX7174170. NET-seq data are available through the SRA with the accession numbers SRX7174164, SRX7174165, SRX7174166 and SRX7174167 (for data presented in Fig. 5) and SRX11385220, SRX11385221 and SRX11385222 (for data presented in Extended Data Fig. 9). The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD027810 (data presented in Extended Data Fig. 6c and Supplementary Table 6). All strains and plasmids constructed in this study are available on request from the authors. Source data are provided with this paper.
Code availability
All codes used in this study are available on request from the corresponding author.
References
Furin, J., Cox, H. & Pai, M. Tuberculosis. Lancet https://doi.org/10.1016/S0140-6736(19)30308-3 (2019).
Guinn, K. M. & Rubin, E. J. Tuberculosis: just the FAQs. Mbio 8, e01910-17 (2017).
Berrada, Z. L. et al. Rifabutin and rifampin resistance levels and associated rpoB mutations in clinical isolates of Mycobacterium tuberculosis complex. Diagn. Microbiol. Infect. Dis. 85, 177–181 (2016).
Campbell, E. A. et al. Structural mechanism for rifampicin inhibition of bacterial RNA polymerase. Cell 104, 901–912 (2001).
Molodtsov, V., Scharf, N. T., Stefan, M. A., Garcia, G. A. & Murakami, K. S. Structural basis for rifamycin resistance of bacterial RNA polymerase by the three most clinically important RpoB mutations found in Mycobacterium tuberculosis. Mol. Microbiol. 103, 1034–1045 (2017).
Lin, W. et al. Structural basis of Mycobacterium tuberculosis transcription and transcription inhibition. Mol. Cell 66, 169–179 (2017).
Jin, D. J. & Gross, C. A. Mapping and sequencing of mutations in the Escherichia coli rpoB gene that lead to rifampicin resistance. J. Mol. Biol. 202, 45–58 (1988).
Jin, D. J. & Gross, C. A. Characterization of the pleiotropic phenotypes of rifampin-resistant rpoB mutants of Escherichia coli. J. Bacteriol. 171, 5229–5231 (1989).
McDowell, J. C., Roberts, J. W., Jin, D. J. & Gross, C. Determination of intrinsic transcription termination efficiency by RNA polymerase elongation rate. Science 266, 822–825 (1994).
Jin, D. J. et al. Effects of rifampicin resistant rpoB mutations on antitermination and interaction with nusA in Escherichia coli. J. Mol. Biol. 204, 247–261 (1988).
Svetlov, V., Belogurov, G. A., Shabrova, E., Vassylyev, D. G. & Artsimovitch, I. Allosteric control of the RNA polymerase by the elongation factor RfaH. Nucleic Acids Res. 35, 5694–5705 (2007).
Zhou, Y. N. & Jin, D. J. The rpoB mutants destabilizing initiation complexes at stringently controlled promoters behave like ‘stringent’ RNA polymerases in Escherichia coli. Proc. Natl Acad. Sci. USA 95, 2908–2913 (1998).
Baym, M., Stone, L. K. & Kishony, R. Multidrug evolutionary strategies to reverse antibiotic resistance. Science 351, aad3292 (2016).
Imamovic, L. & Sommer, M. O. A. Use of collateral sensitivity networks to design drug cycling protocols that avoid resistance development. Sci. Transl. Med. 5, 204ra132 (2013).
Maltas, J. & Wood, K. B. Pervasive and diverse collateral sensitivity profiles inform optimal strategies to limit antibiotic resistance. PLoS Biol. 17, e3000515 (2019).
Lázár, V. et al. Bacterial evolution of antibiotic hypersensitivity. Mol. Syst. Biol. 9, 700 (2013).
Chait, R., Craney, A. & Kishony, R. Antibiotic interactions that select against resistance. Nature 446, 668–671 (2007).
Chevereau, G. & Bollenbach, T. Systematic discovery of drug interaction mechanisms. Mol. Syst. Biol. 11, 807 (2015).
Epshtein, V. et al. UvrD facilitates DNA repair by pulling RNA polymerase backwards. Nature 505, 372–377 (2014).
Goodman, A. L. et al. Identifying genetic determinants needed to establish a human gut symbiont in its habitat. Cell Host Microbe 6, 279–289 (2009).
van Opijnen, T., Bodi, K. L. & Camilli, A. Tn-seq: high-throughput parallel sequencing for fitness and genetic interaction studies in microorganisms. Nat. Methods 6, 767–772 (2009).
Nichols, R. J. et al. Phenotypic landscape of a bacterial cell. Cell 144, 143–156 (2011).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Bean, G. J. et al. A22 disrupts the bacterial actin cytoskeleton by directly binding and inducing a low-affinity state in MreB. Biochemistry 48, 4852–4857 (2009).
Iwai, N., Nagai, K. & Wachi, M. Novel S-benzylisothiourea compound that induces spherical cells in Escherichia coli probably by acting on a rod-shape-determining protein(s) other than penicillin-binding protein 2. Biosci. Biotechnol. Biochem. 66, 2658–2662 (2002).
Singh, V. et al. The complex mechanism of antimycobacterial action of 5-fluorouracil. Chem. Biol. 22, 63–75 (2015).
Bendezú, F. O. & de Boer, P. A. J. Conditional lethality, division defects, membrane involution, and endocytosis in mre and mrd shape mutants of Escherichia coli. J. Bacteriol. 190, 1792–1811 (2008).
Aberg, A., Fernández-Vázquez, J., Cabrer-Panes, J. D., Sánchez, A. & Balsalobre, C. Similar and divergent effects of ppGpp and DksA deficiencies on transcription in Escherichia coli. J. Bacteriol. 191, 3226–3236 (2009).
Kingston, R. E., Nierman, W. C. & Chamberlin, M. J. A direct effect of guanosine tetraphosphate on pausing of Escherichia coli RNA polymerase during RNA chain elongation. J. Biol. Chem. 256, 2787–2797 (1981).
Campodónico, V. L., Rifat, D., Chuang, Y.-M., Ioerger, T. R. & Karakousis, P. C. Altered Mycobacterium tuberculosis cell wall metabolism and physiology associated with RpoB mutation H526D. Front. Microbiol. 9, 494 (2018).
Liu, A. et al. Antibiotic sensitivity profiles determined with an Escherichia coli gene knockout collection: generating an antibiotic bar code. Antimicrob. Agents Chemother. 54, 1393–1403 (2010).
Břinda, K. et al. Rapid inference of antibiotic resistance and susceptibility by genomic neighbour typing. Nat. Microbiol. 5, 455–464 (2020).
Votintseva, A. A. et al. Same-day diagnostic and surveillance data for tuberculosis via whole-genome sequencing of direct respiratory samples. J. Clin. Microbiol. 55, 1285–1298 (2017).
Ferrières, L. et al. Silent mischief: bacteriophage Mu insertions contaminate products of Escherichia coli random mutagenesis performed using suicidal transposon delivery plasmids mobilized by broad-host-range RP4 conjugative machinery. J. Bacteriol. 192, 6418–6427 (2010).
Chiang, S. L. & Rubin, E. J. Construction of a mariner-based transposon for epitope-tagging and genomic targeting. Gene 296, 179–185 (2002).
Langmead, B. Aligning short sequencing reads with Bowtie. Curr. Protoc. Bioinform. https://doi.org/10.1002/0471250953.bi1107s32 (2010).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Moore, S. D. Assembling new Escherichia coli strains by transduction using phage P1. Methods Mol. Biol. 765, 155–169 (2011).
Reisch, C. R. & Prather, K. L. J. The no-SCAR (Scarless Cas9 Assisted Recombineering) system for genome editing in Escherichia coli. Sci. Rep. 5, 15096 (2015).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Rosche, W. A. & Foster, P. L. Determining mutation rates in bacterial populations. Methods 20, 4–17 (2000).
Liu, M. et al. Global transcriptional programs reveal a carbon source foraging strategy by Escherichia coli. J. Biol. Chem. 280, 15921–15927 (2005).
Kamarthapu, V. et al. ppGpp couples transcription to DNA repair in E. coli. Science 352, 993–996 (2016).
Davis, S. et al. Expanding proteome coverage with charge ordered parallel ion analysis (CHOPIN) combined with broad specificity proteolysis. J. Proteome Res. 16, 1288–1299 (2017).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Larson, M. H. et al. A pause sequence enriched at translation start sites drives transcription dynamics in vivo. Science 344, 1042–1047 (2014).
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Dar, D. & Sorek, R. High-resolution RNA 3′-ends mapping of bacterial Rho-dependent transcripts. Nucleic Acids Res. 46, 6797–6805 (2018).
Studier, F. W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Borukhov, S. & Goldfarb, A. Recombinant Escherichia coli RNA polymerase: purification of individually overexpressed subunits and in vitro assembly. Protein Expr. Purif. 4, 503–511 (1993).
Banerjee, R., Rudra, P., Prajapati, R. K., Sengupta, S. & Mukhopadhyay, J. Optimization of recombinant Mycobacterium tuberculosis RNA polymerase expression and purification. Tuberc. Edinb. Scotl. 94, 397–404 (2014).
Acknowledgements
This study was supported by NIH grants T32 AI007180 (K.T.); R01 AI099394, R01 AI121244 and R01 AI105129 (V.J.T.); R01 GM126891 (E.N.); DoD grant PR171734 (E.N.); and the Blavatnik Family Foundation and Howard Hughes Medical Institute (E.N.). We thank L. Ash from the NYU Metabolomics Core Resource Laboratory for technical assistance.
Author information
Authors and Affiliations
Contributions
A.R. and E.N. conceptualized this study. A.R. and E.N. designed the experiments. A.R. constructed bacterial strains. A.R., Y.S., S.K., I.S., A.S. and S.V. performed sequencing experiments, stress-sensitivity assays and fluctuation tests. Y.S. performed phenotypic plate array screening. Z.H. and L.I. performed all protein purifications. N.V. performed proteomic experiments. V.E. performed in vitro transcription experiments. B.P. performed microscopy experiments. G.Q. performed measurements of ppGpp concentrations in cell lysates. A.R., K.T. and V.J.T. designed the human serum and animal experiments and K.T. performed these experiments. A.R., Y.S., V.E., N.V., K.T., C.V., I.S., V.J.T. and E.N. analysed data. A.R. and E.N. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genome-wide fitness cost analysis in RifR rpoB H526Y.
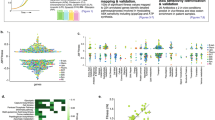
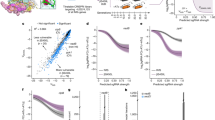
a, Genome-wide normalized transposon insertion density (IPKM) values. Each data point corresponds to a single gene. Genes are ordered on the x-axis based on distance from oriC, and are stratified according to previous annotation of gene essentiality and dispensability in wild type E. coli MG1655. b, Insertion saturation analysis. Each bar provides the number of genes with a given transposon insertion saturation value. Gene essentiality is colour-coded. Total number of reads and overall saturation percentage are indicated. c, Distribution of gene number as a function of IPKM values. Per gene, the IPKM value is the sum of reads mapped to transposon insertion sites per mean number of TA sites in kilobases per million reads. Gene essentiality is colour-coded. d, positive correlation between fold IPKM value of top cost-mitigating genes included in Supplementary Table 1 and fold IPKM values of the same genes from Supplementary Table 2. e, Gene Ontology (GO) enrichment analysis for biological processes that are either cost mitigating (blue) or compounding (red). f, Gene browser comparative view of transposon insertion sites of cost mitigating genes in wild type (black) and rpoB H526Y (blue) at the zapB, dacA, and relA loci. g, same as in (f) for cost-compounding genes in wild type (black) and RifR H526Y (red) at the rsgA, rlmE, and gpp loci. h, Growth in LB of single gene knockouts in genetic background of wild type or H526Y.
Extended Data Fig. 2 Light Microscopy imaging of cell division defect in the synthetic lethal strain.
wild type, H526Y, yebAzapA (ΔΔ), and the synthetic lethal strain (SL) before (ON) and after (OFF) the depletion of yebA from a complementation plasmid (pY). Scale bar denotes 10μm length. The mean for each group is indicated with a solid horizontal line. **** p-value <0.0001. p-values were calculated with the student t-test (two-sided, equal variance). Cell length distribution is based on at least 80 cells from each culture. For all microscopy imaging experiments, n = 3 biological replicates.
Extended Data Fig. 3 Specific synthetic lethal interaction between RifR rpoB H526Y and yebA zapA (ΔΔ).
a, Transduction of RifR rpoB H526Y to yebA zapA depends on a complementation plasmid expressing zapA. b, Poor co-transduction efficiency of RifR rpoB H526Y cells to multiple double knockout combinations of top cost mitigating genes from the Tn-seq analysis. Donor P1 phage lysates were prepared either from RifR rpoB H526Y (with linked kanamycin resistance cassette downstream of the 3’ of rpoC) or from RifS strain with the same linked selectable marker. The efficiency of transducing the Km cassette to indicated recipients (wild type and selected double knockouts) was calculated by dividing the number of transductants to the number of recipient cells. c, serial dilution and plating of the indiciated strains. Unlike the synthetic lethal interaction between RifR rpoB H526Y with yebA zapA (ΔΔ), the RifR rpoB S531F is compatible with the yebAzapA double deletion without any growth defect.
Extended Data Fig. 4 Specific Targeting of the H526Y phenotypic class of RifR mutants.
Increased sensitivity of RifR cells harbouring alleles H526Y, D516G, and S522F to A22 (1 μg/ml), 5FUd (5 μg/ml) and high temperature. The S531F is included as an example of a RifR allele that does not share the same phenotypic profile. Mean and standard errors of all plating efficiency data are based on colony forming unit values (c.f.u) from three biological triplicates. * p-value <0.05, ** p-value <0.01, *** p-value <0.001. p-values are based on the student t-test (two-sided, equal variance) b, Growth of wild type and rpoB H526Y cells in LB supplemented with increasing concentration of human serum, with or without subinhibitory concentration of A22. c, Bacterial load in the peritoneal lavage, spleen, and liver in mice 24 h after infection with either wild type (triangle-shaped symbols) or H526Y cells (circle-shaped symbols). For each strain, the two groups of mice (n = 10 mice/group) received either no drug (clear symbols) or 20 mg/kg A22 one-hour postinfection (filled symbols). The median for each group is indicated with a solid horizontal line. Dotted lines indicate limit of detection. * p-value <0.05, ** p-value <0.01, n.s. not significant. d, Phenotypic profiling of wild type and rpoB H526Y cells against a collection of 56 antibiotics and antimicrobials with a known mode of action. colour code is according to the cellular target of each drug. Abbreviations of each class of drugs: Aminoglycosides (AG), Cell wall synthesis and division inhibitors (CW), Quinolones (QUI), genotoxic compounds (GNTX), DNA polymerase starving molecules (REP), Tetracyclines (TETC), Macrolides (MCLD), rifamycins (RIF), Antimicrobial peptides (AMP), and in a miscellaneous group (MICEL). Arrows point to antibiotics that were selected for MIC analysis across the nine different strains that are presented in Fig.3i.
Extended Data Fig. 5 Rescue of A22 and 5FUd sensitivity of additional RifR mutants with moderate overexpression of relA.
a, Degenerated Shine–Dalgarno (SD) sequence in the ‘weak’ SD vector provides low translation rates in comparison to an optimal ‘strong’ SD sequence. the two vectors are otherwise identical. b, Growth of wt and rpoB H526Y cells with the indicated concentration of inducer (aTc) for the expression of relA from a ‘weak’-SD plasmid. c, same as in (b) except that relA is expressed from a ‘strong’-SD plasmid. d, Plating efficiency values from serial dilution of overnight cultures of wt, rpoB D516G and rpoB D516G carrying prelA (for moderate overexpression of relA) on LB-agar supplemented with either A22 (1 μg/ml) or 5fUd (15 μg/ml) for 16 h at 37 C. e, same as in (a) except with S522F and S522F prelA, and that 5FUd concentration is 5μg/ml. For both d and e, Mean and standard errors of all plating efficiency data are based on colony forming unit values (c.f.u) from three biological triplicates (n = 3). * p-value <0.05, ** p-value <0.01, *** p-value <0.001. p-values are based on the student t-test (two-sided, equal variance).
Extended Data Fig. 6 Indifference of rpoB H526Y cells to steady-state ppGpp concentration underlies its fitness cost.
a, Sensitivity of the indicated strains to A22, 5FUd and high temperature. b, Robust growth of wild type and rpoB H526Y versus auxotrophy of relAspoT on minimal agar plates. c, Proteomic analysis of differentially expressed proteins between rpoB H526Y and relAspoT (ppGpp0). Venn diagram presenting the overlap between differentially expressed proteins in rpoB H526Y and relAspoT cells in comparison to the wild type strain. A full list of genes is given in Supplementary Table 6. The total number of proteins detected across all three strains is 2174. The overlap between differentially regulated proteins in rpoB H526Y and relAspoT cells, in comparison to wild type cells, has p-values of 1.1×10−4 and 1.29×10−8, respectively, for up- and down-regulated proteins. d, ppGpp concentration in lysates prepared from two biological replicates of wild type and rpoB H526Y.
Extended Data Fig. 7 High concentrations of ppGpp slow-down the ‘fast’ H526Y RNAP in vitro.
a, In vitro transcription reactions of E. coli RifR rpoB H526Y RNAP with increasing concentrations of ppGpp (0,0.1 and 1 mM). At the highest ppGpp concentration, reactions were performed with or without DksA. Stalled RNAP elongation complexes (20-mer, denoted with the red arrow) were chased to the end of the template (runoff, denoted with the blue arrow), and the labelled nascent RNA then separated using PAGE. For in vitro transcription experiments, n = 3 independent replicates. b, Quantification plot of signal from gel scans as a function of distance from the initial site of stalling (right-end on the x-axis). Arrows (labelled 1 through 6) correspond to major pause sites.
Extended Data Fig. 8 Desensitization of M. tuberculosis H445Y RNAP to ppGpp in vitro.
a, In vitro transcription reactions of M. tuberculosis wild type and RifR rpoB H445Y RNAP with or without ppGpp (10 mM). Stalled RNAP elongation complexes (20-mer, denoted with the red arrow) were chased to the end of the template (runoff, denoted with the blue arrow), and the labelled nascent RNA then separated using PAGE. For in vitro transcription experiments, n = 3 independent replicates. b, Quantification plot of signal from the gel scans as a function of distance from the initial site of stalling (right-end on the x-axis). Arrows (labelled 1 through 5) correspond to major pause sites.
Extended Data Fig. 9 Mean RNAP pause frequency in the upp genetic background.
NET-seq analysis of RNAP pausing in wild type, upp, and uppH526Y double mutant. For each gene, the mean number of pauses per kilobase gene (the pause frequency) was calculated. The midline in each box denotes the aggregated median frequency for the top 3500 expressed genes for each condition. The lower and upper ends of the box denote Q1 and Q3, respectively. The whiskers span 1.5*(Q3–Q1) from each side of the box. **** p-value <0.0001, N.S not significant. p-values are based on the student t-test (two-sided, equal variance). For each condition, n = 2 biologically independent replicates of NET-seq experiments.
Extended Data Fig. 10 Rifampicin resistance desensitizes elongating RNAP to ppGpp.
a, Same image from Fig.5e with b, quantification plots. In vitro transcription reactions of wild type and rpoB H526Y RNAP with or without ppGpp (100 μM). Stalled RNAP elongation complexes (20-mer, denoted with the red arrow) were chased to the end of the template (runoff, denoted with the blue arrow), and the labelled nascent RNA then separated using PAGE. Numbering corresponds to time points 0, 10, 20, 40 and 60-seconds following the chase reaction. Arrows (labelled 1 through 5) correspond to major pause sites.
Supplementary information
Supplementary Tables
Supplementary Tables 1–3 and 7.
Source data
Source Data Fig. 1
Raw numerical data.
Source Data Fig. 2
Raw numerical data.
Source Data Fig. 3
Raw numerical data.
Source Data Fig. 4
Raw numerical data.
Source Data Fig. 5
Raw numerical data.
Source Data Fig. 5
Unprocessed scan of in vitro transcription gel.
Source Data Extended Data Fig. 1
Raw numerical data.
Source Data Extended Data Fig. 2
Raw numerical data.
Source Data Extended Data Fig. 3
Raw numerical data.
Source Data Extended Data Fig. 4
Raw numerical data.
Source Data Extended Data Fig. 5
Raw numerical data.
Source Data Extended Data Fig. 6
Raw numerical data.
Source Data Extended Data Fig. 6
Unprocessed scan of TLC plate.
Source Data Extended Data Fig. 7
Unprocessed scan of in vitro transcription gel.
Source Data Extended Data Fig. 8
Unprocessed scan of in vitro transcription gel.
Source Data Extended Data Fig. 9
Raw numerical data.
Source Data Extended Data Fig. 10
Unprocessed scan of in vitro transcription gel.
Rights and permissions
About this article
Cite this article
Rasouly, A., Shamovsky, Y., Epshtein, V. et al. Analysing the fitness cost of antibiotic resistance to identify targets for combination antimicrobials. Nat Microbiol 6, 1410–1423 (2021). https://doi.org/10.1038/s41564-021-00973-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-00973-1
This article is cited by
-
General transcription factor from Escherichia coli with a distinct mechanism of action
Nature Structural & Molecular Biology (2024)
-
Gut Distribution, Impact Factor, and Action Mechanism of Bacteriocin-Producing Beneficial Microbes as Promising Antimicrobial Agents in Gastrointestinal Infection
Probiotics and Antimicrobial Proteins (2024)
-
Long-term exposure to food-grade disinfectants causes cross-resistance to antibiotics in Salmonella enterica serovar Typhimurium strains with different antibiograms and sequence types
Antimicrobial Resistance & Infection Control (2023)
-
High-resolution landscape of an antibiotic binding site
Nature (2023)
-
Control of transcription elongation and DNA repair by alarmone ppGpp
Nature Structural & Molecular Biology (2023)